Abstract
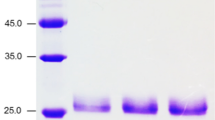
Xylan constitutes the second most abundant source of renewable organic carbon on earth and is located in the cell walls of hardwood and softwood plants in the form of hemicellulose. Based on its availability, there is a growing interest in production of xylanolytic enzymes for industrial applications. β-1,4-xylan xylosidase (EC 3.2.1.37) hydrolyses from the nonreducing end of xylooligosaccharides arising from endo-1,4-β-xylanase activity. This work reports the partial characterization of a purified β-xylosidase from the native strain Aspergillus niger GS1 expressed by means of a fungal system. A gene encoding β-xylosidase, xlnD, was successfully cloned from a native A. niger GS1 strain. The recombinant enzyme was expressed in A. niger AB4.1 under control of A. nidulans gpdA promoter and trpC terminator. β-xylosidase was purified by affinity chromatography, with an apparent molecular weight of 90 kDa, and showed a maximum activity of 4,280 U mg protein−1 at 70°C, pH 3.6. Half-life was 74 min at 70°C, activation energy was 58.9 kJ mol−1, and at 50°C optimum stability was shown at pH 4.0–5.0. β-xylosidase kept residual activity >83% in the presence of dithiothreitol (DTT), β-mercaptoethanol, sodium dodecyl sulfate (SDS), ethylenediaminetetraacetate (EDTA), and Zn2+. Production of a hemicellulolytic free xylosidase showed some advantages in applications, such as animal feed, enzymatic synthesis, and the fruit-juice industry where the presence of certain compounds, high temperatures, and acid media is unavoidable in the juice-making process.







Similar content being viewed by others
References
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (1999) Short protocols in molecular biology. Wiley, New York, pp 2–11
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bendtsen JD, Nielsen H, von Heijne G, Brunak S (2004) Improved prediction of signal peptides: signalP 3.0. J Mol Biol 340:783–795. doi:10.1016/j.jmb.2004.05.028
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
de Vries RP, Visser JAAP (2001) Aspergillus enzymes involved in degradation of plant cell wall polysaccharides. Microbiol Mol Biol Rev 65:497–522
Han Y, Chen H (2010) A β-xylosidase from cell wall of maize: purification, properties and its use in hydrolysis of plant cell wall. J Mol Catal B: Enzym 63:135–140
Kainz E, Gallmetzer A, Hatzl C, Nett JH, Li H, Schinko T, Pachlinger R, Berger H, Reyes-Dominguez Y, Bernreiter A, Gerngross T, Wildt S, Strauss J (2008) N-Glycan modification in Aspergillus Species. Appl Environ Microbiol 74(4):1076–1086. doi:10.1128/AEM.01058-07
Kumar S, Ramón D (1996) Purification and regulation of the synthesis of a β-xylosidase from Aspergillus nidulans. FEMS Microbiol Lett 135:287–293
Knob A, Terrasan CRF, Carmona EC (2010) β-Xylosidases from filamentous fungi: an overview. World J Microbiol Biotechnol 26:389–407. doi:10.1007/s11274-009-0190-4
La Grange DC, Pretorius IS, Claeyssens M, van Zyl WH (2001) Degradation of xylan to d-xylose by recombinant Saccharomyces cerevisiae coexpressing the Aspergillus niger β-xylosidase (xlnD) and the Trichoderma reesei xylanase II (xyn2) genes. Appl Environ Microbiol 67(12):5512–5519. doi:10.1128/AEM.67.12.5512-5519.2001
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lenartovicz V, Marques Giatti, de Souza C, Guillen Moreira F, Marina Peralta R (2003) Temperature and carbon source affect the production and secretion of a thermostable β-xylosidase by Aspergillus fumigatus. Process Biochem 38:1775–1780. doi:10.1016/S0032-9592(02)00261-3
Linton CJ, Borman AM, Cheung C, Holmes AD, Szekely A, Palmer MD, Bridge PD, Campbell CK, Johnson EM (2007) Molecular identification of unusual pathogenic yeast isolates by large ribosomal subunit gene sequencing: 2 years of experience at the United Kingdom mycology reference laboratory. J Clin Microbiol 45(4):1152–1158. doi:10.1128/JCM.02061-06
Liu C, Sun Z, Du J, Wang J (2008) Response surface optimization of fermentation conditions for producing xylanase by Aspergillus niger SL-05. J Ind Microbiol Biotechnol 35:703–711
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31(3):426–428. doi:10.1021/ac60147a030
Okafor U, Okochi V, Onyegeme-Okerenta B, Nwodo-Chinedu S (2007) Xylanase production by Aspergillus niger ANL 301 using agro-wastes. Afr J Biotechnol 6:1710–1714
Pachlinger R, Mitterbauer R, Adam G, Strauss J (2005) Metabolically independent and accurately adjustable Aspergillus sp. expression system. Appl Environ Microbiol 71(2):672–678. doi:10.1128/AEM.71.2.672-678.2005
Pedersen M, Lauritzen HK, Frisvad JC, Meyer AS (2007) Identification of thermostable β-xylosidase activities produced by Aspergillus brasiliensis and Aspergillus niger. Biotechnol Lett 29:743–748
Pérez-González JA, van Peij NNME, Bezoen A, Maccabe AP, Ramón D, De Graaff LH (1998) Molecular cloning and transcriptional regulation of the Aspergillus nidulans xlnD gene encoding a β-xylosidase. Appl Environ Microbiol 64(4):1412–1419
Polizeli MLTM, Rizzatti ACS, Monti R, Terenzi HF, Jorge JA, Amorim DS (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67:577–591. doi:10.1007/s00253-005-1904-7
Prathumpai W, McIntyre M, Nielsen J (2004) The effect of CreA in glucose and xylose catabolism in Aspergillus nidulans. Appl Microbiol Biotechnol 63:748–753. doi:10.1007/s00253-003-1409-1
Rasmussen LE, Sørensen HR, Vind J, Viksø-Nielsen A (2006) Mode of action and properties of the β-xylosidases from Talaromyces emersonii and Trichoderma reesei. Biotechnol Bioeng 94(5):871–876. doi:10.1002/bit.20908
Regalado C, García-Almendárez B, Venegas-Barrera L, Tellez-Jurado A, Rodríguez-Serrano G, Huerta-Ochoa S, Whitaker J (2000) Production, partial purification and properties of-mannanases obtained by solid substrate fermentation of spent soluble coffee wastes and copra paste using Aspergillus oryzae and Aspergillus niger. J Sci Food Agric 80:1343–1350
Rizzatti A, Jorge J, Terenzi H, Rechia C, Polizeli M (2001) Purification and properties of a thermostable extracellular β-d-xylosidase produced by a thermotolerant Aspergillus phoenicis. J Ind Microbiol Biotechnol 26:156–160
Sanchez O, Aguirre J (1996) Efficient transformation of Aspergillus nidulans by electroporation of germinated conidia. Fungal Genet Newsl 43:48–51
Selig MJ, Knoshaug EP, Decker SR, Baker JO, Himmel ME, Adney WS (2008) Heterologous expression of Aspergillus niger β-d-Xylosidase (XlnD): characterization on lignocellulosic substrates. Appl Biochem Biotechnol 146:57–68
Shallom D, Shoham Y (2003) Microbial hemicellulases. Curr Op Microbiol 6:219–228. doi:10.1016/S1369-5274(03)00056-0
Singh S, Madlala AM, Prior BA (2003) Thermomyces lanuginosus: properties of strains and their hemicellulases. FEMS Microbiol Rev 27:3–16
van Hartingsveldt W, Mattern IE, van Zeijl CMJ, Pouwels PH, van den Hondel CAMJJ (1978) Development of a homologous transformation system for Aspergillus niger based on the pyrG gene. Mol Gen Genet 206:71–75
Wakiyama M, Yoshihara K, Hayashi S, Ohta K (2008) Purification and properties of an extracellular β-xylosidase from Aspergillus japonicus and sequence analysis of the encoding gene. J Biosci Bioeng 106(4):398–404. doi:10.1263/jbb.106.398
Yuan XL, van der Kaaij RM, van den Hondel CAMJJ, Punt PJ, van der Maarel MJEC, Dijkhuizen L, Ram AFJ (2008) Aspergillus niger genome-wide analysis reveals a large number of novel alpha-glucan acting enzymes with unexpected expression profiles. Mol Genet Genomics 279:545–561. doi:10.1007/s00438-008-0332-7
Acknowledgments
Thanks are given to Consejo Nacional de Ciencia y Tecnología for PhD grant to AAR.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Amaro-Reyes, A., García-Almendárez, B.E., Vázquez-Mandujano, D.G. et al. Homologue expression of a β-xylosidase from native Aspergillus niger . J Ind Microbiol Biotechnol 38, 1311–1319 (2011). https://doi.org/10.1007/s10295-010-0912-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-010-0912-5