Abstract
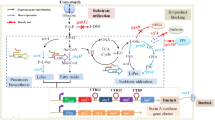
Sporolactobacillus inulinus, a homofermentative lactic acid bacterium, is a species capable of efficient industrial d-lactic acid production from glucose. Glucose phosphorylation is the key step of glucose metabolism, and fine-tuned expression of which can improve d-lactic acid production. During growth on high-concentration glucose, a fast induction of high glucokinase (GLK) activity was observed, and paralleled the patterns of glucose consumption and d-lactic acid accumulation, while phosphoenolpyruvate phosphotransferase system (PTS) activity was completely repressed. The transmembrane proton gradient of 1.3–1.5 units was expected to generate a large proton motive force to the uptake of glucose. This suggests that the GLK pathway is the major route for glucose utilization, with the uptake of glucose through PTS-independent transport systems and phosphorylation of glucose by GLK in S. inulinus d-lactic acid production. The gene encoding GLK was cloned from S. inulinus and expressed in Escherichia coli. The amino acid sequence revealed significant similarity to GLK sequences from Bacillaceae. The recombinant GLK was purified and shown to be a homodimer with a subunit molecular mass of 34.5 kDa. Strikingly, it demonstrated an unusual broad substrate specificity, catalyzing phosphorylation of 2-deoxyglucose, mannitol, maltose, galactose and glucosamine, in addition to glucose. This report documented the key step concerning glucose phosphorylation of S. inulinus, which will help to understand the regulation of glucose metabolism and d-lactic acid production.



Similar content being viewed by others
References
Albery WJ, Knowles JR (1976) Evolution of enzyme function and the development of catalytic efficiency. Biochemistry 15:5631–5640
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Buckley ND, Hamilton IR (1994) Vesicles prepared from Streptococcus mutans demonstrate the presence of a second glucose transport system. Microbiology 140:2639–2648
Conejo MS, Thompson SM, Miller BG (2010) Evolutionary bases of carbohydrate recognition and substrate discrimination in the ROK protein family. J Mol Evol 70:545–556
Dorr C, Zaparty M, Tjaden B, Brinkmann H, Siebers B (2003) The hexokinase of the hyperthermophile Thermoproteus tenax—ATP-dependent hexokinases and ADP-dependent glucokinases, two alternatives for glucose phosphorylation in Archaea. J Biol Chem 278:18744–18753
Goward CR, Hartwell R, Atkinson T, Scawen MD (1986) The purification and characterization of glucokinase from the thermophile Bacillus stearothermophilus. Biochem J 237:415–420
Han B, Liu HZ, Hu XM, Cai YJ, Zheng DS, Yuan ZM (2007) Molecular characterization of a glucokinase with broad hexose specificity from Bacillus sphaericus strain C3-41. Appl Environ Microbiol 73:3581–3586
Hansen T, Reichstein B, Schmid R, Schonheit P (2002) The first archaeal ATP-dependent glucokinase, from the hyperthermophilic crenarchaeon Aeropyrum pernix, represents a monomeric, extremely thermophilic ROK glucokinase with broad hexose specificity. J Bacteriol 184:5955–5965
Helmann JD (1995) Complication and analysis of Bacillus subtilis σA-dependent promoter sequence: evidence for extended contact between RNA polymerase and upstream promoter DNA. Nucleic Acids Res 23:2351–2360
Kamata K, Mitsuya M, Nishimura T, Eiki JI, Nagata Y (2004) Structural basis for allosteric regulation of the monomeric allosteric enzyme human glucokinase. Structure 12:429–438
Karst D, Yang Y (2006) Molecular modeling study of the resistance of PLA to hydrolysis based on the blending of PLLA and PDLA. Polymer 47:4845–4850
Kawai S, Mukai T, Mori S, Mikami B, Murata K (2005) Hypothesis: structures, evolution, and ancestor of glucose kinases in the hexokinase family. J Biosci Bioeng 99:320–330
Kornberg HL, Reeves RE (1972) Inducible phosphoenolpyruvate-dependent hexose phosphotransferase activities in Escherichia coli. Biochem J 128:1339–1344
Len MCL, Harty DWS, Jacques NA (2004) Proteome analysis of Streptococcus mutans metabolic phenotype during acid tolerance. Microbiology 150:1353–1366
Meyer D, Schneider-Fresenius C, Horlacher R, Peist R, Boos W (1997) Molecular characterization of glucokinase from Escherichia coli K-12. J Bacteriol 179:1298–1306
Mishra RN, Singla-Pareek SL, Nair S, Sopory SK, Reddy MK (2002) Directional genome walking using PCR. Biotechniques 33:830–834
Mukai T, Kawai S, Matsukawa H, Matuo Y, Murata K (2003) Characterization and molecular cloning of a novel enzyme, inorganic polyphosphate/ATP-glucomannokinase, of Arthrobacter sp. strain KM. Appl Environ Microbiol 69:3849–3857
Nakayama T, Soma M, Rahmutula D, Ozawa Y, Kanmatsuse K (2001) Isolation of the 5′-flanking region of genes by thermal asymmetric interlaced polymerase chain reaction. Med Sci Monit 7:345–349
Panneman H, Ruijter GJ, van den Broeck HC, Driever ET, Visser J (1996) Cloning and biochemical characterisation of an Aspergillus niger glucokinase. Evidence for the presence of separate glucokinase and hexokinase enzymes. Eur J Biochem 240:518–525
Paulsen IT, Chauvaux S, Choi P, Saier MH Jr (1998) Characterization of glucose-specific catabolite repression-resistant mutants of Bacillus subtilis: identification of a novel hexose:H+ symporter. J Bacteriol 180:498–504
Paulsen IT, Nguyen L, Sliwinski MK, Rabus R, Saier MH Jr (2000) Microbial genome analyses: comparative transport capabilities in eighteen prokaryotes. J Mol Biol 301:75–100
Porter EV, Chassy BM, Holmlund CE (1982) Purification and kinetic characterization of a specific glucokinase from Streptococcus mutans OMZ70 cells. Biochim Biophys Acta 709:178–186
Sakuraba H, Yoshioka I, Koga S, Takahashi M, Kitahama Y, Satomura T, Kawakami R, Ohshima T (2002) ADP-dependent glucokinase/phosphofructokinase, a novel bifunctional enzyme from the hyperthermophilic archaeon Methanococcus jannaschii. J Biol Chem 277:12495–12498
Siegumfeldt H, Björn Rechinger K, Jakobsen M (2000) Dynamic changes of intracellular pH in individual lactic acid bacterium cells in response to a rapid drop in extracellular pH. Appl Environ Microbiol 66:2330–2335
Skarlatos P, Dahl MK (1998) The glucose kinase of Bacillus subtilis. J Bacteriol 180:3222–3226
Stülke J, Hillen W (2000) Regulation of carbon catabolism in Bacillus species. Annu Rev Microbiol 54:849–880
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Ureta T (1982) The comparative isozymology of vertebrate hexokinases. Comp Biochem Physiol B 71:549–555
Vadeboncoeur C, St Martin S, Brochu D, Hamilton IR (1991) Effect of growth rate and pH on intracellular levels and activities of the components of the phosphoenolpyruvate: sugar phosphotransferase system in Streptococcus mutans Ingbritt. Infect Immun 59:900–906
Xu H, Teng CQ, Yu MH (2006) Improvements of thermal property and crystallization behavior of PLLA-based multiblock copolymer by forming stereocomplex with PDLA oligomer. Polymer 47:3922–3928
Xu TT, Bai ZZ, Wang LJ, He BF (2010) Breeding of D(−)-lactic acid high producing strain by low-energy ion implantation and preliminary analysis of related metabolism. Appl Biochem Biotechnol 160:314–321
Yu B, Su F, Wang LM, Xu K, Zhao B, Xu P (2011) Draft genome sequence of Sporolactobacillus inulinus strain CASD, an efficient d-lactic acid-producing bacterium with high-concentration lactate tolerance capability. J Bacteriol 193:5864–5865
Zhao B, Wang LM, Li FS, Hua DL, Ma CQ, Ma YH, Xu P (2010) Kinetics of d-lactic acid production by Sporolactobacillus sp. strain CASD using repeated batch fermentation. Bioresour Technol 101:6499–6505
Zheng HJ, Gong JX, Chen T, Chen X, Zhao XM (2010) Strain improvement of Sporolactobacillus inulinus ATCC 15538 for acid tolerance and production of d-lactic acid by genome shuffling. Appl Microbiol Biotechnol 85:1541–1549
Acknowledgments
We acknowledge the support of the National Basic Research Program of China (2011CB707400) and the National High Technology Research and Development Program of China (2011AA02A202). This work was also supported by the projects funded by PCSIRT, PAPD and CXZZ11_0358.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zheng, L., Bai, Z., Xu, T. et al. Glucokinase contributes to glucose phosphorylation in d-lactic acid production by Sporolactobacillus inulinus Y2-8. J Ind Microbiol Biotechnol 39, 1685–1692 (2012). https://doi.org/10.1007/s10295-012-1176-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-012-1176-z