Abstract
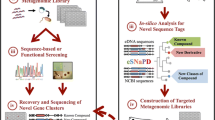
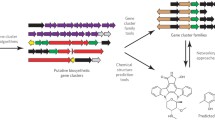
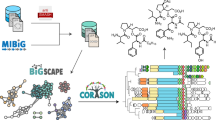
Technological improvements have accelerated natural product (NP) discovery and engineering to the point that systematic genome mining for new molecules is on the horizon. NP biosynthetic potential is not equally distributed across organisms, environments, or microbial life histories, but instead is enriched in a number of prolific clades. Also, NPs are not equally abundant in nature; some are quite common and others markedly rare. Armed with this knowledge, random ‘fishing expeditions’ for new NPs are increasingly harder to justify. Understanding the ecological and evolutionary pressures that drive the non-uniform distribution of NP biosynthesis provides a rational framework for the targeted isolation of strains enriched in new NP potential. Additionally, ecological theory leads to testable hypotheses regarding the roles of NPs in shaping ecosystems. Here we review several recent strain prioritization practices and discuss the ecological and evolutionary underpinnings for each. Finally, we offer perspectives on leveraging microbial ecology and evolutionary biology for future NP discovery.





Similar content being viewed by others
Reference
Abdelmohsen UR, Bayer K, Hentschel U (2014) Diversity, abundance and natural products of marine sponge-associated actinomycetes. Nat Prod Rep 31:381–399. doi:10.1039/c3np70111e
Andrianasolo EH, Haramaty L, Rosario-Passapera R, Vetriani C, Falkowski P, White E, Lutz R (2012) Ammonificins C and D, hydroxyethylamine chromene derivatives from a cultured marine hydrothermal vent bacterium, Thermovibrio ammonificans. Mar Drugs 10:2300–2311. doi:10.3390/md10102300
Bakker MG, Otto-Hanson L, Lange AJ, Bradeen JM, Kinkel LL (2013) Plant monocultures produce more antagonistic soil Streptomyces communities than high-diversity plant communities. Soil Biol Biochem 65:304–312. doi:10.1016/j.soilbio.2013.06.007
Baltz RH (2006) Marcel Faber roundtable: is our antibiotic pipeline unproductive because of starvation, constipation or lack of inspiration? J Ind Microbiol Biotechnol 33:507–513
Barton LL, Mandl M, Loy A (2010) Geomicrobiology: molecular and environmental perspective. Geomicrobiol Mol Environ Perspect. doi:10.1007/978-90-481-9204-5
Bernier SP, Surette MG (2013) Concentration-dependent activity in natural environments. Front Microbiol 4:20. doi:10.3389/fmicb.2013.00020
Bertrand S, Bohni N, Schnee S, Schumpp O, Gindro K, Wolfender J-L (2014) Metabolite induction via microorganism co-culture: a potential way to enhance chemical diversity for drug discovery. Biotechnol Adv 32:1180–1204. doi:10.1016/j.biotechadv.2014.03.001
Bhullar K, Waglechner N, Pawlowski A, Koteva K, Banks ED, Johnston MD, Barton HA, Wright GD (2012) Antibiotic resistance is prevalent in an isolated cave microbiome. PLoS One 7:e34953. doi:10.1371/journal.pone.0034953
Blunt JW, Copp BR, Keyzers Ra, Munro MHG, Prinsep MR (2014) Marine natural products. Nat Prod Rep 31:160–258. doi:10.1039/c3np70117d
Brendel N, Partida-Martinez LP, Scherlach K, Hertweck C (2007) A cryptic PKS-NRPS gene locus in the plant commensal Pseudomonas fluorescens Pf-5 codes for the biosynthesis of an antimitotic rhizoxin complex. Org Biomol Chem 5:2211–2213. doi:10.1039/b707762a
Bull AT (2004) ASM Press, Washington
Cafaro MJ, Currie CR (2005) Phylogenetic analysis of mutualistic filamentous bacteria associated with fungus-growing ants. Can J Microbiol 51:441–446. doi:10.1139/w05-023
Caldera EJ, Currie CR (2012) The population structure of antibiotic-producing bacterial symbionts of Apterostigma dentigerum ants: Impacts of coevolution and multipartite symbiosis. Am Nat 180:604–617. doi:10.1086/667886
De Candolle AP (1804) Essai sur les proprietes medicales de plantes, comparees ave leur formes exterieures et leur classification naturelle. Mequignon, Paris 1
Challis GL, Hopwood DA (2003) Synergy and contingency as driving forces for the evolution of multiple secondary metabolite production by Streptomyces species. Proc Natl Acad Sci 100:14555–14561. doi:10.1073/pnas.1934677100
Charlop-Powers Z, Owen JG, Reddy BVB, Ternei MA, Brady SF (2014) Chemical-biogeographic survey of secondary metabolism in soil. Proc Natl Acad Sci USA 111:3757–3762. doi:10.1073/pnas.1318021111
Charlop-Powers Z, Owen JG, Reddy BVB, Ternei MA, Guimarães DO, de Frias Ua, Pupo MT, Seepe P, Feng Z, Brady SF (2015) Global biogeographic sampling of bacterial secondary metabolism. Elife 4:1–10. doi:10.7554/eLife.05048
Cheng K, Rong X, Pinto-Tomás AA, Fernández-Villalobos M, Murillo-Cruz C, Huang Y (2015) Population genetic analysis of Streptomyces albidoflavus reveals habitat barriers to homologous recombination in the diversification of streptomycetes. Appl Environ Microbiol 81:966–975. doi:10.1128/AEM.02925-14
Cimermancic P, Medema MH, Claesen J, Kurita K, Wieland Brown LC, Mavrommatis K, Pati A, Godfrey PA, Koehrsen M, Clardy J, Birren BW, Takano E, Sali A, Linington RG, Fischbach MA (2014) Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158:412–421. doi:10.1016/j.cell.2014.06.034
Cordero OX, Polz MF (2014) Explaining microbial genomic diversity in light of evolutionary ecology. Nat Rev Microbiol 12:263–273. doi:10.1038/nrmicro3218
Cordero OX, Wildschutte H, Kirkup B, Proehl S, Ngo L, Hussain F, Le Roux F, Mincer T, Polz MF (2012) Ecological populations of bacteria act as socially cohesive units of antibiotic production and resistance. Science 337:1228–1231. doi:10.1126/science.1219385
Czaran TL (2002) Chemical warfare between microbes promotes biodiversity. Proc Natl Acad Sci 99:786–790. doi:10.1073/pnas.012399899
Davelos AL, Kinkel LL, Samac DA (2004) Spatial variation in frequency and intensity of antibiotic interactions among streptomycetes from prairie soil. Appl Environ Microbiol 70:1051–1058. doi:10.1128/AEM.70.2.1051-1058.2004
Davelos Baines AL, Xiao K, Kinkel LL (2007) Lack of correspondence between genetic and phenotypic groups amongst soil-borne streptomycetes. FEMS Microbiol Ecol 59:564–575. doi:10.1111/j.1574-6941.2006.00231.x
Davidson B (1995) New dimensions in natural products research: cultured marine microorganisms. Curr Opin Biotechnol 6:284–291. doi:10.1016/0958-1669(95)80049-2
Ding ZG, Li MG, Zhao JY, Ren J, Huang R, Xie MJ, Cui XL, Zhu HJ, Wen ML (2010) Naphthospironone A: an unprecedented and highly functionalized polycyclic metabolite from an alkaline mine waste extremophile. Chem-A Eur J 16:3902–3905. doi:10.1002/chem.200903198
Doroghazi JR, Albright JC, Goering AW, Ju K-S, Haines RR, Tchalukov KA, Labeda DP, Kelleher NL, Metcalf WW (2014) A roadmap for natural product discovery based on large-scale genomics and metabolomics. Nat Chem Biol 10:963–968. doi:10.1038/nchembio.1659
Doroghazi JR, Buckley DH (2010) Widespread homologous recombination within and between Streptomyces species. ISME J 4(1136–1143):29
Doroghazi JR, Buckley DH (2014) Intraspecies comparison of Streptomyces pratensis genomes reveals high levels of recombination and gene conservation between strains of disparate geographic origin. BMC Genom 15:970. doi:10.1186/1471-2164-15-970->
Fenical W (1993) Chemical studies of marine bacteria: developing a new resource. Chem Rev 93:1673–1683. doi:10.1021/cr00021a001
Firn RD, Jones CG (2000) The evolution of secondary metabolism—a unifying model. Mol Microbiol 37:989–994
Gabriel CR, Northup DE (2013) Cave microbiomes: a novel resource for drug discovery. Cave Microbiomes. doi:10.1007/978-1-4614-5206-5
Galm U, Shen B (2007) Natural product drug discovery: the times have never been better. Chem Biol 14:1098–1104
Genilloud O, González I, Salazar O, Martín J, Tormo JR, Vicente F (2011) Current approaches to exploit actinomycetes as a source of novel natural products. J Ind Microbiol Biotechnol 38:375–389. doi:10.1007/s10295-010-0882-7
Groll M, Huber R, Potts BCM (2006) Crystal structures of salinosporamide A (NPI-0052) and B (NPI-0047) in complex with the 20S proteasome reveal important consequences of β-lactone ring opening and a mechanism for irreversible binding. J Am Chem Soc 128:5136–5141. doi:10.1021/ja058320b
Haas D, Défago G (2005) Biological control of soil-borne pathogens by fluorescent pseudomonads. Nat Rev Microbiol 3:307–319. doi:10.1038/nrmicro1129
Heinig U, Scholz S, Jennewein S (2013) Getting to the bottom of Taxol biosynthesis by fungi. Fungal Divers 60:161–170. doi:10.1007/s13225-013-0228-7
Hentschel U, Piel J, Degnan SM, Taylor MW (2012) Genomic insights into the marine sponge microbiome. Nat Rev Microbiol 10:641–654. doi:10.1038/nrmicro2839
Hibbing ME, Fuqua C, Parsek MR, Peterson SB (2009) Bacterial competition: surviving and thriving in the microbial jungle. Nat Rev Microbiol 8:15–25. doi:10.1038/nrmicro2259
Huang T, Yang D, Xie P, Xie G, Teng Q, Lohman JR, Zhu X, Huang Y, Zhao L, Jiang Y, Duan Y, Shen B (2014) Strain prioritization for natural product discovery by a high- throughput real-time PCR method. J Nat Prod 77:2296–2303
Janvier C, Villeneuve F, Alabouvette C, Edel-Hermann V, Mateille T, Steinberg C (2007) Soil health through soil disease suppression: which strategy from descriptors to indicators? Soil Biol Biochem 39:1–23. doi:10.1016/j.soilbio.2006.07.001
Jensen PR, Williams PG, Oh DC, Zeigler L, Fenical W (2007) Species-specific secondary metabolite production in marine actinomycetes of the genus Salinispora. Appl Environ Microbiol 73:1146–1152. doi:10.1128/AEM.01891-06
Jiang W, Ye P, Chen CTA, Wang K, Liu P, He S, Wu X, Gan L, Ye Y, Wu B (2013) Two novel hepatocellular carcinoma cycle inhibitory cyclodepsipeptides from a hydrothermal vent crab-associated fungus Aspergillus clavatus C2WU. Mar Drugs 11:4761–4772. doi:10.3390/md11124761
Kaltenpoth M, Engl T (2014) Defensive microbial symbionts in Hymenoptera. Funct Ecol 28:315–327. doi:10.1111/1365-2435.12089
Kim BM, Lee JY, Hwang BK (1998) Diversity of actinomycetes antogonistic to plant pathogenic fungi in cave and sea-mud soils of Korea. J Microbiol 36:86–92
Kinkel LL, Bakker MG, Schlatter DC (2011) A coevolutionary framework for managing disease-suppressive soils. Annu Rev Phytopathol 49:47–67. doi:10.1146/annurev-phyto-072910-095232
Kinkel LL, Schlatter DC, Bakker MG, Arenz BE (2012) Streptomyces competition and co-evolution in relation to plant disease suppression. Res Microbiol 163:490–499. doi:10.1016/j.resmic.2012.07.005
Kinkel LL, Schlatter DC, Xiao K, Baines AD (2014) Sympatric inhibition and niche differentiation suggest alternative coevolutionary trajectories among Streptomycetes. ISME J 8:249–256. doi:10.1038/ismej.2013.175
Kirst HA, Michel KH, Martin JW, Creemer LC, Chio EH, Yao RC, Nakatsukasa WM, Boeck LD, Occolowitz JL, Paschal JW, Deeter JB, Jones ND, Thompson GD (1991) A83543A-D, unique fermentation-derived tetracyclic macrolides. Tetrahedron Lett 32:4839–4842. doi:10.1016/S0040-4039(00)93474-9
Larsen PE, Field D, Gilbert JA (2012) Predicting bacterial community assemblages using an artificial neural network approach. Nat Methods 9:621–625. doi:10.1038/nmeth.1975
Larsen TO, Smedsgaard J, Nielsen KF, Hansen ME, Frisvad JC (2005) Phenotypic taxonomy and metabolite profiling in microbial drug discovery. Nat Prod Report 22:672–695
Ligon JM, Hill DS, Hammer PE, Torkewitz NR, Hofmann D, Kempf H, Van Pe K (2000) Natural products with antifungal activity from Pseudomonas biocontrol bacteria. Pest Manag Sci 56:688–695
Little AEF, Murakami T, Mueller UG, Currie CR (2006) Defending against parasites: fungus-growing ants combine specialized behaviours and microbial symbionts to protect their fungus gardens. Biol Lett 2:12–16. doi:10.1098/rsbl.2005.0371
Liu M, Abdel-Mageed WM, Ren B, He W, Huang P, Li X, Bolla K, Guo H, Chen C, Song F, Dai H, Quinn RJ, Grkovic T, Liu X, Zhang X, Zhang L (2014) Endophytic Streptomyces sp Y3111 from traditional Chinese medicine produced antitubercular pluramycins. Appl Microbiol Biotechnol 98:1077–1085. doi:10.1007/s00253-013-5335-6
Long RA, Azam F (2001) Antagonistic interactions among marine pelagic bacteria. Appl Environ Microbiol 67:4975–4983. doi:10.1128/AEM.67.11.4975-4983.2001
Long RA, Rowley DC, Zamora E, Liu J, Bartlett DH, Azam F (2005) Antagonistic interactions among marine bacteria impede the proliferation of Vibrio cholerae. Appl Environ Microbiol 71:8531–8536. doi:10.1128/AEM.71.12.8531-8536.2005
Lu C, Shen Y (2007) A novel ansamycin, naphthomycin K from Streptomyces sp. J Antibiot (Tokyo) 60:649–653
Macherla VR, Mitchell SS, Manam RR, Reed KA, Chao TH, Nicholson B, Deyanat-Yazdi G, Mai B, Jensen PR, Fenical WF, Neuteboom STC, Lam KS, Palladino MA, Potts BCM (2005) Structure-activity relationship studies of salinosporamide A (NPI-0052), a novel marine derived proteasome inhibitor. J Med Chem 48:3684–3687. doi:10.1021/jm048995+
Marmann A, Aly A, Lin W, Wang B, Proksch P (2014) Co-cultivation—a powerful emerging tool for enhancing the chemical diversity of microorganisms. Mar Drugs 12:1043–1065. doi:10.3390/md12021043
Martiny JBH, Bohannan BJM, Brown JH, Colwell RK, Fuhrman Ja, Green JL, Horner-Devine MC, Kane M, Krumins JA, Kuske CR, Morin PJ, Naeem S, Ovreås L, Reysenbach A-L, Smith VH, Staley JT (2006) Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4:102–112. doi:10.1038/nrmicro1341
Martiny JBH, Eisen JA, Penn K, Allison SD, Horner-Devine MC (2011) Drivers of bacterial beta-diversity depend on spatial scale. Proc Natl Acad Sci USA 108:7850–7854. doi:10.1073/pnas.1016308108
Mazzola M (2004) Assessment and management of soil microbial community structure for disease suppression. Annu Rev Phytopathol 42:35–59. doi:10.1146/annurev.phyto.42.040803.140408
Mendes R, Kruijt M, de Bruijn I, Dekkers E, van der Voort M, Schneider JHM, Piceno YM, DeSantis TZ, Andersen GL, Bakker PAHM, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100. doi:10.1126/science.1203980
Michelsen CF, Watrous J, Glaring MA, Kersten R, Koyama N, Dorrestein PC (2015) Non-ribosomal peptides, key biocontrol components for Pseudomonas fluorescens In5, isolated from a Greenlandic suppressive soil. Mbio 6:2e0079–2e0115
Mitova MI, Lang G, Wiese J, Imhoff JF (2008) Subinhibitory concentrations of antibiotics induce phenazine production in a marine Streptomyces sp. J Nat Prod 71:824–827. doi:10.1021/np800032a
Moore BS (2005) Biosynthesis of marine natural products: microorganisms (Part A). Nat Prod Rep 22:580–593. doi:10.1039/b404737k
Morlon H, O’Connor TK, Bryant JA, Charkoudian LK, Docherty KM, Jones E, Kembel SW, Green JL, Bohannan BJM (2015) The biogeography of putative microbial antibiotic production. PLoS One 10:e0130659. doi:10.1371/journal.pone.0130659
Mousa WK, Raizada MN (2015) Biodiversity of genes encoding anti-microbial traits within plant associated microbes. Front Plant Sci. doi:10.3389/fpls.2015.00231
Newman DJ, Cragg GM (2012) Natural products as sources of new drugs over the 30 years from 1981 to 2010. J Nat Prod 75:311–335. doi:10.1021/np200906s
Nguyen T, Ishida K, Jenke-Kodama H, Dittmann E, Gurgui C, Hochmuth T, Taudien S, Platzer M, Hertweck C, Piel J (2008) Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection. Nat Biotechnol 26:225–233. doi:10.1038/nbt1379
Nielsen TH, Christophersen C, Anthoni U, Sørensen J (1999) Viscosinamide, a new cyclic depsipeptide with surfactant and antifungal properties produced by Pseudomonas fluorescens DR54. J Appl Microbiol 87:80–90. doi:10.1046/j.1365-2672.1999.00798.x
Oh D-C, Poulsen M, Currie CR, Clardy J (2009) Dentigerumycin: a bacterial mediator of an ant-fungus symbiosis. Nat Chem Biol 5:391–393. doi:10.1038/nchembio.159
Oh DC, Scott JJ, Currie CR, Clardy J (2009) Mycangimycin, a polyene peroxide from a mutualist Streptomyces sp. Org Lett 11:633–636. doi:10.1021/ol802709x
Ohta E, Ohta S, Kubota NK, Suzuki M, Ogawa T, Yamasaki A, Ikegami S (2001) Micromonospolide A, a new macrolide from Micromonospora sp. Tetrahedron Lett 42:4179–4181. doi:10.1016/S0040-4039(01)00683-9
Passari AK, Mishra VK, Saikia R, Gupta VK, Singh BP (2015) Isolation, abundance and phylogenetic affiliation of endophytic actinomycetes associated with medicinal plants and screening for their in vitro antimicrobial biosynthetic potential. Front Microbiol. doi:10.3389/fmicb.2015.00273
Pettit G, Herald C, Doubek DL, Herald DL, Arnold E, Clardy J (1982) Isolation and structure of bryostatin 1. J Am Chem Soc 104:6846–6848. doi:10.1021/ja00388a092
Piel J (2002) A polyketide synthase-peptide synthetase gene cluster from an uncultured bacterial symbiont of Paederus beetles. Proc Natl Acad Sci USA 99:14002–14007. doi:10.1073/pnas.222481399
Poulsen M, Cafaro M, Boomsma JJ, Currie CR (2005) Specificity of the mutualistic association between actinomycete bacteria and two sympatric species of Acromyrmex leaf-cutting ants. Mol Ecol 14:3597–3604. doi:10.1111/j.1365-294X.2005.02695.x
Poulsen M, Oh D-C, Clardy J, Currie CR (2011) Chemical analyses of wasp-associated Streptomyces bacteria reveal a prolific potential for natural products discovery. PLoS One 6:e16763. doi:10.1371/journal.pone.0016763
Qin S, Xing K, Jiang JH, Xu LH, Li WJ (2011) Biodiversity, bioactive natural products and biotechnological potential of plant-associated endophytic actinobacteria. Appl Microbiol Biotechnol 89:457–473. doi:10.1007/s00253-010-2923-6
Reveillaud J, Maignien L, Eren Ma, Huber Ja, Apprill A, Sogin ML, Vanreusel A (2014) Host-specificity among abundant and rare taxa in the sponge microbiome. ISME J 8:1198–1209. doi:10.1038/ismej.2013.227
Rodriguez RJ, Henson J, Van Volkenburgh E, Hoy M, Wright L, Beckwith F, Kim Y-O, Redman RS (2008) Stress tolerance in plants via habitat-adapted symbiosis. ISME J 2:404–416. doi:10.1038/ismej.2007.106
Schlatter DC, Kinkel LL (2014) Global biogeography of Streptomyces antibiotic inhibition, resistance, and resource use. FEMS Microbiol Ecol 88:386–397. doi:10.1111/1574-6941.12307
Seipke RF (2015) Strain-level diversity of secondary metabolism in Streptomyces albus. PLoS One 10:e0116457. doi:10.1371/journal.pone.0116457
Seipke RF, Kaltenpoth M, Hutchings MI (2012) Streptomyces as symbionts: an emerging and widespread theme? FEMS Microbiol Rev 36:862–876. doi:10.1111/j.1574-6976.2011.00313.x
Shaaban KA, Singh S, Elshahawi SI, Wang X, Ponomareva LV, Sunkara M, Copley GC, Hower JC, Morris AJ, Kharel MK, Thorson JS (2014) Venturicidin C, a new 20-membered macrolide produced by Streptomyces sp. TS-2-2. J Antibiot (Tokyo) 67:223–230. doi:10.1038/ja.2013.113
Shapiro BJ, Friedman J, Cordero OX, Preheim SP, Timberlake SC, Szabo G, Polz MF, Alm EJ (2012) Population genomics of early events in the ecological differentiation of bacteria. Science 336:48–51. doi:10.1126/science.1218198
Shapiro BJ, Polz MF (2014) Ordering microbial diversity into ecologically and genetically cohesive units. Trends Microbiol 22:235–247. doi:10.1016/j.tim.2014.02.006
Sipkema D, de Caralt S, Ja Morillo, Al-Soud WA, Sørensen SJ, Smidt H, Uriz MJ (2015) Similar sponge-associated bacteria can be acquired via both vertical and horizontal transmission. Environ Microbiol. doi:10.1111/1462-2920.12827 in press
Skropeta D (2008) Deep-sea natural products. Nat Prod Rep 25:1131–1166. doi:10.1039/b808743a
Stierle A, Strobel G, Stierle D (1993) Taxol and taxane production by Taxomyces andreanae, an endophytic fungus of Pacific yew. Science 260:214–216. doi:10.1126/science.8097061
Stierle AA, Stierle DB, Kelly K (2006) Berkelic acid, a novel spiroketal with selective anticancer activity from an acid mine waste fungal extremophile. J Org Chem 71:5357–5360. doi:10.1021/jo060018d
Strobel G, Daisy B (2003) Bioprospecting for microbial endophytes and their natural products. Microbiol Mol Bio Rev 67:491–502. doi:10.1128/MMBR.67.4.491
Takeuchi K, Noda N, Katayose Y, Mukai Y, Numa H, Yamada K, Someya N (2015) Rhizoxin analogs contribute to the biocontrol activity of a newly isolated Pseudomonas strain. Mol Plant Microbe Interact 28:333–342
Taylor MW, Schupp PJ, De Nys R, Kjelleberg S, Steinberg PD (2005) Biogeography of bacteria associated with the marine sponge Cymbastela concentrica. Environ Microbiol 7:419–433. doi:10.1111/j.1462-2920.2004.00711.x
Thompson JN (2005) The geographic mosaic of coevolution. University of Chicago Press, Chicago
Tiwari K, Gupta RK (2012) Diversity and isolation of rare actinomycetes: an overview. Crit Rev Microbiol 39:1–39. doi:10.3109/1040841X.2012.709819
Tiwari K, Gupta RK (2012) Rare actinomycetes: a potential storehouse for novel antibiotics. Crit Rev Biotechnol 32:108–132. doi:10.3109/07388551.2011.562482
Torsvik V, Goksøyr J, Daae FL (1990) High diversity in DNA of soil bacteria. Appl Environ Microbiol 56:782–787
Traxler MF, Watrous JD, Alexandrov T, Dorrestein PC, Kolter R (2013) Interspecies interactions stimulate diversification of the Streptomyces coelicolor secreted metabolome. mBio 4:e00459–13. doi:10.1128/mBio.00459-13
Truyens S, Weyens N, Cuypers A, Vangronsveld J (2015) Bacterial seed endophytes: genera, vertical transmission and interaction with plants: bacterial seed endophytes. Environ Microbiol Rep 7:40–50. doi:10.1111/1758-2229.12181
Vaz Jauri P, Bakker MG, Salomon CE, Kinkel LL (2013) Subinhibitory antibiotic concentrations mediate nutrient use and competition among soil Streptomyces. PLoS One 8:e81064. doi:10.1371/journal.pone.0081064
Vaz Jauri P, Kinkel LL (2014) Nutrient overlap, genetic relatedness and spatial origin influence interaction-mediated shifts in inhibitory phenotype among Streptomyces spp. FEMS Microbiol Ecol 90:264–275. doi:10.1111/1574-6941.12389
Verma VC, Prakash S, Singh RG, Gange AC (2014) Host-mimetic metabolomics of endophytes: looking back into the future. Adv Endophytic Res. doi:10.1007/978-81-322-1575-2
Waksman S, Schatz A (1943) Strain specificity and production of antibiotic substances. Proc Natl Acad Sci USA 29:74–79
Wang J, Kodali S, Lee SH, Galgoci A, Painter R, Dorso K, Racine F, Motyl M, Hernandez L, Tinney E, Colletti SL, Herath K, Cummings R, Salazar O, González I, Basilio A, Vicente F, Genilloud O, Pelaez F, Jayasuriya H, Young K, Cully DF, Singh SB (2007) Discovery of platencin, a dual FabF and FabH inhibitor with in vivo antibiotic properties. Proc Natl Acad Sci USA 104:7612–7616. doi:10.1073/pnas.0700746104
Wang J, Soisson SM, Young K, Shoop W, Kodali S, Galgoci A, Painter R, Parthasarathy G, Tang YS, Cummings R, Ha S, Dorso K, Motyl M, Jayasuriya H, Ondeyka J, Herath K, Zhang C, Hernandez L, Allocco J, Basilio A, Tormo JR, Genilloud O, Vicente F, Pelaez F, Colwell L, Lee SH, Michael B, Felcetto T, Gill C, Silver LL, Hermes JD, Bartizal K, Barrett J, Schmatz D, Becker JW, Cully D, Singh SB (2006) Platensimycin is a selective FabF inhibitor with potent antibiotic properties. Nature 441:358–361. doi:10.1038/nature04784
Wang W, Ji J, Li X, Wang J, Li S, Pan G, Fan K, Yang K (2014) Angucyclines as signals modulate the behaviors of Streptomyces coelicolor. Proc Natl Acad Sci 111:5688–5693. doi:10.1073/pnas.1324253111
Wang X, Elshahawi SI, Shaaban KA, Fang L, Ponomareva LV, Zhang Y, Copley GC, Hower JC, Zhan CG, Kharel MK, Thorson JS (2014) Ruthmycin, a new tetracyclic polyketide from Streptomyces sp. RM-4-15. Org Lett 16:456–459. doi:10.1021/ol4033418
Wang X, Shaaban KA, Elshahawi SI, Ponomareva LV, Sunkara M, Copley GC, Hower JC, Morris AJ, Kharel MK, Thorson JS (2014) Mullinamides A and B, new cyclopeptides produced by the Ruth Mullins coal mine fire isolate Streptomyces sp. RM-27-46. J Antibiot (Tokyo) 67:571–575
Wani ZA, Ashraf N, Mohiuddin T, Riyaz-Ul-Hassan S (2015) Plant-endophyte symbiosis, an ecological perspective. Appl Microbiol Biotechnol 99:2955–2965. doi:10.1007/s00253-015-6487-3
Wawrik B, Kutliev D, Abdivasievna UA, Kukor JJ, Zylstra GJ, Kerkhof L (2007) Biogeography of actinomycete communities and type II polyketide synthase genes in soils collected in New Jersey and Central Asia. Appl Environ Microbiol 73:2982–2989. doi:10.1128/AEM.02611-06
Weinstein MJ, Luedemann GM, Oden EM, Wagman GH (1963) Gentamicin, a new broad-spectrum antibiotic complex. Antimicrob Agents Chemother 161:1–7
Weller DM, Raaijmakers JM, Gardener BBM, Thomashow LS (2002) Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annu Rev Phytopathol 40:309–348. doi:10.1146/annurev.phyto.40.030402.110010
Wiener P (1996) Experimental studies on the ecological role of antibiotic production in bacteria. Evol Ecol 10:405–421. doi:10.1007/BF01237726
Williams S, Vickers J (1986) The ecology of antibiotic production. Microb Ecol 12:43–52
Wilson MC, Mori T, Rückert C, Uria AR, Helf MJ, Takada K, Gernert C, Steffens UAE, Heycke N, Schmitt S, Rinke C, Helfrich EJN, Brachmann AO, Gurgui C, Wakimoto T, Kracht M, Crüsemann M, Hentschel U, Abe I, Matsunaga S, Kalinowski J, Takeyama H, Piel J (2014) An environmental bacterial taxon with a large and distinct metabolic repertoire. Nature 506:58–62. doi:10.1038/nature12959
Yim G, Wang HH, Davies J (2007) Antibiotics as signalling molecules. Philos Trans R Soc Lond B Biol Sci 362:1195–1200. doi:10.1098/rstb.2007.2044
Zein N, Solomon W, Colson KL, Schroeder DR (1995) Maduropeptin: an antitumor chromoprotein with selective protease activity and DNA cleaving properties. Biochemistry 34:11591–11597. doi:10.1021/bi00036a035
Zhao LX, Huang SX, Tang SK, Jiang CL, Duan Y, Beutler Ja, Henrich CJ, McMahon JB, Schmid T, Blees JS, Colburn NH, Rajski SR, Shen B (2011) Actinopolysporins A-C and tubercidin as a pdcd4 stabilizer from the halophilic actinomycete Actinopolyspora erythraea YIM 90600. J Nat Prod 74:1990–1995. doi:10.1021/np200603g
Zinger L, Boetius A, Ramette A (2014) Bacterial taxa-area and distance-decay relationships in marine environments. Mol Ecol 23:954–964. doi:10.1111/mec.12640
Acknowledgements
This work has been supported by the National Institute of Food and Agriculture, U.S. Department of Agriculture, under agreement No. 2011-67019-30330 and the University of Minnesota Agricultural Experiment Station Project (#MIN 22-018). Resources from the University of Minnesota Supercomputing Institute are gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Additional information
Special Issue: Natural Product Discovery and Development in the Genomic Era. Dedicated to Professor Satoshi Ōmura for his numerous contributions to the field of natural products.
Rights and permissions
About this article
Cite this article
Smanski, M.J., Schlatter, D.C. & Kinkel, L.L. Leveraging ecological theory to guide natural product discovery. J Ind Microbiol Biotechnol 43, 115–128 (2016). https://doi.org/10.1007/s10295-015-1683-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-015-1683-9