Abstract
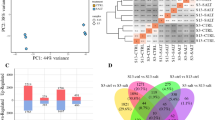
The small genome size of rice relative to wheat and barley, together with its salt sensitivity, make it an ideal candidate for studies of salt stress response. Transcriptomics has emerged as a powerful technique to study salinity responses in many crop species. By identifying a large number of differentially expressed genes (DEGs) simultaneously after the stress induction, it can provide crucial insight into the immediate responses towards the stressor. In this study, a Malaysian salt-tolerant indigenous rice variety named Bajong and one commercial rice variety named MR219 were investigated for their performance in plant growth and ion accumulation properties after salt stress treatment. Bajong was further investigated for the changes in leaf’s transcriptome after 6 h of stress treatment using 100 mM NaCl. Based on the results obtained, Bajong is found to be significantly more salt tolerant than MR219, showing better growth and a lower sodium ion accumulation after the stress treatment. Additionally, Bajong was analysed by transcriptomic sequencing, generating a total of 130 millions reads. The reads were assembled into de novo transcriptome and each transcript was annotated using several pre-existing databases. The transcriptomes of control and salt-stressed samples were then compared, leading to the discovery of 4096 DEGs. Based on the functional annotation results obtained, the enrichment factor of each functional group in DEGs was calculated in relation to the total reads obtained. It was found that the group with the highest gene modulation was involved in the secondary metabolite biosynthesis of plants, with approximately 2.5% increase in relation to the total reads obtained. This suggests an extensive transcriptional reprogramming of the secondary metabolic pathways after stress induction, which could be directly responsible for the salt tolerance capability of Bajong.





Similar content being viewed by others
Abbreviations
- COG:
-
Clusters of orthologous groups
- DEGs:
-
Differentially expressed genes
- DPPH:
-
Di(phenyl)-(2,4,6-trinitrophenyl) iminoazanium
- GMO:
-
Genetically modified organism
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of gene and genome
- LEA:
-
Late embryogenesis abundant
- ROS:
-
Reactive oxygen species
- RWC:
-
Relative water content
- SOS:
-
Salt overly sensitive
- TFC:
-
Total flavonoid content
- TPC:
-
Total phenolic content
References
Aronson JA (1989) HALOPH: a data base of salt tolerant plants of the world. Office of Arid Land Studies, University of Arizona, Tuscon, AZ
Ashraf M, Foolad MR (2007) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59:206–216
Bharti N, Yadav D, Barnawal D, Maji D, Kalra A (2013) Exiguobacterium oxidotolerans, a halotolerant plant growth promoting rhizobacteria, improves yield and content of secondary metabolites in Bacopa monnieri (L.) Pennell under primary and secondary salt stress. World J Microbiol Biotechnol 29:379–387
Bohnert HJ, Ayoubi P, Borchert C, Bressan RA, Burnap RL, Cushman JC, Cushman MA, Deyholos M, Fischer R, Galbraith DW (2001) A genomics approach towards salt stress tolerance. Plant Physiol Biochem 39:295–311
Brondani C, Borba TCO, Rangel PHN, Brondani RPV (2006) Determination of genetic variability of traditional varieties of Brazilian rice using microsatellite markers. Genet Mol Biol 29:676–684
D’Auria JC, Gershenzon J (2005) The secondary metabolism of Arabidopsis thaliana: growing like a weed. Curr Opin Plant Biol 8:308–316
Dharmawardhana P, Ren L, Amarasinghe V, Monaco M, Thomason J, Ravenscroft D, McCouch S, Ware D, Jaiswal P (2013) A genome scale metabolic network for rice and accompanying analysis of tryptophan, auxin and serotonin biosynthesis regulation under biotic stress. Rice 6:1–15
Garg R, Verma M, Agrawal S, Shankar R, Majee M, Jain M (2014) Deep transcriptome sequencing of wild halophyte rice, Porteresia coarctata, provides novel insights into the salinity and submergence tolerance factors. DNA Res 21:69–84
Gilroy S, Suzuki N, Miller G, Choi WG, Toyota M, Devireddy AR, Mittler R (2014) A tidal wave of signals: calcium and ROS at the forefront of rapid systemic signaling. Trends Plant Sci 19:623–630
Golldack D, Li G, Mohan H, Probst N (2014) Tolerance to drought and salt stress in plants: unraveling the signaling networks. Mol Genet Genom 1:15–25
Gou JY, Yu XH, Liu CJ (2009) A hydroxycinnamoyltransferase responsible for synthesizing suberin aromatics in Arabidopsis. Proc Natl Acad Sci USA 106:18,855–18,860
Herald TJ, Gadgil P, Tilley M (2012) High-throughput micro plate assays for screening flavonoid content and DPPH-scavenging activity in sorghum bran and flour. J Sci Food Agric 92:2,326–2,331
Hernández JA, Corpas FJ, Gomez M, Río LA, Sevilla F (1993) Salt-induced oxidative stress mediated by activated oxygen species in pea leaf mitochondria. Physiol Plant 89:103–110
Ismail C (2005) The role of potassium in alleviating detrimental effects of abiotic stresses in plants. J Plant Nutr Soil Sci 168:521–530
Joshi SH, Gupta VS, Aggarwal RK, Ranjekar PK, Brar DS (2000) Genetic diversity and phylogenetic relationship as revealed by inter simple sequence repeat (ISSR) polymorphism in the genus Oryza. Theor Appl Genet 100:1,311–1,320
Kim JH, Kim WT (2013) The Arabidopsis RING E3 ubiquitin ligase AtAIRP3/LOG2 participates in positive regulation of high-salt and drought stress responses. Plant Physiol Biochem 162:1,733–1,749
Lee HH, Neoh PPN, Bong WST, Puvaneswaran J, Wong SC, Yiu PH, Rajan A (2011) Genotyping of Sarawak rice cultivars using microsatellite markers. Pertan J Trop Agric Sci 34:123–136
Leigh RA, Wyn Jones RG (1984) A hypothesis relating critical potassium concentrations for growth to the distribution and functions of this ion in the plant cell. New Phytol 97:1–3
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinf 12:323
Li T, Li H, Zhang YX, Liu JY (2011) Identification and analysis of seven H2O2-responsive miRNAs and 32 new miRNAs in the seedlings of rice (Oryza sativa L. subsp. indica). Nucleic Acids Res 39:2,821–2,833
Li L, Li N, Song SF, Li YX, Xia XJ, Fu XQ, Chen GH, Deng HF (2014) Cloning and characterization of the drought-resistance OsRCI2-5 gene in rice (Oryza sativa L.). GMR Genet Mol Res 13:4,022–4,035
Lugan R, Niogret MF, Kervazo L, Larher FR, Kopka J, Bouchereau A (2009) Metabolome and water status phenotyping of Arabidopsis under abiotic stress cues reveals new insight into ESK1 function. Plant Cell Environ 32:95–108
Mandhania S, Madan S, Sawhney V (2006) Antioxidant defense mechanism under salt stress in wheat seedlings. Biol Plant 50:227–231
Mekawy AM, Assaha DV, Yahagi H, Tada Y, Ueda A, Saneoka H. Growth (2015) Physiological adaptation, and gene expression analysis of two Egyptian rice cultivars under salt stress. Plant Physiol Biochem 87:17–25
Meloni DA, Gulotta MR, Martínez CA, Oliva MA (2004) The effects of salt stress on growth, nitrate reduction and proline and glycinebetaine accumulation in Prosopis alba. Braz J Plant Physiol 16:39–46
Menzel U, Lieth H (2003) HALOPHYTE Database Ver. 2.0 update. In: Cash crop halophytes, pp 221–223
Munns R (2002) Comparative physiology of salt and water stress. Plant Cell Environ 25:239–250
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Rahman MA, Thomson MJ, Shah-E-Alam M, de Ocampo M, Egdane J, Ismail AM (2016) Exploring novel genetic sources of salinity tolerance in rice through molecular and physiological characterization. Ann Bot 117:1,083–1,097
Ren HY, Wei Q (2015) Isolation and functional analysis of phosphate-responsive gene OsRCI2-9 in Oryza sativa. Zhongguo Nongye Kexue 48:831–840
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1,101–1,108
Singleton VL, Rossi JA (1965) Colorimetry of total phenolics with phosphomolybdic-phosphotungstic acid reagents. Am J Enol Vitic 16:144–158
Smart RE, Bingham GE (1974) Rapid estimates of relative water content. Plant Physiol 53:258–260
Tavakkoli E, Rengasamy P, McDonald GK (2010) High concentrations of Na+ and Cl− ions in soil solution have simultaneous detrimental effects on growth of faba bean under salinity stress. J Exp Biol 61:4,449–4,459
Tester M, Davenport R (2003) Sodium ion tolerance and sodium ion transport in higher plants. Ann Bot 91:503–527
Thomson WW (1975) The structure and function of salt glands. Plants Saline Environ 1:118–146
Van Poecke RM, Posthumus MA, Dicke M (2001) Herbivore-induced volatile production by Arabidopsis thaliana leads to attraction of the parasitoid Cotesia rubecula: chemical, behavioral, and gene-expression analysis. J Chem Ecol 27:1,911–1,928
Walia H, Wilson C, Condamine P, Liu X, Ismail AM, Zeng L, Wanamaker SI, Mandal J, Xu J, Cui X, Close TJ (2005) Comparative transcriptional profiling of two contrasting rice genotypes under salinity stress during the vegetative growth stage. Plant Physiol 139:822–835
Wang M, Zheng Q, Shen Q, Guo S (2013) The critical role of potassium in plant stress response. Int J Mol Sci 14:7,370–7,390
Winkel-Shirley B (2002) Biosynthesis of flavonoids and effects of stress. Curr Opin Plant Biol 5:218–223
Yeo AR, Flowers TJ (1984) Nonosmotic effects of polyethylene glycols upon sodium transport and sodium-potassium selectivity by rice roots. Plant Physiol 75:298–303
Acknowledgements
We greatly thank Swinburne University of Technology Sarawak Campus for providing the Strategic Research Grant (StraRG 2-5605).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yeo, B.P.H., Bhave, M. & Hwang, S.S. Effects of acute salt stress on modulation of gene expression in a Malaysian salt-tolerant indigenous rice variety, Bajong. J Plant Res 131, 191–202 (2018). https://doi.org/10.1007/s10265-017-0977-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-017-0977-6