Abstract
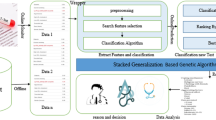
This paper introduces a simple strategy for diagnosing disease, which is called improved gray wolf optimization (IGWO) and ensemble classification. The proposed strategy consists of two sequential phases, which are; (i) Feature Selection Phase (FSP) and (ii) Ensemble Classification Phase (ECP). During the former, the most effective features for diagnosing disease are selected, while during the latter, the actual diagnosis takes place depending on voting of five different classifiers. The main contribution of this paper is a suggested modification for the traditional Gray Wolf Optimization (GWO), which is called Improved Gray Wolf Optimization (IGWO). As an optimization technique, the proposed IGWO is employed in the FSP for selecting the effective features. For evaluating, IGWO has been implemented using recent feature selection techniques as well as the proposed method. To accomplish the classification phase; ensemble classification has been used which uses several classification techniques such as; Naïve Bayes (NB), Support Vector Machines (SVM), Deep Neural Network (DNN), Decision Tree (DT), and K-Nearest Neighbors (KNN). Ensemble classification integrate several classifiers for improving prediction performance. Experimental results have shown that employing IGWO promotes the performance of the diagnosing strategy of different diseases in terms of precision, recall, and accuracy.

































Similar content being viewed by others
References
Moreno-Ibarra, M.-A., Y. Villuendas-Rey, et al. Classification of diseases using machine learning algorithms: a comparative study. Mathematics. 2021:9, 1817. https://doi.org/10.3390/math9151817.
Elghamrawy, S. M., and A. E. Hassanien. A hybrid Genetic-Grey Wolf Optimization algorithm for optimizing Takagi–Sugeno–Kang fuzzy systems. Neural Comput. Appl. 34(19):17051–69, 2022.
Kumar, Yogesh, Apeksha Koul, et al. Artificial intelligence in disease diagnosis: a systematic literature review, synthesizing framework and future research agenda. J. Ambient Intell. Hum. Comput. 2022. https://doi.org/10.1007/s12652-021-03612-z.
Mirbabaie, M., S. Stieglitz, and N. R. Frick. Artificial intelligence in disease diagnostics: A critical review and classification on the current state of research guiding future direction. Health Technol. 11(4):693–731, 2021.
Singh, A., J. C. Mehta, D. Anand, P. Nath, B. Pandey, and A. Khamparia. An intelligent hybrid approach for hepatitis disease diagnosis: combining enhanced k -means clustering and improved ensemble learning. Expert Syst. 2020. https://doi.org/10.1111/exsy.12526.
Frick, N. R. J., H. L. Möllmann, M. Mirbabaie, and S. Stieglitz. Driving digital transformation during a pandemic: case study of virtual collaboration in a German Hospital. JMIR Med. Inform. 9(2):e25183, 2021.
Mirbabaie, M., S. Stieglitz, and N. R. Frick. Hybrid intelligence in hospitals: towards a research agenda for collaboration. Electron. Markets. 31:365–87, 2021.
ani SU, Khan NA, Thakur G, Gautam SP, Ali M, Alam P, Alshehri S, Ghoneim MM, Shakeel F. Utilization of artificial intelligence in disease prevention: Diagnosis, treatment, and implications for the healthcare workforce. In Healthcare 2022 Mar 24 (Vol. 10, No. 4, p. 608). MDPI. https://doi.org/10.3390/healthcare10040608.
Sarkar, Tanmay, Molla Salauddin, et al. Application of bio-inspired optimization algorithms in food processing. Curr. Res. Food Sci. 5:432–450, 2022.
Johnvictor, A. C., V. Durgamahanthi, R. M. Pariti Venkata, and N. Jethi. Critical review of bio-inspired optimization techniques. Wiley Interdiscip. Rev. Comput. Stat. 14(1):1528, 2022.
Mirjalili, S., and A. Lewis. S-shaped versus V-shaped transfer functions for binary particle swarm optimization. Swarm Evol. Comput. 9:1–14, 2013.
Luan, F., Z. Cai, S. Wu, T. Jiang, F. Li, and J. Yang. Improved whale algorithm for solving the flexible job shop scheduling problem. Mathematics. 7(5):384, 2019.
Pan, C., A. Jin, W. Yang, and Y. Zhang. Early detection of network fault using improved Gray Wolf Optimization and wavelet neural network. Hindawi Math. Probl. Eng. 2022. https://doi.org/10.1155/2022/1235229.
Mirjalili, S. Moth-flame optimization algorithm: a novel nature-inspired heuristic paradigm. Knowl. Based Syst. 89:228–249, 2015.
Wang G. A comparative study of cuckoo algorithm and ant colony algorithm in optimal path problems. Paper presented at MATECweb of conferences, 232, 03003. EITCE (2018).
Mirjalili, S. The ant lion optimizer. Adv. Eng. Softw. 83:80–98, 2015.
Mahdy, A.M.S., Lotfy, et al., “Analytical solution of magneto-photothermal theory during variable thermal conductivity of a semiconductor material due to pulse heat flux and volumetric heat source”, Waves Random Complex Media, 2021, 31, 2040–2057.
Khamis, A. K., K. Lotfy, et al. Thermal-piezoelectric problem of a semiconductor medium during photo-thermal excitation. Waves Random Complex Media. 31:2499–2513, 2021.
Hou, Y., H. Gao, et al. Improved Grey Wolf optimization algorithm and application. Sensors. 22:3810, 2022. https://doi.org/10.3390/s22103810.
Shen, C., and K. Zhang. Two-stage improved Grey Wolf optimization algorithm for feature selection on high-dimensional classification. Complex Intell. Syst. 2021. https://doi.org/10.1007/s40747-021-00452-4.
Feature selection using binary particle swarm optimization and support vector machines for medical diagnosis, Biomedizinische Technik/Biomedical Engineering, 2012.
Predicting the cognitive states of the subjects in functional magnetic resonance imaging signals using the combination of feature selection strategies, Brain Topography, 2012.
VijayAnand, M., B. KiranBala, et al. Gaussian Naïve Bayes Algorithm: a reliable technique involved in the assortment of the segregation in cancer. Hindawi Mob. Inf. Syst. 2022. https://doi.org/10.1155/2022/2436946.
Rodríguez-Pérez, R., and J. Bajorath. Evolution of support vector machine and regression modeling in chemoinformatics and drug discovery. J. Comput. Aided Mol. Des. 2022. https://doi.org/10.1007/s10822-022-00442-9.
Song, Y., X. Kong, and C. Zhang. A large-scale-nearest neighbor classification algorithm based on neighbor relationship preservation. Wirel. Commun. Mob. Comput. 2022. https://doi.org/10.1155/2022/7409171.
Salehi, A., P. Baglat, and G. Gupta. Review on machine and deep learning models for the detection and prediction of Coronavirus. Mater. Today: Proc. 2020. https://doi.org/10.1016/j.matpr.2020.06.245.
Gupta, B., A. Rawat, et al. Analysis of various decision tree algorithms for classification in data mining. Int. J. Comput. Appl. 163(8):15–19, 2017.
Frick, N. R. J., M. Mirbabaie, S. Stieglitz, and J. Salomon. Maneuvering through the stormy seas of digital transformation: the impact of empowering leadership on the AI readiness of enterprises. J. Decis. Syst. 2021. https://doi.org/10.1080/12460125.2020.1870065.
Rauschert, S., K. Raubenheimer, P. E. Melton, and R. C. Huang. Machine learning and clinical epigenetics: a review of challenges for diagnosis and classification. Clin. Epigenet. 2020. https://doi.org/10.1186/s13148-020-00842-4.
Mishra S, Yamasaki T, Imaizumi H. Supervised classifcation of Dermatological diseases by Deep learning. 2018,1–6.
Jin, Y., C. Qin, Y. Huang, W. Zhao, and C. Liu. Multi-domain modeling of atrial fibrillation detection with twin attentional convolutional long short-term memory neural networks. Knowl. Based Syst. 193:5460, 2020.
Lu, J., E. Song, A. Ghoneim, and M. Alrashoud. Machine learning for assisting cervical cancer diagnosis: An ensemble approach. Futur. Gener. Comput. Syst. 106:199–205, 2020.
Ding, S., S. Hu, et al. A homogeneous ensemble method for predicting gastric cancer based on gastroscopy reports. Expert Syst. 37:1–14, 2020.
Dutta, A., T. Batabyal, M. Basu, and S. T. Acton. An efficient convolutional neural network for coronary heart disease prediction. Expert Syst. Appl. 159:113408, 2020. https://doi.org/10.1016/j.eswa.2020.113408.
Hamedan, F., A. Orooji, H. Sanadgol, and A. Sheikhtaheri. Clinical decision support system to predict chronic kidney disease: a fuzzy expert system approach. Int. J. Med. Inform. 138:104134, 2020. https://doi.org/10.1016/j.ijmedinf.2020.
Karabayir, I., S. M. Goldman, S. Pappu, and O. Akbilgic. Gradient boosting for Parkinson’s disease diagnosis from voice recordings. BMC Med. Inform. Decis. Mak. 20:228, 2020.
Senturk, Z. K. Early diagnosis of Parkinson’s disease using machine learning algorithms. Med. Hypotheses. 138:109603, 2020. https://doi.org/10.1016/j.mehy.2020.109603.
Erkan, U., and D. N. H. Thanh. Autism spectrum disorder detection with machine learning methods. Curr. Psychiatry Res. Rev. 15:297–308, 2019. https://doi.org/10.2174/2666082215666191111121115.
Yuan, K. C., L. W. Tsai, et al. The development an artificial intelligence algorithm for early sepsis diagnosis in the intensive care unit. Int. J. Med. Inform. 141:104176, 2020. https://doi.org/10.1016/j.ijmedinf.2020.104176.
Uchino, E., K. Suzuki, et al. Classification of glomerular pathological findings using deep learning and nephrologist–AI collective intelligence approach. Int. J. Med. Inform. 141:104231, 2020. https://doi.org/10.1016/j.ijmedinf.2020.104231.
Steinbuss, G., K. Kriegsmann, and M. Kriegsmann. Identifcation of gastritis subtypes by convolutional neuronal networks on histological images of antrum and corpus biopsies. Int. J. Mol. Sci. 21:6652, 2020. https://doi.org/10.3390/ijms21186652.
Laurentinus et al., “Design Fuzzy Expert System And Certainty Factor In Early Detection Of Stroke Disease”, 2020 8th International Conference on Cyber and IT Service Management (CITSM), 2020, pp. 1–7. https://doi.org/10.1109/CITSM50537.2020.9268830.
Chen Y, Li M, et al. Classification of glomerular spikes using Convolutional Neural Network. Proc. Conf Artif Intell Healthc. New York, NY, USA: ACM. 2020; 2020:254–8, https://doi.org/10.1145/3433996.3434043.
Nithya, A., A. Ahilan, N. Venkatadri, D. Ramji, and A. Palagan. Kidney disease detection and segmentation using artifcial neural network and multi kernel k-means clustering for ultrasound images. Measurement. 149:106952, 2019. https://doi.org/10.1016/j.measurement.2019.106952.
Khan, A., M. Khan, F. Ahmed, et al. Gastrointestinal diseases segmentation and classification based on duo-deep architectures. Pattern Recognit Lett. 131:193–204, 2020. https://doi.org/10.1016/j.patrec.2019.12.024.
Gouda, W., and R. Yasin. COVID-19 disease: CT pneumonia analysis prototype by using artificial intelligence, predicting the disease severity. J Radiol Nucl Med. 51:196, 2020. https://doi.org/10.1186/s43055-020-00309-9.
Vasal, S., S. Jain, and A. Verma. COVID-AI: an artificial intelligence system to diagnose COVID 19 disease. J Eng Res Technol. 9:1–6, 2020.
Kanegae, H., K. Suzuki, et al. Highly precise risk prediction model for new onset hypertension using artificial neural network techniques. J Clin Hypertens. 22:445–450, 2020. https://doi.org/10.1111/jch.13759.
Lai, N., W. Shen, C. Lee, J. Chang, M. Hsu, et al. Comparison of the predictive outcomes for anti-Alzheimer drug-induced hepatotoxicity by different machine learning techniques. Comput Methods Programs Biomed. 188:307, 2020. https://doi.org/10.1016/j.cmpb.2019.105307.
Sarao, V., D. Veritti, and L. Paolo. Automated diabetic retinopathy detection with two diferent retinal imaging devices using artificial intelligence. Graefe’s Arch Clin Exp Opthamol. 2020. https://doi.org/10.1007/s00417-020-04853-y.
Khan, A., and S. Zubair. An improved multi-modal based machine learning approach for the prognosis of Alzheimer’s disease. J. King Saud Univ. Comput. Inf. Sci. 2020. https://doi.org/10.1016/j.jksuci.2020.04.004.
Isravel, D. P., and S. V. P. D. Silas. Improved heart disease diagnostic IoT model using machine learning techniques. Neuroscience. 9:4442–4446, 2020.
Rodrigues, D. A., R. F. Ivo, S. C. Satapathy, S. Wang, J. Hemanth, and P. P. R. Filho. A new approach for classification skin lesion based on transfer learning, deep learning, and IoT system. Pattern Recogn. Lett. 136:8–15, 2020. https://doi.org/10.1016/j.patrec.2020.05.019.
http://covid19.nilehi.edu.eg.
https://www.kaggle.com/datasets/abhi8923shriv/liver-disease-patient-dataset.
Abasabadi, S., H. Nematzadeh, H. Motameni, et al. Hybrid feature selection based on SLI and genetic algorithm for microarray datasets. J. Supercomput. 2022. https://doi.org/10.1007/s11227-022-04650-w.
Kamel, S. R., and R. Yaghoubzadeh. Feature selection using grasshopper optimization algorithm in diagnosis of diabetes disease. Inform. Med. Unlocked. 2021. https://doi.org/10.1016/j.imu.2021.100707.
Rabie, A. H., A. I. Saleh, and N. A. Mansour. A Covid-19’s integrated herd immunity (CIHI) based on classifying people vulnerability. Comput. Biol. Med. 2021. https://doi.org/10.1016/j.compbiomed.2021.105112.
Kundu, R., S. Chattopadhyay, E. Cuevas, and R. Sarkar. AltWOA: Altruistic Whale Optimization Algorithm for feature selection on microarray datasets. Comput. Biol. Med. 2022. https://doi.org/10.1016/j.compbiomed.2022.105349,Volume144.
Pashaei, E., and E. Pashaei. An efficient binary chimp optimization algorithm for feature selection in biomedical data classification. Neural Comput. Appl. 34:6427–6451, 2022. https://doi.org/10.1007/s00521-021-06775-0.
Lan, P., K. Xia, et al. An improved GWO algorithm optimized RVFL model for oil layer prediction. Electronics. 10:3178, 2021. https://doi.org/10.3390/electronics10243178.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Associate Editor Mona Kamal Marei oversaw the review of this article.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Saleh, A.I., Hussien, S.A. Disease Diagnosis Based on Improved Gray Wolf Optimization (IGWO) and Ensemble Classification. Ann Biomed Eng 51, 2579–2605 (2023). https://doi.org/10.1007/s10439-023-03303-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10439-023-03303-0