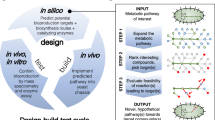
Abstract
Recent advances and emerging technologies for metabolic pathway engineering and synthetic biology have transformed the field of natural product discovery, production, and engineering. Despite these advancements, there remain many challenges in understanding how biosynthetic gene clusters are silenced or activated, including changes in the transcription of key biosynthetic and regulatory genes. This knowledge gap is highlighted by the success and failed attempts of manipulating regulatory genes within biosynthetic gene clusters in both native producers and heterologous hosts. These complexities make the choice of native producers versus heterologous hosts, fermentation medium, and supply of precursors crucial factors in achieving the production of the target natural products and engineering designer analogs. Nature continues to serve as inspiration for filling the knowledge gaps and developing new research strategies. By exploiting the evolutionary power of nature, alternative producers, with the desired genetic amenability and higher titers of the target natural products, and new strains, harboring gene clusters that encode evolutionary optimized congeners of the targeted natural product scaffolds, can be discovered. These newly identified strains can serve as an outstanding biotechnology platform for the engineered production of sufficient quantities of the target natural products and their analogs, enabling biosynthetic studies and potential therapeutic applications. These challenges and opportunities are showcased herein using fredericamycin, iso-migrastatin, platencin and platensimycin, the enediynes of C-1027, tiancimycin, and yangpumicin, and the leinamycin family of natural products.






Similar content being viewed by others
References
Anton N, Mendes MV, Martin JF, Aparicio JF (2004) Identification of PimR as a positive regulator of pimaricin biosynthesis in Streptomyces natalensis. J Bacteriol 186:2567–2575. https://doi.org/10.1128/JB.186.9.2567-2575.2004
Byrne KM, Hilton BD, White RJ, Misra R, Pandey RC (1985) Biosynthesis of fredericamycin A, a new antitumor antibiotic. Biochemistry 24:478–486. https://doi.org/10.1021/bi00323a035
Casini A, Chang F-Y, Eluere R, King AM, Young EM, Dudley QM, Karim A, Pratt K, Bristol C, Forget A, Ghodasara A, Warden-Rothman R, Gan R, Cristofaro A, Borujeni AE, Ryu M-H, Li J, Kwon Y-C, Wang H, Tatsis E, Rodriguez-Lopez C, O’Connor S, Medema MH, Fischbach MA, Jewett MC, Voigt C, Gordon DB (2018) A pressure test to make 10 molecules in 90 days: external evaluation of methods to engineer biology. J Am Chem Soc 140:4302–4316. https://doi.org/10.1021/jacs.7b13292
Chen Y, Wendt-Pienkowski E, Shen B (2008) Identification and utility of FdmR1 as a Streptomyces antibiotic regulatory protein activator for fredericamycin production in Streptomyces griseus ATCC 49344 and heterologous hosts. J Bacteriol 190:5587–5596. https://doi.org/10.1128/JB.00592-08
Chen Y, Yin M, Horsman GP, Huang S, Shen B (2010) Manipulation of pathway regulation in Streptomyces globisporus for overproduction of the enediyne antitumor antibiotic C-1027. J Antibiot 63:482–485. https://doi.org/10.1038/ja.2010.55
Chen Y, Yin M, Horsman GP, Shen B (2011) Improvement of the enediyne antitumor antibiotic C-1027 production by manipulating its biosynthetic pathway regulation in Streptomyces globisporus. J Nat Prod 74:420–424. https://doi.org/10.1021/np100825y
Cheng Y-Q, Tang G-L, Shen B (2002) Identification and localization of the gene cluster encoding biosynthesis of the antitumor macrolactam leinamycin in Streptomyces atroolivaceus S-140. J Bacteriol 184:7013–7024. https://doi.org/10.1128/JB.184.24.7013-7024.2002
Davies J, Wang H, Taylor T, Warabi K, Huang X-H, Andersen RJ (2005) Uncialamycin, a new enediyne antibiotic. Org Lett 7:5233–5236. https://doi.org/10.1021/ol052081f
Dong L-B, Rudolf JD, Shen B (2016) A mutasynthetic library of platensimycin and platencin analogues. Org Lett 18:4606–4609. https://doi.org/10.1021/acs.orglett.6b02248
Edo K, Mizugaki M, Koide Y, Seto H, Furihata K, Otake N, Ishida N (1985) The structure of neocarzinostatin chromophore possessing a novel bicylco-[7,3,0]dodecadiyne system. Tetrahedron Lett 26:331–334. https://doi.org/10.1016/S0040-4039(01)80810-8
Feng Z, Wang L, Rajski SR, Xu Z, Coeffet-LeGal MF, Shen B (2009) Engineered production of iso-migrastatin in heterologous Streptomyces hosts. Bioorg Med Chem 17:2147–2153. https://doi.org/10.1016/j.bmc.2008.10.074
Hara M, Asano K, Kawamoto I, Takiguchi T, Katsumata S, Takahashi K, Nakano H (1989) Leinamycin, a new antitumor antibiotic form Streptomyces: producing organism, fermentation, and isolation. J Antibiot 42:1768–1774. https://doi.org/10.7164/antibiotics.42.1768
Hara M, Saitoh Y, Nakano H (1990) DNA strand scission by the novel antitumor antibiotic leinamycin. Biochemistry 29:5676–5681. https://doi.org/10.1021/bi0047a005
Hara M, Takahashi I, Yoshida M, Asano K, Kawamoto I, Morimoto M, Nakano H (1989) DC 107, a novel antitumor antibiotic produced by a Streptomyces sp. J Antibiot 42:333–335. https://doi.org/10.7164/antibiotics.42.33
Herath KB, Zhang C, Jayasuriya H, Ondeyka JG, Zink DL, Burgess B, Wang J, Singh SB (2008) Structure and semisynthesis of platensimide A produced by Streptomyces platensis. Org Lett 10:1699–1702. https://doi.org/10.1021/ol800251v
Hindra, Huang T, Dong Y, Rudolf JD, Xie P, Xie G, Teng Q, Lohman JR, Zhu X, Huang Y, Zhao L-X, Jang Y, Duan Y, Shen B (2014) Strain prioritization for natural product discovery by a high-throughput real-time PCR method. J Nat Prod 77:2296–2303. https://doi.org/10.1021/np5006168
Hirayama N, Shimizu ME (1993) Molecular structure of a novel antitumor antibiotic leinamycin. Chem Lett 22:1957–1958. https://doi.org/10.1246/cl.1993.1957
Hochlowski JE, Whittern DN, Hill P, McAlpine JB (1994) Dorrigocins: novel antifungal antibiotics that change the morphology or ras-transformed NIH/3T3 cells to that of normal cells II isolation and elucidation of structures. J Antibiot 47:870–874. https://doi.org/10.7164/antibiotics.47.870
Ju J, Lim S-K, Jiang H, Seo J-W, Her Y, Shen B (2006) Thermolysis of iso-migrastatin and its congeners via [3]-sigmatropic rearrangement: a new route to the synthesis of migrastatin and its analogues. Org Lett 8:5868. https://doi.org/10.1021/ol062470p
Ju J, Lim S-K, Jiang H, Seo J-W, Shen B (2005) Iso-migrastatin congeners from Streptomyces platensis and generation of a glutarimide polyketide library featuring the dorrigocin, lactimidomycin, migrastatin, and nk30424 scaffolds. J Am Chem Soc 127:11930–11931. https://doi.org/10.1021/ja053118u
Ju J, Lim S-K, Jiang H, Shen B (2005) Migrastatin and dorrigocins are shunt metabolites of iso-migrastatin. J Am Chem Soc 127:1623–1662. https://doi.org/10.1021/ja043808i
Karwowski JP, Jackson M, Sunga G, Sheldon P, Poddig JB, Kohl WL, Kadan S (1994) Dorrigocins: novel antifungal antibiotics that change the morphology or ras-transformed NIH/3T3 cells to that of normal cells I taxonomy of the producing organism, fermentation and biological activity. J Antibiot 47:862–869. https://doi.org/10.7164/antibiotics.47.862
Konishi M, Ohkuma H, Matsumoto K, Tsuno T, Kamei H, Miyaki T, Oki T, Kawaguchi H, VanDuyne GD, Clardy J (1989) Dynemicin A, a novel antibiotic with the anthraquinone and 1,5-diyn-3-ene subunit. J Antibiot 42:1449–1452. https://doi.org/10.7164/antibiotics.42.1449
Liang Z-X (2010) Complexity and simplicity in the biosynthesis of enediyne natural products. Nat Prod Rep 27:499–528. https://doi.org/10.1039/B908165H
Lim S-K, Ju J, Zazopoulos E, Jiang H, Seo J-W, Chen Y, Feng Z, Rajski SR, Farnet CM, Shen B (2009) iso-Migrastatin, migrastatin, and dorrigocin production in Streptomyces platensis NRRL18993 is governed by a single biosynthetic machinery featuring an acyltransferase-less type I polyketide synthase. J Biol Chem 284:29746–29756. https://doi.org/10.1074/jbc.M109.046805
Ma M, Lohman JR, Liu T, Shen B (2015) C-S bon cleavage by a polyketide synthase domain. Proc Natl Acad Sci USA 112:10359–10364. https://doi.org/10.1073/pnas.1508437112
Medema MH, Trefzer A, Kovalchuk A, Berg MVD, Muller U, Heijne W, Wu L, Alam MT, Ronning CM, Nierman WC, Bovenberg RA, Breitling R, Takano R (2010) The sequence of a 1.8-Mb bacterial linear plasmid reveals a rich evolutionary reservoir of secondary metabolic pathways. Genome Biol Evol 2:212–224. https://doi.org/10.1093/gbe/evq013/570411
Misra R, Pandey RC, Silverton JV (1982) Fredericamycin A, an antitumor antibiotic of novel skeletal type. J Am Chem Soc 104:4478–4479. https://doi.org/10.1021/ja00380a025
Nakae K, Yoshimoto Y, Sawa T, Homma Y, Hamada M, Takeuchi T, Imoto M (2000) Migrastatin, a new inhibitor of tumor cell migration from Streptomyces s. MK929-43F1. Taxonomy, fermentation, isolation and biological activites. J Antibiot 53:1130–1136. https://doi.org/10.7164/antibiotics.53.1130
Newman DJ, Cragg GM (2016) Natural products as sources of new drugs from 1981 to 2014. J Nat Prod 79:629–661. https://doi.org/10.1021/acs.jnatprod.5b01055
Pan G, Xu Z, Guo Z, Hindra, Ma M, Yang D, Zhou H, Gansemans Y, Zhu X, Huang Y, Zhao L-X, Jiang Y, Cheng J, Van Nieuwerburgh F, Suh J-W, Duan Y, Shen B (2017) Discovery of the leinamycin family of natural products by mining actinobacterial genomes. Proc Natl Acad Sci USA 114:E11131–E11140. https://doi.org/10.1073/pnas.1716245115
Pandey RC, Toussaint MW, Stroshane RM, Kalita CC, Aszalos AA, Garretson AL, Wei TT, Byrne KM, Geoghegan RF Jr (1981) Fredericamycin A, a new antitumor antibiotic I. production, isolation and physicochemical properties. J Antibiot 34:1389–1401. https://doi.org/10.7164/antibiotics.34.1389
Romero-Rodriguez A, Robledo-Casados I, Sanchez S (2015) An overview on transcriptional regulators in Streptomyces. Biochimica et Biophysica Acta 1849:1017–1039. https://doi.org/10.1016/j.bbagrm.2015.06.007
Rudolf JD, Dong L-B, Huang T, Shen B (2015) A genetically amenable platensimycin- and platencin-overproducer as a platform for biosynthetic explorations: a showcase of PtmO4. A long-chain acyl-CoA dehydrogenase. Mol Biosyst 11:2717–2726. https://doi.org/10.1039/c5mb00303b
Rudolf JD, Yan X, Shen B (2016) Genome neighborhood network reveals insights into enediyne biosynthesis and facilitates prediction and prioritization for discovery. J Ind Microbiol Biotechnol 43:261–276. https://doi.org/10.1007/s10295-015-1671-0
Shen B (2015) A new golden age of natural products drug discovery. Cell 163:1297–1300. https://doi.org/10.1016/j.cell.2015.11.031
Shi J, Pan J, Yang D, Lu S, Zhu X, Shen B, Duan Y, Huang Y (2016) Titer improvement and pilot-scale production of platensimycin from Streptomyces platensis SB12026. J Ind Microbiol Biotechnol 43:1027–1035. https://doi.org/10.1007/s10295-016-1769-z
Smanski MJ, Casper J, Peterson RM, Yu Z, Rajski SR, Shen B (2012) Expression of the platencin biosynthetic gene cluster in heterologous hosts yielding new platencin congeners. J Nat Prod 75:2158–2167. https://doi.org/10.1021/np3005985
Smanski MJ, Peterson RM, Rajski SR, Shen B (2009) Engineered Streptomyces platensis strains that overproduce antibiotics platensimycin and platencin. Antimicrob Agents Chemother 53:1299–1304. https://doi.org/10.1128/AAC.01358-08
Smanski MJ, Yu Z, Casper J, Lin S, Peterson RM, Chen Y, Wendt-Pienkowski E, Rajski SR, Shen B (2011) Dedicated ent-kaurene and ent-atiserene synthases for platensimycin and platencin biosynthesis. Proc Natl Acad Sci USA 108:13498–13503. https://doi.org/10.1073/pnas.1106919108
Smanski MJ, Zhou H, Claesen J, Shen B, Fishbach MA, Voigt CA (2016) Synthetic biology to access and expand nature’s chemical diversity. Nat Rev Micobiol 14:135–149. https://doi.org/10.1038/nrmicro.2015.24
Stontag B, Muller JG, Hansske FG (2004) Fredericamycin B and C from Streptomyces griseus: structure elucidation after 23 years. J Antibiot 57:823–828. https://doi.org/10.7164/antibiotics.57.823
Tang G-L, Cheng Y-Q, Shen B (2004) Leinamycin biosynthesis revealing unprecedented architectural complexity for a hybrid polyketide synthase and nonribosomal peptide synthetase. Chem Biol 11:33–45. https://doi.org/10.1016/j.chembiol.2003.12.014
Van Lanen SG, Shen B (2008) Biosynthesis of enediyne antitumor antibiotics. Curr Top Med Chem 8:448–459. https://doi.org/10.2174/156802608783955656
Wang J, Kodali S, Lee SH, Galgoci A, Painter R, Dorso K, Racine F, Motyl M, Hernandez L, Tinney E, Colletti SL, Herath K, Cummings R, Salazar O, Gonzalez I, Basillo A, Vicente F, Genilloud O, Pelaez F, Jayasuriya H, Young K, Cully DF, Singh SB (2007) Discovery of platencin, a dual FabF and FabH inhibitor with in vivo antibiotic properties. Proc Natl Acad Sci USA 104:7612–7616. https://doi.org/10.1073/pnas.0700746104
Wang J, Soisson SM, Young K, Shoop W, Kodali S, Galgoci A, Painter R, Parthasarathy G, Tang YS, Cummings R, Ha S, Dorso K, Motyl M, Jayasuriya H, Ondeyka J, Herath K, Zhang C, Hernandez L, Allocco J, Basilio A, Tormo JR, Genilloud O, Vicente F, Pelaez F, Colwell L, Lee SH, Michael B, Felcetto T, Gill C, Silver LL, Hermes JD, Bartizal K, Barrett J, Schmatz D, Becker JW, Cully D, Singh SB (2006) Platensimycin is a selective FabF inhibitor with potent antibiotic properties. Nature 441:358–361. https://doi.org/10.1038/nature04784
Wang L, Wang S, He Q, Yu T, Li Q, Hong B (2012) Draft genome sequence of Streptomyces globisporus C-1027, which produces an antitumor antibiotic consisting of a nine-membered enediyne with a chromoprotein. J Bacteriol 194:4144. https://doi.org/10.1128/jb.00797-12
Wendt-Pienkowski E, Huang Y, Zhang J, Li B, Jiang H, Kwon H, Hutchinson R, Shen B (2005) Cloning, sequencing, analysis, and heterologous expression of the fredericamycin biosynthetic gene cluster from Streptomyces griseus. J Am Chem Soc 127:16442–16452. https://doi.org/10.1021/ja054376u
Wenzel SC, Müller R (2005) Recent developments towards the heterologous expression of complex bacterial natural product biosynthetic pathways. Curr Opin Biotechnol 16:594–606. https://doi.org/10.1016/j.copbio.2005.10.001
Woo EJ, Starks CM, Carney JR, Arslanin R, Cadapan L, Zavala S, Licari P (2002) Migrastatin and a new compound, isomigrastatin, from Streptomyces platensis. J Antibiot 55:141–146. https://doi.org/10.7164/antibiotics.55.141
Wu X, Yang D, Zhu X, Feng Z, Lv Z, Zhang Y, Shen B, Xu Z (2010) Iso-migrastatin titer improvement in the engineered Streptomyces lividans SB11002 strain by optimization of fermentation conditions. Biotechnol Bioprocess Eng 15:664–669. https://doi.org/10.1007/s12257-009-3129-6
Yan X, Chen J-J, Adhikari A, Yang D, Crnovic I, Wang N, Chang C-Y, Rader C, Shen B (2017) Genome mining of Micromonospora yangpuensis DSM 45577 as a producer of an anthraquinone-fused enediyne. Org Lett 19:6192–6196. https://doi.org/10.1021/acs.orglett.7b03120
Yan X, Ge H, Huang T, Hindra, Yang D, Teng Q, Crnovcic I, Li X, Rudolf JD, Lohman JR, Gansemans Y, Zhu X, Huang Y, Zhao L-X, Jiang Y, Van Nieuwerburgh F, Rader C, Duan Y, Shen B (2016) Strain prioritization and genome mining for enediyne natural products. mBio 7:e02104–e02116. https://doi.org/10.1128/mbio.02104-16
Yan X, Hindra, Ge H, Yang D, Huang T, Crnovcic I, Chang C-Y, Fang S-M, Annaval T, Zhu X, Huang Y, Zhao L-X, Jiang Y, Duan Y, Shen B (2018) Discovery of alternative producers of the enediyne antitumor antibiotic C-1027 with high titers. J Nat Prod 81:594–599. https://doi.org/10.1021/acs.jnatprod.7b01013
Yang D, Zhu X, Wu X, Fend Z, Huang L, Shen B, Xu Z (2011) Titer improvement of iso-migrastatin in selected heterologous Streptomyces hosts and related analysis of mRNA expression by quantitative RT-PCR. Appl Microbial Biotechnol 89:1709–1719. https://doi.org/10.1007/s00253-010-3025-1
Yu Z, Smanski MJ, Peterson RM, Marchillo K, Andes D, Rajski SR, Shen B (2011) Engineering of Streptomyces platensis MA7339 for overproduction of platencin and congeners. Org Lett 12:1744–1747. https://doi.org/10.1021/ol100342m
Zhang B, Yang D, Yan Y, Pan G, Xiang W, Shen B (2016) Overproduction of lactimidomycin by cross-overexpression of genes encoding Streptomyces antibiotic regulatory proteins. Appl Microbiol Biotechnol 100:2267–2277. https://doi.org/10.1007/s00253-015-7119-7
Zhang H, Wang Y, Pfeifer BA (2008) Bacterial hosts for natural product production. Mol Pharm 5:212–225. https://doi.org/10.1021/mp70013
Acknowledgements
Research on natural product discovery, biosynthesis, engineering and drug discovery in the Shen lab is currently supported by NIH Grants CA106150, GM114353, GM115575, and the Natural Products Library Initiative at The Scripps Research institute. We thank the past and current members of the Shen Lab for their dedication, creativity, and contributions to research on natural product discovery, biosynthesis, engineering, and genome mining, and Guohui Pan, Jeffery Rudolf, and Andrew Steele for their critical reading and valued inputs to this review. This is manuscript no. 29761 from The Scripps Research Institute.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the Special Issue “Natural Product Discovery and Development in the Genomic Era 2019”.
Rights and permissions
About this article
Cite this article
Teijaro, C.N., Adhikari, A. & Shen, B. Challenges and opportunities for natural product discovery, production, and engineering in native producers versus heterologous hosts. J Ind Microbiol Biotechnol 46, 433–444 (2019). https://doi.org/10.1007/s10295-018-2094-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-018-2094-5