Abstract
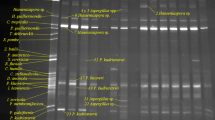
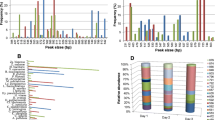
We compared pyrosequencing technology with the PCR-ITS-RFLP analysis of yeast isolates and denaturing gradient gel electrophoresis (DGGE). These methods gave divergent findings for the yeast population. DGGE was unsuitable for the quantification of biodiversity and its use for species detection was limited by the initial abundance of each species. The isolates identified by PCR-ITS-RFLP were not fully representative of the true population. For population dynamics, high-throughput sequencing technology yielded results differing in some respects from those obtained with other approaches. This study demonstrates that 454 pyrosequencing of amplicons is more relevant than other methods for studying the yeast community on grapes and during alcoholic fermentation. Indeed, this high-throughput sequencing method detected larger numbers of species on grapes and identified species present during alcoholic fermentation that were undetectable with the other techniques.




Similar content being viewed by others
References
Beltran G, Torija MJ, Novo M, Ferrer N, Poblet M, Guillamon JM, Rozes N, Mas A (2002) Analysis of yeast populations during alcoholic fermentation: a six-year follow-up study. Syst Appl Microbiol 25:287–293
Berry D, Mahfoudh KB, Wagner M, Loy A (2011) Barcoded primers used in multiplex amplicon pyrosequencing bias amplification. Appl Environ Microbiol 77:7846–7849
Bisson LF, Joseph CML (2009) Yeasts. In: König H (ed) Biology of microorganisms on grapes in must and in wine. Springer, Berlin Heidelberg New York, pp 47–60
Bokulich NA, Joseph CM, Allen G, Benson AK, Mills DA (2012) Next-generation sequencing reveals significant bacterial diversity of botrytized wine. PLoS ONE 7:e36357
Bokulich NA, Bamforth CW, Mills DA (2012) Brewhouse-resident microbiota are responsible for multi-stage fermentation of American coolship ale. PLoS ONE 7(4):e35507
Bokulich NA, Ohta M, Richardson PM, Mills DA (2013) Monitoring seasonal changes in winery-resident microbiota. PLoS ONE 6:e66437
Borman AM, Linton CJ, Miles SJ, Campbell CK, Johnson EM (2006) Ultra-rapid preparation of total genomic DNA from isolates of yeast and mould using Whatman FTA filter paper technology: a re-usable DNA archiving system. Med Mycol 44:389–398
Cavalieri D, Barberio C, Casalone E, Pinzauti F, Sebastiani F, Mortimer R, Polsinelli M (1998) Genetic and molecular diversity in Saccharomyces cerevisiae natural populations. Food Technol Biotechnol 36:45–50
Chemidlin Prévost-Bouré N, Christen R, Dequiedt S, Mougel C, Lelièvre M, Jolivet C, Shahbazkia HR, Guillou L, Arrouays D, Ranjard L (2011) Validation and application of a PCR primer set to quantify fungal communities in the soil environment by real-time quantitative PCR. PLoS ONE 6:e24166
Cocolin L, Bisson LF, Mills DA (2000) Direct profiling of the yeast dynamics in wine fermentations. FEMS Microbiol Lett 189:81–87
Comitini F, Ciani M (2008) Influence of fungicide treatments on the occurrence of yeast flora associated with wine grapes. Ann Microbiol 58:489–493
Cordero-Bueso G, Arroyo T, Serrano A, Tello J, Aporta I, Vélez MD, Valero E (2011) Influence of the farming system and vine variety on yeast communities associated with grape berries. Int J Food Microbiol 145:132–139
Dobson A, O’sullivan O, Cotter PD, Ross P, Hill C (2011) High-throughput sequence-based analysis of the bacterial composition of kefir and an associated kefir grain. FEMS Microbiol Lett 320:56–62
Esteve-Zarzoso B, Belloch C, Uruburu F, Querol A (1999) Identification of yeasts by RFLP analysis of the 5 8S rRNA gene and the two ribosomal internal transcribed spacers. Int J Syst Bacteriol 49:329–337
Fleet GH (2003) Yeast interactions and wine flavor. Int J Food Microbiol 86:11–22
Fowell RR (1965) The identification of wild yeast colonies on lysine agar. J Appl Bacteriol 28:373–383
Gil JV, Mateo JJ, Jimenez M, Pastor A, Huerta T (1996) Aroma compounds in wines as influenced by apiculate yeasts. J Food Sci 61:1247–1249
Heard G (1999) Novel yeasts in winemaking—looking to the future. Food Aust 51:347–352
Ivey ML, Phister TG (2011) Detection and identification of microorganisms in wine: a review of molecular techniques. J Ind Microbiol Biotechnol 38:1619–1634
Jolly N, Augustyn O, Pretorius I (2003) The occurrence of non-Saccharomyces yeast species over three vintages on four vineyards and grape musts from four production regions of the Western Cape South Africa. S Afr J Enol Vitic 24:35–42
Kunin V, Engelbrektson A, Ochman H, Hugenholtz P (2010) Wrinkles in the rare biosphere: pyrosequencing errors can lead to artificial inflation of diversity estimates. Environ Microbiol 12:118–123
Leite AMO, Mayoa B, Rachid CTCC, Peixoto RS, Silva JT, Paschoalin VMF, Delgado S (2012) Assessment of the microbial diversity of Brazilian kefir grains by PCR-DGGE and pyrosequencing analysis. Food Microbiol 31:215–221
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer ML, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu P, Begley RF, Rothberg JM (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–380
Martini A, Ciani M, Scorzetti G (1996) Direct enumeration and isolation of wine yeast from grape surfaces. Am J Enol Vitic 47:435–440
Milanović V, Comitini F, Ciani M (2013) Grape berry yeast communities: influence of fungicide treatments. Int J Food Microbiol 161:240–246
Mills DA, Johannsen EA, Cocolin L (2002) Yeast diversity and persistence in Botrytis affected wine fermentations. Appl Environ Microbiol 68:4884–4893
Nawrocki EP, Kolbe DL, Eddy SR (2009) Infernal 1.0: inference of RNA alignments. Bioinformatics 25:1713
Nisiotu AA, Spiropoulos AE, Nychas G-JE (2007) Yeast community structures and dynamics in healthy and Botrytis-affected grape must fermentations. Appl Environ Microbiol 21:6705–6713
Pascault N, Ranjard L, Kaisermann A, Bachar D, Christen R, Terrat S, Mathieu O, Lévêque J, Mougel C, Henault C, Lemanceau P, Péan M, Boiry S, Fontaine S, Maron PA (2013) Stimulation of different functional groups of bacteria by various plant residues as a driver of soil priming effect. Ecosystems. doi:10.1007/s10021-013-9650-7
Prakitchaiwattana CJ, Fleet GH, Heard GM (2004) Application and evaluation of denaturing gradient gel electrophoresis to analyse the yeast ecology of wine grapes. FEMS Yeast Res 4:865–877
Ribéreau-Gayon P, Peynaud E (1960) Traité d’Oenologie Paris et Liège. Librairie Polytechnique Ch Béranger Paris, pp 293–298
Regueiro LA, Costas CL, Lopez Rubio JE (1993) Influence of viticultural and enological practices on the development of yeast populations during winemaking. Am J Enol Vitic 44:405–408
Renouf V, Claisse O, Lonvaud-Funel A (2007) Inventory and monitoring of wine microbial consortia. J Appl Microbiol 75:149–164
Sadoudi M, Tourdot-Maréchal R, Rousseaux S, Steyer D, Gallardo-Chacón JJ, Ballester J, Vichi S, Guérin-Schneider R, Caixach J, Alexandre H (2012) Yeast-yeast interactions revealed by aromatic profile analysis of Sauvignon Blanc wine fermented by single or co-culture of non-Saccharomyces and Saccharomyces yeasts. Food Microbiol 32:243–253
Solieri L, Dakal TC, Giudici P (2013) Next-generation sequencing and its potential impact on food microbial genomics. Ann Microbiol 63:21–37
Setati ME, Jacobson D, Andong UC, Bauer F (2013) The vineyard yeast microbiome a mixed model microbial map. PLoS ONE. doi:10.1371/j.pone.0052609
Sipiczki M (2006) Metschnikowia strains isolated from botrytized grapes antagonize fungal and bacterial growth by iron depletion. Appl Environ Microbiol 72:6716–6724
Soden A, Francis IL, Oakey H, Henschke PA (2000) Effects of co-fermentation with Candida stellata and Saccharomyces cerevisiae on the aroma and composition of Chardonnay wine. Aust J Grape Wine Res 6:21–30
Terrat S, Christen R, Dequiedt S, Lelievre M, Nowak V, Regnier T, Bachar D, Plassart P, Wincker P, Jolivet C, Bispo A, Lemanceau P, Maron PA, Mougel C, Ranjard L (2012) Molecular biomass and MetaTaxogenomic assessment of soil microbial communities as influenced by soil DNA extraction procedure. Microb Biotechnol 5:135–141
White T, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfarnd DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Zott K, Thibon C, Bely M, Lonvaud-Funel A, Dubourdieu D, Masneuf-Pomarede I (2011) The grape must non-Saccharomyces microbial community: impact on volatile thiol release. Int J Food Microbiol 151:210–215
Acknowledgments
This work, through the involvement of the technical facilities of the GenoSol platform of the ANAEE-Services infrastructure, received a grant from the French state through the National Research Agency, as part of the “Investments for the Future” program (reference ANR-11-INBS-0001), and a grant from the Regional Council of Burgundy.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
David, V., Terrat, S., Herzine, K. et al. High-throughput sequencing of amplicons for monitoring yeast biodiversity in must and during alcoholic fermentation. J Ind Microbiol Biotechnol 41, 811–821 (2014). https://doi.org/10.1007/s10295-014-1427-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-014-1427-2