Abstract
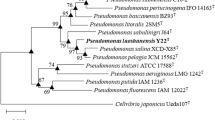
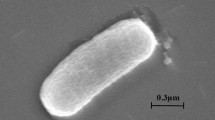
A Gram-negative, non-motile, catalase-positive and oxidase-positive, aerobic bacterium designated as NII-0918T was isolated from soil sample in Western ghat forest, India. 16S rRNA gene sequence analysis showed that strain NII-0918T belongs to the subclass α-Proteobacteria, being related to the genus Paracoccus, and sharing highest sequence similarity with Paracoccus chinensis NBRC 104937T (99.4%), Paracoccus marinus NBRC 100640T (97.3%), Paracoccus koreensis Ch05T (97.1%) and Paracoccus kondratievae GBT (97.0%). Other members of Paracoccus showed below 97.0% similarity. The DNA–DNA hybridization values between these four strains and NII-0918T were 44.7, 28, 32 and 41%, respectively. The major fatty acids of strain NII-0918T were summed feature 7 (C18:1 ω7c/ω 9t/ω 12t) (83.0%) and C18:0 (12.5%). Ubiquinone Q-10 was detected as the major respiratory quinone. The G+C content of genomic DNA of NII-0918T was 66.6 mol%. On the basis of physiological, morphological, chemotaxonomical and DNA–DNA hybridization data, it is proposed that strain NII-0918T should be placed as a novel species, for which we propose Paracoccus niistensis sp. nov. The type strain is NII-0918T (CCTCC AA 209055T = NCIM 5340T = KCTC 22789T).

Similar content being viewed by others
References
Chen MH, Sheu SY, Chen CA, Wang JT, Chen WM (2010) Paracoccus isoporae sp. nov., isolated from the reef-building coral Isopora palifera. Int J Syst Evol Microbiol. doi:ijs.0.023333-0v1-ijs.0.023333-0
Chun J, Lee J-H, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequence. Int J Syst Evol Microbiol 57:2259–2261
Daneshvar MI, Hollis DG, Weyant RS, Steigerwalt AG, Whitney AM (2003) Paracoccus yeeii sp. nov. (formerly CDC Group EO-2), a novel bacterial species associated with human infection. J Clin Microbiol 41:1289–1294
Davis DH, Doudoroff M, Stanier RY, Mandel M (1969) Proposal to reject the genus Hydrogenomonas: taxonomic implications. Int J Syst Bacteriol 19:375–390
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Deng ZS, Zhao LF, Xu L, Kong ZY, Zhao P, Qin W, Chang JL, Wei GH (2010) Paracoccus sphaerophysae sp. nov., a siderophore-producing, endophytic bacterium isolated from root nodules of a Sphaerophysa salsula growing in northwestern China. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.021071-0
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Conference limits on phylogenies: an approach using the bootstrap. Evolution 39:783–789
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specified tree topology. Syst Zool 20:406–416
Fitch WM, Margoliash E (1967) Construction of a phylogenetic tree. Science 155:279–284
Gerhardt P, Krieg NR (1981) General characterization. In: Krieg NR (ed) Manual of methods for general bacteriology. American Society for Microbiology, Washington, DC, pp 409–443
Huß VAR, Festl H, Schleifer KH (1983) Studies on the spectrophotometric determination of DNA hybridization from renaturation rates. Syst Appl Microbiol 4:184–192
Jahnke KD (1992) BASIC computer program for evaluation of spectroscopic DNA renaturation data from GILFORD SYSTEM 2600 spectrophotometer on a PC/XT/AT type personal computer. J Microbiol Methods 15:61–73
John P, Whatley FR (1975) Paracoccus denitrificans and the evolutionary origin of the mitochondrion. Nature 254:495–498
Kelly DP, Rainey FA, Wood AP (2006) The genus Paracoccus. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes. A handbook on the biology of bacteria, vol. 5, 3rd edn. Springer, New York, pp 232–249
Khan ST, Takaichi S, Harayama S (2008) Paracoccus marinus sp. nov., an adonixanthin diglucoside-producing bacterium isolated from coastal seawater in Tokyo Bay. Int J Syst Evol Microbiol 58:383–386
Kim YO, Kong HJ, Park S, Kang SJ, Kim KK, Moon DY, Oh TK, Yoon JH (2010) Paracoccus fistulariae sp. nov., a lipolytic bacterium isolated from bluespotted cornetfish, Fistularia commersonii. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.021808-0
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kluge AG, Farris FS (1969) Quantitative phyletics and the evolution of anurans. Syst Zool 18:1–32
Kroppenstedt RM (1982) Separation of bacterial menaquinones by HPLC using reverse phase (RP 18) and a silver loaded ion exchange as stationary phases. J Liq Chromatogr 5:2359–2387
Li WJ, Xu P, Schumann P, Zhang YQ, Pukall R, Xu LH, Stackebrandt E, Jiang CL (2007) Georgenia ruanii sp. nov., a novel actinobacterium isolated from forest soil in Yunnan (China) and emended description of the genus Georgenia. Int J Syst Evol Microbiol 57:1424–1428
Li H-F, Qu J-H, Yang J-S, Li J-Z, Yuan H-L (2009) Paracoccus chinensis sp. nov., isolated from sediment of a reservoir. Int J Syst Evol Microbiol 59:2670–2674
Liu ZP, Wang BJ, Liu XY, Dai X, Liu YH, Liu SJ (2008) Paracoccus halophilus sp. nov., isolated from marine sediment of the South China Sea, China, and emended description of the genus Paracoccus Davis 1969. Int J Syst Evol Microbiol 58:257–261
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from microorganisms. J Mol Biol 3:208–218
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Bacteriol 39:159–167
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of isoprenoid quinines and polar lipids. J Microbiol Methods 2:233–241
Roh SW, Nam YD, Chang HW, Kim KH, Kim MS, Shin KS, Yoon JH, Oh HM, Bae JW (2009) Paracoccus aestuarii sp. nov., isolated from tidal flat sediment. Int J Syst Evol Microbiol 59:790–794
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic tree. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. USFCC Newsl 20:1–6
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general, molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Stackebrandt E, Goebel BM (1994) Taxonomic Note: a place for DNA–DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evolu 24:1596–1599
Wang Y, Tang SK, Lou K, Mao PH, Jin X, Jiang CL, Xu LH, Li WJ (2009) Paracoccus saliphilus sp. nov., a halophilic bacterium isolated from a saline soil. Int J Syst Evol Microbiol 59:1924–1928
Acknowledgment
The authors would like to thank CSIR Task force network programme on Exploration of India’s Rich Microbial Diversity (NWP 0006) for providing the financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dastager, S.G., Deepa, C.K., Li, WJ. et al. Paracoccus niistensis sp. nov., isolated from forest soil, India. Antonie van Leeuwenhoek 99, 501–506 (2011). https://doi.org/10.1007/s10482-010-9515-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-010-9515-4