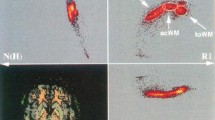
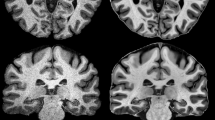
Abstract
Segmentation of MRI scans is a critical part of the workflow process before we can further analyze neuroimaging data. Although there are several automatic tools for segmentation, no segmentation software is perfectly accurate, and manual correction by visually inspecting the segmentation errors is required. The process of correcting these errors is tedious and time-consuming, so we present a novel method of performing this task in a head-mounted virtual reality interactive system with a new software, Virtual Brain Segmenter (VBS). We provide the results of user testing on 30 volunteers to show the benefits of our tool as a more efficient, intuitive, and engaging alternative compared with the current method of correcting segmentation errors.




Similar content being viewed by others
References
Anand KS, Dhikav V: Hippocampus in health and disease: An overview. Ann Indian Acad Neurol 15:239–246, 2012
Bayyari A, Tudoreanu ME: The impact of immersive virtual reality displays on the understanding of data visualization. In: Proc. ACM Sym. on Virtual Reality Software and Technology:368–371, 2006
Cherbuin N, Anstey K, Reglade-Meslin C, Sachdev PS: In vivo hippocampal measurement and memory: A comparison of manual tracing and automated segmentation in a large community-based sample. PLoS ONE. https://doi.org/10.1371/journal.pone.0005265, April 16, 2009
Cho I, Dou W, Wartell Z, Ribarsky W, Wang X: Evaluating depth perception of volumetric data in semi-immersive VR. In: IEEE Virtual Reality Workshops. https://doi.org/10.1145/2254556.2254606, 2012
Chupin M, Hammers A, Liu RS Colliot O, Burdett J, Bardinet E, Duncan JS, Garnero L, Lemieux L: Automatic segmentation of the hippocampus and the amygdala driven by hybrid constraints: method and validation. Neuroimage, 46:749–761, 2009
Cooper S, Khatib F, Treuille A, Barbero J, Lee J, Beenen M, Leaver-Fay A, Baker D, Popović Z, players F: Predicting protein structures with a multiplayer online game. Nature, 466:756–760, 2010
Cordero P, Lucks JB, Das R: An RNA mapping database for curating RNA structure mapping experiments. Bioinformatics, 28:3006–3008, 2012
Fairén M, Farres M, Moyes J, Insa E: Virtual reality to teach anatomy. In: Proc. Eurographics. https://doi.org/10.2312/eged.201710262017, 2017
Fischl B. FreeSurfer. Neuroimage, 62:774–81, 2012
Fox NC, Warrington EK, Freeborough PA, Hartikainen P, Kennedy AM, Stevens JM, Rossor MN: Presymptomatic hippocampal atrophy in Alzheimer’s disease. Brain, 119:2001–2007, 1996
Ganz M, Kondermann D, Andrulis J, Knudsen GM, Maier-Hein L: Crowdsourcing for error detection in cortical surface delineations. Int J Comput Assist Radiol Surg 12:161–166, 2017
Garcia-Palacios A, Hoffman HG, Richards TR, Seibel EJ, Sharar SR: Use of virtual reality distraction to reduce claustrophobia symptoms during a mock magnetic resonance imaging brain scan: A case report. Cyberpsychol Behav 10:485–488, 2007
Herbelin B, Riquier F, Vexo F, Thalmann D. Virtual reality in cognitive behavioral therapy: A preliminary study on social anxiety disorder. In: Proc. Int. Conf. on Virtual Systems and Multimedia, 2002
Jack, Jr CR, Theodore WH, Cook M, McCarthy G: MRI-based hippocampal volumetrics: data acquisition, normal ranges, and optimal protocol. Magn Reson Imaging 13:1057–1064, 1995
Kim JS, Greene MJ, Zlateski A, Lee K, Richardson M, Turaga SC, Purcaro M, Balkam M, Robinson A, Behabadi BF, Campos M, Denk W, Seung HS, Eyewirers: Space-time wiring specificity supports direction selectivity in the retina. Nature 509:331–336, 2014
Li Z, Wang Y, Yu J, Shi Z, Guo Y, Chen L, Mao Y. Low-grade glioma segmentation based on CNN with fully connected CRF. J Healthc Eng, https://doi.org/10.1155/2017/9283480, June 13, 2017
Lötjönen JM, Wolz R, Koikkalainen JR, Thurfjell L, Waldemar G, Soininen H, Rueckert D, Alzheimer’s Disease Neuroimaging Initiative: Fast and robust multi-atlas segmentation of brain magnetic resonance images. Neuroimage 49:2352–2365, 2010
Marx V: Neuroscience waves to the crowd. Nature methods 10:1069–1074, 2013
Morey RA, Petty CM, Xu Y, Pannu Hayes J, Wagner, II HR, Lewis DV, LaBar KS, Styner M, McCarthy G: A comparison of automated segmentation and manual tracing for quantifying hippocampal and amygdala volumes. Neuroimage 45:855–866, 2009
Pardoe HR, Pell GS, Abbott DF, Jackson GD: Hippocampal volume assessment in temporal lobe epilepsy: How good is automated segmentation? Epilepsia 50:2586–2592, 2009
Pronk NP, Katz AS, Lowry M, Payfer JR: Reducing Occupational Sitting Time and Improving Worker Health: The Take-a-Stand Project, 2011. Prev Chronic Dis, https://doi.org/10.5888/pcd9.110323, October 11, 2012
Rowles TA: Power to the people: Does Eterna signal the arrival of a new wave of crowd-sourced projects? BMC Biochem 14:1, 2013
Shamir L, Yerby C, Simpson R, von Benda-Beckmann AM, Tyack P, Samarra F, Miller P, Wallin J: Classification of large acoustic datasets using machine learning and crowdsourcing: Application to whale calls. J. Acoust. Soc. Am. 135:953–962, 2014
Shoemaker D, Buss C, Head K, Sandman CA, Davis EP, Chakravarty MM, Gauthier S, Pruessner JC: Hippocampus and amygdala volumes from magnetic resonance images in children: Assessing accuracy of FreeSurfer and FSL against manual segmentation. Neuroimage 129:1–14, 2016
Sun C, Rampalli N, Yang F, Doan AH: Large-scale classification using machine learning, rules, and crowdsourcing. In: Proc. VLDB Endowment 7:1529–1540, 2014
Van Dam A, Laidlaw DH, Simpson RM: Experiments in immersive virtual reality for scientific visualization. Computer and Graphics 26:535–555, 2002
Van de Pol LA, Hensel A, van der flier WM, Visser PJ, Pijnenburg YA, Barkhof F, Gerz HJ, Scheltens P: Hippocampal atrophy on MRI in frontotemporal lobar degeneration and Alzheimer’s disease. J Neurol Neurosurg Psychiatry 77:439–442, 2006
Wang H, Das SR, Suh JW, Altinay M, Pluta J, Craige C, Avants B, Yushkevich PA, Alzheimer’s Disease Neuroimaging Initiative: A learning-based wrapper method to correct systematic errors in automatic image segmentation: Consistently improved performance in hippocampus, cortex and brain segmentation. Neuroimage 55:968–985, 2011
Wang JY, Ngo M, Hessl D, Hagerman RJ, Rivera SM: Robust machine learning-based correction on automatic segmentation of the cerebellum and brainstem. PLoS ONE, https://doi.org/10.1371/journal.pone.0156123, May 23, 2016
Wang Y, Otitoju K, Liu T, Bowman DA: Evaluating the effect of real-world distraction on user performance in virtual environments. In: Proc. ACM Symp. Virtual Reality Software and Technology;19–26, 2006
Williams D. Cubiquity for Unity (Voxel Engine). Retrieved September 26, 2016 from https://www.assetstore.unity3d.com/en/#!/content/12689
Wu H, Sun H, Fang Y, Hu K, Xie Y, Song Y, Liu X: Combining machine learning and crowdsourcing for better understanding commodity reviews. In: AAAI; 4220–4221, 2015
Xue H, Srinivasan L, Jiang S, Rutherford M, Edwards AD, Rueckert D, Hajnal JV: Automatic segmentation and reconstruction of the cortex from neonatal MRI. Neuroimage 38:461–477, 2007
Zheng W, Chee MWL, Zagorodnov V: Improvement of brain segmentation accuracy by optimizing non-uniformity correction using N3. Neuroimage 48:73–83, 2009
Funding
This work is supported by the National Institutes of Health grants P41-EB015922, U54EB0020406, and the University of Southern California Provost’s Postdoctoral Scholar Research Grant.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
ESM 1
(MP4 14868 kb)
Rights and permissions
About this article
Cite this article
Duncan, D., Garner, R., Zrantchev, I. et al. Using Virtual Reality to Improve Performance and User Experience in Manual Correction of MRI Segmentation Errors by Non-experts. J Digit Imaging 32, 97–104 (2019). https://doi.org/10.1007/s10278-018-0108-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-018-0108-5