Abstract
The prevalence of HCV infection in Egypt has decreased following the introduction of direct-acting antiviral therapy. However, treatment response is influenced by various factors, particularly host immunogenetics such as IL-28B and FOXP3 polymorphisms. The current study examined the impact of SNPs in the FOXP3 gene promoter region on HCV-infected Egyptian patients, along with SNPs in the IL28B gene.This study involved 99 HCV patients who achieved SVR12 after a 12 week DAA treatment while 63 HCV patients experienced treatment failure. IL28B rs12979860 SNP was identified using real-time PCR, while IL28B rs8099917, FOXP3 rs3761548, and rs2232365 SNPs were analyzed using RFLP-PCR. Serum levels of IL28B and FOXP3 were quantified using ELISA technique in representative samples from both groups. The IL28B rs12979860 T > C (P = 0.013) and FOXP3 rs2232365 A > G polymorphisms (P = 0.008) were found to significantly increase the risk of non-response. Responders had higher IL28B serum levels (P = 0.046) and lower FOXP3 levels (P < 0.001) compared to non-responders. Regression analysis showed an association between IL28B rs12979860 and FOXP3 rs2232365 with treatment response, independent of age and gender. A predictive model was developed with 76.2% sensitivity and 91.9% specificity for estimating DAAs response in HCV patients.Our findings confirmed the IL28B rs12979860 T > C and FOXP3 rs2232365 A > G polymorphisms significantly affect DAA treatment response in HCV Egyptian patients. Lower levels of IL-28B along with higher levels of FOXP3 are linked to poor response. Our results may lead to new insights into DAA responsiveness contributing to personalized medicine and improving therapeutic decision-making for HCV patients.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Based on the latest estimate of the World Health Organization (WHO), 58 million people have chronic hepatitis C virus (HCV) infection worldwide. Chronic HCV is an urgent worldwide health issue leading to liver cirrhosis which increase the development of hepatocellular carcinoma (HCC), and liver transplantation [1, 2]. Due to the use of poorly sanitized glass syringes for parenteral anti-schistosomal treatment between 1920 and 1980, Egypt was among the Eastern Mediterranean regions with higherrates of HCV infection, with an estimated prevalence of around 14% [3, 4]. Currently, healthcare-related transmission is the main route of HCV infection in Egypt, with approximately 150,000 new infections per year [5]. Among the eight confirmed genotypes of HCV, genotype 4 (GT4) accounts for 91% of HCV-infected Egyptian patients [6,7,8].
Previously, pegylated interferon (PEG-IFN) and ribavirin (RBV) were considered the gold standard for treating all genotypes of HCV. However, this treatment was non-specific and associated with several adverse events. In contrast to nonspecific treatment with PEG-IFN, the introduction of oral direct-acting antiviral (DAA) therapy has revolutionized the management of HCV patients [2]. DAAs target specific regions of the HCV genome and are categorized as inhibitors of NS3/4A proteases, NS5B nucleoside polymerase (such as Sofosbuvir [SOF]), NS5B non-nucleoside polymerase, and NS5A inhibitors (such as Daclatasvir [DAC]) [9, 10]. Over 90% sustained virological response at 12 weeks (SVR12) has been achieved by DAAs combinations [11,12,13]. The substantial cost of DAA therapy coupled with the high occurrence of HCV infection in Egypt, leads to animmense economic burden that necessitates adequate cure rates [14]. Moreover, it is critical to address the issue of patients who have failed DAA therapy, making recovery more challenging.
Recent evidence highlights that in addition to the viral factor, host genetic factors may be involved in the success or failure of antiviral therapy and HCV clearance [15]. Three genome-wide association studies (GWAS) have identified single nucleotide polymorphisms (SNPs) near the interleukin 28B (IL-28B) gene that are strongly associated with SVR to treatment with pegylated interferon alpha (PEG-IFN-α) and/or ribavirinalone. They stated that the most significant polymorphisms of the IL-28B gene were rs12979860, rs8099917, and rs12980275 [16,17,18]. Research on IL-28B polymorphisms in various populations has demonstrated that individuals who harbor the CC genotype of the rs12979860 have a better response to DAAs as a treatment compared to other genotypes [19,20,21].
IL-28B is encoded on the interferon lambda (IFN-λ) gene which plays a significant role in the natural antiviral defenses against HCV [22, 23]. The IFN-λs have increased particular attention due to their association with the spontaneous clearance and inhibition of HCV replication via upregulation of the interferon signaling gene (ISG) [24]. Thus far, there have been limited investigations to study the association of IL-28B serum levels and the different genotypes of IL-28B in chronic HCV patients undergoing treatment with DAAs [25, 26].
Persistent antigen exposure leads to the exhaustion of T cells, which lessens their capacity to control the infection. Additionally, it can lead to regulatory T cell (Treg) activation, which suppresses the immune response. These dysregulations in the immune system contribute to the sustained presence of HCV infection leading to chronicity [27]. Treg activation may also affect the progress of liver disease by maintaining fibrogenesis and inflammatory tissue activity [28]. A pivotal marker of Tregs modulation is the transcription factor known asforkhead box P3 (FOXP3). It is a key factor in the expansion of diverse Treg lines and the maintenance of immunoregulation in different pathological conditions that are both autoimmune and infectious. FOXP3-expressing Tregs possess immunosuppressive properties and exert inhibitory effects on various effector immune cells. Therefore, the polymorphism in FOXP3 and the level of FOXP3 are vital in regulating immune responses [29, 30]. Given that the FOXP3 gene plays a crucial role in regulating the gene expression and activating Treg, its promoter region may include significant SNPs [31]. Among these SNPs, the C > T rs3761549, C > A rs3761548, and A > G rs2232365 SNPs are functionally defined as immunomodulatory [32]. Few studies have investigated FOXP3 polymorphisms in conditions other than autoimmune diseases.One study proposed that the immunomodulatory effects on individuals with viral hepatitis are influenced by FOXP3 SNPs, specifically, rs2232365, rs3761549, and rs3761548 [33]. Another recent study investigated the influence of rs3761547 SNP of FOXP3 in the inflammatory responses accompanying viral hepatitis [32].
Accordingly, our objective was to enhance our understanding of immune-related genes that may impact a patient’s response to DAA therapy. Specifically, we aimed to examine the effects of IL-28B SNPs (rs12979860, rs8099917) and FOXP3 SNPs (rs2232365 and rs3761548) on the response to Sofosbuvir and Daclatasvir combination therapy in chronic HCV Egyptian patients. Additionally, we evaluated the serum levels of IL-28B and FOXP3 in relation to treatment response and their associations with the studied SNPs.
Subjects and methods
Subject recruitment and study design
A retrospective cohort study was conducted, analyzing data from 162 patients with chronic HCV infection who attended the outpatient unit of the Hepatology and Gastroenterology Department, National LiverInstituteHospital, Menoufia University, Egypt from 2021 to 2022.All cases who participated in the study completed a 12 week treatment course of DAC 60 mg/day and SOF 400 mg/day, orally. The cases were then grouped into responders and non-responders based on their sustained virologic response (SVR) at the end of therapy.
Based on the inclusion criteria, a clinical examination and laboratory testing were performed on all selected patients. These tests included liver enzyme level tests such as alanine aminotransferase (ALT), aspartate aminotransferase (AST), total bilirubin, albumin, creatinine, and a complete blood count (including red blood cells (RBCs), white blood cells (WBCs), platelets, and hemoglobin (HB). Additionally, the AST to Platelet Ratio Index (APRI) Score and Fibrosis-4 (FIB-4) were calculated, and the international normalized ratio (INR), alpha-fetoprotein (AFP), serological tests like surface antigen of HBV (HBsAg) and anti-HCV, HCV RNA quantification, and fibro scan for the diagnosis of liver cirrhosis.
The following individuals were excluded; those diagnosed with decompensated liver disease, Child–Pugh B and C cirrhosis, ascites or history of ascites, hepatic encephalopathy or history of hepatic encephalopathy, patients complicated with hepatocellular carcinoma, serum creatinine > 2.5 mg/dL, pregnancy, and poorly controlled diabetes (HbA1c ≥ 8), INR ≥ 1.7, serum albumin < 2.8 g/dL, total serum bilirubin ≥ 3 mg/dL, platelets < 50 000/mm3. Additionally, patients with Hepatitis B virus (HBV), Human Immunodeficiency virus (HIV) co-infections, other liver disease causes, renal impairment,non-adherence to treatment, and those who declined to be enrolled in the study were excluded.
Ethics consideration
The protocol of our study follows the ethical guidelines of the Declaration of Helsinki (2013) and has been approved by the Institutional Review Board of the National Liver Institute (NLI IRB 00003143 ; protocol number 00252/2021) and Faculty of Pharmacy, University of Ain Shams (ACUC-FP-ASU.RHDIRB2020110301REC#37). We adhered to the STROBE (Strengthening the Reporting of Observational Studies in Epidemiology) guideline for this observational study.
Genetic analysis
Peripheral venous blood samples were divided into one vacutainer for serum preparation and one Na-EDTA vacutainer for DNA extraction. Genomic DNA extraction was conducted using a modified method of a previously described protocol [34].
Polymorphism genotyping
Genotyping of IL-28B C/T (rs12979860)
IL-28B genotyping was determined by Eco 48 Real-Time Polymerase chain reaction (PCR) using SYBR green assay (PCRmax Limited, Staffordshire, UK) using 25 ng extracted DNA and with 0.5 µl of each primer (Supp. Table 1) in 20 µl reaction mixture. We followed the PCR protocol by Zakaria et.al.[35]. (Fig. 1A–C).
Amplification plots of the genotypes for Interleukin IL28B (rs12979860) polymorphism. The amplification curves are based on the fluorescence intensity variation (ΔRn) according to the number of PCR cycles for the wild-type Heterozygous CT (A), Homozygous CC (B), and Mutant-Type Homozygous TT (C). RFLP of the amplified product of (D) For the genotyping IL28B rs8099917, the 552 bp PCR product was digested by BsrDI enzyme. The T allele is not cut by the enzyme where the G allele yields 322 and 230 bp products. (E) For the genotyping FOXP3 rs3761548, the 487 bp product was digested with PstI enzyme. The C allele is not cut by the enzyme, whereas the A allele yields 329 and 158 bp products (F) For the genotyping FOXP3 rs2232365, the 374 bp PCR product was digested with BsMBI enzyme. The A allele is not cut by the enzyme, whereas the G allele yields 188 and 186 bp products
Genotyping of IL-28B (rs8099917), FOXP3 [rs3761548], and FOXP3 (rs2232365)
Genotyping of IL-28B (rs8099917), FOXP3 intron 1 (rs3761548), and (rs2232365)
polymorphisms were conducted using the polymerase chain reaction-restriction fragment-length polymorphism (PCR–RFLP) method. The primer sequences for forward and reverse are provided in Table S1.The target sequences for each gene were amplified using a thermocycler 006 (a&e™, UK) with 50 ng of extracted DNA and 1 μl of each primer in a 20 μl PCR mixture (TOP simpleTM DyeMIX-nTaq Polymerase,Enzynomics, INC, South Korea). The calculated amount of PCR product was then digested using the relevant enzyme specified in Table S1 which were purchased from NEW ENGLAND BioLabsInc, MA. The digestion conditions were as follows:
For IL-28B G/T (rs8099917): Three μl of PCR product (552 bp in length) was digested using BsrDI enzyme at 65 °C for 30 min then separated on a 1.5% agarose gel stained with 5% ethidium bromide and examined under ultraviolet light (UV transilluminator, PHASE, Germany). The BsrDI digestion yielded 552 bp for the undigested allele G and 322, 230 bp for the T allele as shown in Fig. 1D.
For FOXP3 A/C (rs3761548): PstuI enzyme was used to digest 5 μl of PCR product (487 bp in length) at 37 °C for 40 min and separated on 1.5% agarose gel stained with ethidium bromide. The PstI digestion of the PCR product yielded 487 bp for the undigested allele A, and 329 and 158 bp for allele C as shown in Fig. 1E.
For FOXP3 A/G (rs2232365): A 5 μl aliquot of PCR product (374 bp in length) was digested using 200 nl BsMBI enzyme at 55 °C for 30 min and separated on a 1.5% agarose gel. The BsMBI digestion of PCR products yielded 374 bp for the A allele, whereas for allele G, 188,186 bp fragments were observed as presented in Fig. 1F.
To ensure result repeatability, a 10% sample of the subjects from both groups was genotyped twice, with 100% reproducibility.
Assessment of human serum levels of FOXP3 and IL-28B
For quantitative determination of IL-28B and FOXP3 serum levels, the concentrations of circulating IL-28B (Cat.No E0692Hu) and FOXP3 (Cat.No E4773Hu) in serum were determined using commercially available ELISA kits (Bioassay Technology Laboratory, BT LAB, England), according to the manufacturer’s instructions. The minimum level of detection for IL-28B and FOXP3 were 0.25 ng/L and 0.09 ng/ml, respectively. The developed color reaction was measured at OD 450 units on an ELISA reader (ELx 808, BIO-TEKInstruments, US).
Sample size calculation
Following the study of Zakaria et al. (2019); the percentage of TT homo-mutant genotype of IL-28B rs12979860 in HCV-positive cases is 13%. Assuming a population size of 500 HCV-infected individuals, a minimum sample size of 130 individuals is required with a margin of error of 0.05 and a 95% confidence interval. To compensate for the loss in follow-up; the sample size was increased by 15% to 150 cases. The sample size was estimated using the NQuery statistical package, version 7.0, Los Angeles, CA [35].
Statistical analysis
Once the data was collected and reviewed, it was coded and entered into the Statistical Package for Social Science (IBM SPSS) version 26. Parametric data were presented as mean and standard deviation, while non parametric data were presented as median with range. Qualitative variables were presented as frequency and percentages. Normality was examined with the application of the Kolmogorov–Smirnov test or Shapiro–Wilk test. Pearson’s Chi-square test was used to examine the relation between qualitative variables. A comparison of quantitative variables between the two groups was done using the Student t-test for normally distributed data. Mann–Whitney test or Kruskal–Wallis test were used for not normally distributed data. Correlation between numerical variables was tested using Spearman-rho correlation. Simple and multiple binary logistic regression was performed with the forward approach to adjust for significant covariates. The receiver operating characteristic curve (ROC) was used to examine the prediction ability of the model and presented as an area under the curve (AUC) with its 95% confidence interval (CI). The odds ratio (OR) with its 95% CI was used for risk assessment. All tests were two-tailed. A p-value < 0.05 was considered significant. All the genotype frequencies of the selected SNPs in ourstudy followed the Hardy–Weinberg equilibrium.
We used Haploview 4.1 software [36] to identify haplotype blocks by analyzing the linkage disequilibrium (LD) between polymorphisms. We employed the linkage disequilibrium coefficient (D’). The proportion of haplotypes in each group was compared usinga Chi-squared test.
Results
Clinical traits of enrolled subjects
In the present study, 162 participants infected with HCV-GT4 and treated with SOF/DAC combination were enrolled. Ninety-nine Egyptian participants (61.1%) were responders, and sixty-three participants (38.9%) were non-responders. Table 1 provides the summary statistics for the baseline characteristics of the studypopulation.The age mean of all participants was 46 ± 12 years. Of the response group, the mean age was 43.2 ± 12.9 years which was significantly less than the mean ageforthe non-responder group (50.1 ± 8.5, P Value < 0.001). The percentages of males and females in the non-responder group were comparable, while the percentage of females was significantly higher than males in the responder group (P Value = 0.01). Among the non-responder group, the levels of ALT, AFP, platelets count, and HCV quantitationwere statisticallysignificantlyhigher than the responder group, P values = 0.002, 0.001, 0.015, < 0.001, respectively.
Association of the IL-28B and FOXP3 polymorphisms with response to DAAs in HCV treated subjects
First, we investigated the association between four SNPs—IL-28B rs12979860, IL-28B rs8099917, FOXP3 rs3761548, and FOXP3 rs2232365—and treatment response in 162 HCV-infected subjects treated with (SOF/DAC) for 12 weeks.
Table 2 and Fig. S1 provide an overview of the frequencies of genotypes and alleles for the SNPs in responders and non-responders. For the responder and non-responder groups, each polymorphism was in Hardy–Weinberg equilibrium.
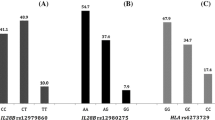
For IL-28B rs12979860 polymorphism, the CT frequency exhibited the highest frequency genotype in the study subjects; 53.5% in the responder group and 50.8% in non-responder patients. A comparable pattern was observed in the subjects of CC genotype where responders showed 2.06 times more sensitivity to SOF/DAC treatment compared to non-responders. In contrast, the TT genotype was significantly more resistant to treatment than other genotypes (P = 0.02). It was found that the risk allele “T” was the most common (57.14%) among the non-responder group and significantly higher as compared to the responder group (42.93%) with an OR of 1.77 (p = 0.013).
Meanwhile, for IL-28B rs8099917, the “T” allele frequency was the predominant allele between the studied subjects, with a frequency of 84.8% in responders and 84.1% in non-responders. Yet, the genotypes and allele distributions of this polymorphic variant did not differ significantly across the groups studied.
For the FOXP3 rs3761548 polymorphism, the “C” allele had the highest percentageamong the two groups, accounting for 52.02% of responders and 53.17% of non-responders. The genotype and allele frequencies of the variations in this polymorphism did not show statistically significant differences.
Furthermore, for FOXP3 rs2232365 polymorphism, a significant difference was identified in the different genotypes (P = 0.045). Table 2 presented that the “A” allele distribution frequencies were 62.12% and 76.19% among the responder and non-responder subjects, respectively. The distribution of FOXP3 rs2232365 genotypes in responders was as follows: AA and AG were the most predominant genotypes accounting for 41.41% of the population, while the GG genotype represented 17.17%. Contrarily, the frequencies of the AA, AG, and GG genotypes in the non-responder group were 60.32%, 31.75%, and 7.94%, respectively. Interestingly, responders exhibited a significantly higher prevalence of the risk allele "G", with an OR of 0.51 (P = 0.008) as compared to non-responders.
A haplotype block was allocated to the studied polymorphisms (Fig. 2). It was shown that among HCV patients, the CTAA haplotype predominated. The polymorphic variation in FOXP3 and IL-28B did not exhibit linkage disequilibrium. However, a weak linkage disequilibrium was observed between the two SNPs of IL-28B and FOXP3 (D′: 0.484, r2: 0.045/ D′:0.447, r2: 0.087; respectively). Table 3 represented the haplotype frequency for each block.
Linkage disequilibrium between genotyped SNPs. The number in each square represents the pairwise LD relationship (r2) between the IL28B (rs12979860, rs8099917) and FOXP3 (rs2232365, rs3761548) SNPs in Egyptian HCV patients treated with Daclatasvir (DAC) 60 mg/day and Sofosbuvir (SOF) 400 mg/day and varying red color represent the linkage disequilibrium values for that pair as measured by D′ (bright red shows high D′). The numbers in the gray-shaded area represent inferred listed in Table 3. The inferred in each haplotype block and the linkage between blocks are shown. The lines between haplotype blocks indicate the linkage frequency with a thick line for > 10% and a thin line for a frequency from 1–10%. Haplotype frequency within each haplotype block is shown next to each haplotype
Relationship between the genotypes of IL-28B and FOXP3 SNPs and their serum levels in response to DAA therapy
Next, to assess the relationship between serum IL-28B and FOXP3 levels and the various genotypes identified in the studied SNPs, serum samples from both groups were randomly selected for analysis. The results (Table 4), as displayed in Fig. 3A, indicate that there was a statistically significant difference in serum levels of IL-28B between patients who responded to DAA therapy and those who did not (P = 0.046). Remarkably, a 146.6% rise was detected in serum levels for the TT genotype of IL-28B rs12979860 among responders as compared to non-responders (Fig. 3B).
Comparison of (A) the serum level of IL28B between responder and non-responder subject; (B) IL28B (rs12979860) genotypes against average serum IL28B concentration; (C) the serum level of FOXP3 between responder and non-responder subjects and FOXP3 rs2232365 (D) and rs3761548 (E) genotypes against average serum FOXP3 concentration (F) Boxplot for the serum level of ALT among C and T Alleles of responders and non-responders a, b: Statistically Significant of AA and AG genotypes of FOXP3 rs2232365 in non-responder compared to GG genotype in responders using Kruskal-Wallis test followed by Dunn’s test for multiple comparison. c, d: Statistically Significant of AA and AC genotypes of FOXP3 rs3761548 in non-responder compared to AC genotype in responders using Kruskal-Wallis test followed by Dunn’s test for multiple comparison. e: Statistically Significant of TT genotype of IL28B rs12979860 in non-responder compared to TT genotype in responders using Kruskal-Wallis test followed by Dunn’s test for multiple comparison *Statistically Significant between C allele in responder and T allele in non-responder using Kruskal-Wallis test followed by Dunn’s test for multiple comparison
On the other hand, the serum level of FOXP3 was significantly decreased by 31% in the responder group compared to the non-responder group (P < 0.001) as shown in Fig. 3C. Post hoc analysis revealed that responders had the lowest concentration of FOXP3, with a median of 6.11 (3.68–12.89), in comparison to non-responders. This was particularly evident in subjects with the FOXP3 (rs2232365) GG genotype (Fig. 3D). In the same manner, the responder patients with the AC genotype of FOXP3 rs3761548 showed the lowest level of FOXP3 with a median of 7.91 (3.6–24.48) compared to other genotypes in non-responder patients (Fig. 3E).
The association between IL-28B and FOXP3 serum levels and different clinical characteristics in responsive and non-responsive subjects
Higher pretreatment ALT levels have been linked to a slower virological response [37]. Further analysis was conducted to evaluate the correlation between ALT levels and IL-28B rs12979860 alleles. Interestingly, the ALT levels were significantly lower in responders with the “C” allele compared to non-responders carrying the “T” allele (Fig. 3F).
Additionally, correlation analysis was conducted to assess the relationship between the serum levels of FOXP3 and IL-28B in terms of several clinical features as shown in Table S2. In the responder group, a noteworthy moderate positive correlation was observed (P = 0.001) between the serum levels of FOXP3 and IL-28B, with a correlation coefficient of 0.44. Furthermore, a weak positive correlation was found between the IL-28B serum level and HCV quantification (r = 0.35, P = 0.007), in addition to platelet count (r = 0.33, P = 0.013). No significant associations were found between the serum levels of IL-28B or FOXP3 and other clinical characteristics. Similarly, the results of the correlational analysis in the non-responder group showed that the serum levels of IL-28B or FOXP3 and other clinical characteristics were not associated. Otherwise, there was a moderate negative correlation (r = -0.48, P = 0.015) between the serum concentration of FOXP3 and ALT.
Univariate and multivariate logistic regression analysis with DAAs treatment response
For variables that showed statistically significant differences between the two groups, a univariate regression analysis was conducted. The purpose of this analysis was to investigate the independent (predictor) and dependent (outcome) variables associated with the response to DAAtherapy. As shown in Table 5, the TT genotype in IL-28B rs12979860 was identified to be a risk factor, increasing the possibility of DAA therapy failure by 3.4 times. Furthermore, this mutation remained a reliable predictor of the treatment response regardless of the patient’s gender or age. Conversely, there was nosignificant relationship disclosed between the serum IL-28B level and the treatment response.
On the contrary, the mutation in FOXP3 rs2232365 (GG) was found to be associated with the response to DAAs, making it a reliable predictor, with an OR of 0.327 and a statistically significant p-value of 0.049. Interestingly, there is a strong relationship between the serum level of FOXP3 and the response to DAAs with an OR of 1.29 (95% CI: 1.13–1.47) and a highly significant P- value of less than 0.001.
A comparable pattern was observed with HCV quantification, ALT concentration, AFP, and platelet count. These parameters significantly correlated with the responsiveness to DAA therapy, independent of the subject’s age or gender (Table 5).
All the relevant univariate variables described earlier were involved in the multivariate logistic regression analysis, adjusting for confounders, to identify the key predictors of treatment response to DAA therapy. Interestingly as shown in Table 6, the response to DAA therapy was shown to be correlated with a number of variables, including TT variation in IL-28B rs12979860, age, gender, ALT, and AFP serum levels, and virological baseline. The following formula was created to develop a prediction equation for estimating the probability (P) of response to DAA therapy in HCV-infected Egyptian patients:
P = 1/1 + e− (−7.408+0.050*age+0.0000005*HCV DNA+0.014*ALT+0.547*AFP+1.062*gender+1.096*IL−28B. rs12979860)
Where Male = 1, female = 0. IL-28B rs12979860 TT = 1, CC + CT = 0.
Receiver operating characteristic curve for the best cut-off point between responder and non-responder groups regarding gender, age, ALT, IL-28B rs12979860, AFP, and HCV quantification
The ROC curve was plotted and showed that our predictors have a reliable predictive capacity, with an area under the ROC curve of 0.917 (95% CI = 0.871- 0.963) (Fig. 4). Where the best cut-off value for this model was 5.00, this means that patients above this cut-off value were more likely to exhibit resistance to DAAs treatment. This model presented a sensitivity of 76.2%, specificity of 91.9%, and total accuracy of 85.8%.
Receiving operating curve (ROC) of the multivariate analysis for DAA treatment response in chronic HCV Egyptian patients treated with Daclatasvir (DAC) 60 mg/day and Sofosbuvir (SOF) 400 mg/day. Area under the curve (AUC) was 0.917 (95% CI = 0.871- 0.963). It is represented a sensitivity of 76.2%, specificity of 91.9%, and total accuracy of 85.8%
Discussion
The introduction of DAA therapy has changed the epidemiological impact of HCV infection management in the Middle East and North Africa (MENA) region [38]. However, HCV-infected patients exhibit varying responses to DAA therapy, and some may not derive any benefit from it [15]. Consequently, it is imperative to explore the factors that may influence the disease and treatment outcomes. Besides the cost burden of DAAs, predicting disease progression, effectively treating patients, and accurately assessing risks and benefits would pave the way toward personalized medicine for HCV patients. In the present study, it was shown that the response to DAA therapy is reduced in HCV Egyptian patients with IL28B rs12979860 T > C and FOXP3 rs2232365 A > G variants, as well as lower serum IL28B and higher serum FOXP3 levels.
Multiple studies have revealed a significant influence of IL-28B polymorphic variations on the treatment outcomes of chronic hepatitis C individuals [17, 39, 40]. Subsequently, several investigators have published similar observations in different populations at different positions of the IL-28B gene [26, 35, 41,42,43,44]. In particular to the MENA region, the IL-28B rs12979860 SNP was found to predict the consequence of HCV genotype 3a infection in the Iranian population, along with the IL-28B rs8099917 and IL-28B rs8103142 SNPs among HCV GT4-infected Egyptians [26, 35, 45]. A Tunisian study further confirmed the significance of IL-28B rs12979860 variation in predicting treatment response and toxicity [46].The same association results were reported among Caucasians, African Americans, and Hispanics [41, 43, 44].
In the era of oral interferon-free therapy,several studies have attempted to determine the impact of IL-28B SNPsin DAA treatment response [47]. Therefore, in this study, we explored the treatment outcome of two different SNPs within IL-28B among chronic HCV Egyptian patients receiving SOF/DAC combination therapy. In addition to IL-28B polymorphisms, we also expanded our investigation to revealvariations in the promoter region of the FOXP3 gene. It is worth noting that polymorphisms in the FOXP3 gene can affect the expression of the FOXP3 protein, leading tochanges in the normal activation of Treg cells [27]. Accordingly, it is crucial to conduct studies to assess the traits of immunogenetic interactionsrelated to disease in various contexts to advance our knowledge and pave the way for personalized medicine.
Recent studies have suggested that the IL-28B gene polymorphic variations significantly influence the treatment response of individuals infected with HCV who are receiving DAA therapy. IL-28B gene polymorphisms, specifically the SNPs rs12979860 and rs8099917 on chromosome 19, are predictiveof viral eradication after 12 weeks of therapy [19, 48, 49]. In the current study, we revealed that the IL-28B rs12979860 “T” allele was approximately twice as likely to be a predisposing factor for non-responders among Egyptian HCV patients. Patients with favorable response genotypes (CC or CT) are more susceptible to respond well to DAAs treatment compared to those with the resistant response genotype (TT). Additionally, we confirmed that the IL-28B rs12979860 “T” allele remains a reliable predictor of the responsiveness of DAA therapy, despite the patient’s gender or age. In agreement with our findings, the first case–control study concluded that IL-28B rs12979860 is an effective marker for predicting treatment response. The CT genotype had the highest response rate at 62.5%, followed by 30% in CC and 7.5% in TTgenotypes among Egyptian patients with GT4 [50]. However, their findings need to be confirmed, by further research, due to the small sample size and the use of only one DAAs regimen (SOF/RBV) in their study. Our observation is consistent with the results reported by El-Garawani in 2020 which showed a noteworthy rise in the percentage of the CT genotype of IL-28B rs12979860 among responders as compared to non-responders receiving SOF/DAC. Additionally, non-responders had a significantly higher frequency of TT genotypes (42.85%) compared to responders [20]. The same results were reported in 131 chronic HCV Egyptian patients with genotype 4 who received SOF and DAC with or without RBV therapy for 3 months [49].
Several researchers have examined the IL-28B rs8099917 polymorphism in chronic viral hepatitis, however, this study represents an initial exploration ofthe impact of this SNP within the Egyptian HCV population receiving the SOF 400 mg/day and DAC 60 mg/day regimen [16, 51]. In an Indian cohort study investigating the impactof IFNL3 in genotype 3 chronic HCV patients receiving SOF/DAC combination, they found that TT (major) genotype of IL-28B rs8099917 were linked to best SVR rates and GG (minor) genotype of IL-28B rs8099917 were more predominant in relapses or non-responses of this regimen [19]. In line with these results, a study conducted on chronic HCV GT4 Egyptian patients who were treated with a combination of PEG-IFN and oral RBV showed that the TT genotype of IL-28B rs8099917 was linked to achieving SVR [52]. Conversely, the present observation revealed that in HCV Egyptian patients, the SOF/DAC combination treatment response was not influenced by the mutation in IL-28B rs8099917. These results could potentially be attributed to variances in IL-28B gene polymorphisms and ethnicity where IL-28B polymorphisms accounted for just 50% of inter-individual variability in SVR that were attributed to ethnic differences and only 15% of HCV infection spontaneous clearance [17, 53]. In a recent meta-analysis, the probability of successful SVR was greater for individuals with the CC genotype of rs12979860 CC genotype compared to those with the TT genotype of rs8099917. The positive predictive value of the CC genotype of rs12979860 was 76% whereas that of the TT genotype of rs8099917was 71% [54]. Multiple pieces of evidence emphasized that the existence of the “G” variant allele on the IL-28B rs8099917 gene may be regarded as a risk allele for persistent HCV infection rather than predicting treatment response [19, 51]. Alternatively, a cohort study of Australian individuals with chronic HCV genotype 1 who were treated with PEG-IFNα/RBV combination therapy reported an association between SVR and the IL-28B rs8099917 genotypes [16]. The current study hypothesized that the contradiction in results could be attributed to the variations in HCV genotype and host ethnicity.
It is interesting to note that the ALT levels, which are known to follow treatment response, were also found to be correlated with the IL-28B genotypes. Patients with high inflammatory activity typically have elevated liver enzyme levels such as ALT and AST [55]. The present study found that patients with the favorable “C” allele for IL-28B rs12979860 had reduced ALT levels. This reduced level of ALT in our investigation has been accompaniedby a favorable response to DAA therapy. This results in compliance with a recent retrospective case–control analysis reporting that abnormal ALT after SVR following treatment with DAAs is rare [56]. In line with the current results, a recent study revealed that surpassing ALT or AST levels after DAA therapy may indicate treatment failure in mild liver disease [37].
Enduring HCV infection is manifested by weak and impaired cellular immune responses due to dysfunctional dendritic cells. These cells play a vital role in influencing the infection outcome by releasing IFNs and triggering the activation of T-lymphocytes [57,58,59]. Nevertheless, the mechanism of how polymorphisms in the region of IL-28B genes affect treatment responses has remained elusive. The present study hypothesized that as a polymorphism located 3 kb upstream of thepromoter region for the IL-28B gene, it may theoretically influence IFN-lambda genes.Therefore, the present study investigated the IL-28B serum level in responder and non-responder patients and showed a significant difference between the two groups. It is worth noting that patients who carried the TT genotype of IL-28B rs12979860 and still responded to the treatment exhibited elevated levels of IL-28B compared to those who remained non-responders.These findings supported the results reported by Langhans et al. (2011) who found a correlation between serum levels of IL-28B and the “C” allele of IL-28B rs12979860 SNP [60]. Additionally, two Iranian studies have quantified that the expression level of IL-28B mRNA increased in HCV-GT1 patients treated with PEG-IFNα /RBV compared to untreated patients. Additionally, the serum level of IL-28B in HCV-GT4 patients was higher compared to subjects who achieved immediate clearance [26, 61]. It is well established that IFN λ ultimately exerts its functions through the upregulation of interferon signaling genes, leading to the control and eradication of infection [24]. It is rationally accepted that patients who respond to DAA combination therapy typically have higher concentrations of IL-28B in their serum [62]. Accordingly, our findings suggest that low IL-28B levels may predispose patients to resist DAAs treatment.
In chronic infections, continuous stimulation by the antigen contributes toexhaustion and dysfunction of T cells, which results in a lessened immune response against the virus [63]. In this scenario, Regulatory T cells (Treg) are the key regulator in sustaining the infection by easing the viral antigens to evade the immune system through the suppression of antiviral responses. FOXP3, a transcription factor for Treg cells, is a crucial marker for the development and function of Treg cells [27]. Consequently, the current study aimed to investigate the impact of SNPs in the promoter region of the FOXP3 gene on FOXP3 expression levels, which in turn influence Treg cell activation [64, 65]. There is evidence to suggest that the FOXP3 gene’s ‘‘G’’ allele increases the risk of developing an autoimmune disease. This allele is linked to reduced FOXP3 expression, which in turn leads to an imbalance in the immune system [66, 67]. In the present study, we investigated two SNPs, rs2232365 and rs3761548, in the promoter of the FOXP3 gene. We found that the “G” allele in FOXP3 rs2232365 is considered to have a protective effect in HCV patients who respond to DAAs. On the contrary, the presence of the FOXP3 rs3761548 polymorphism did not influence the response to DAAs in the studied groups. Strikingly, the individuals who did not respond to DAA therapy showed significantly elevated levels of FOXP3 in their serum compared to those who responded to the therapy. In addition, the serum levels of FOXP3 for the rs2232365 AG and AA genotypes along with the rs3761548 AC and CC genotypes elevated in the non-responder individuals. It is implicated that the increased levels of FOXP3 may initiate a robust hepatic regulatory response potentially leading to a lessening in the immune response against the virus. Consequently, this might ultimately lead to viral reactivation and the continuation of chronic infection [68, 69].
As previously reported, SOF and DAC inhibit HCV replication by targeting RNA polymerase NS5B and NS5A non-structural protein, respectively [70]. Accordingly, FOXP3 regulates Treg cell function and may suppress antiviral immune responses; moreover, its increased expression in some chronic HCV patients may contribute to a reduced ability to control the viral infection and resistance to DAAs treatment [71]. Limited research has explored the factors that regulate Treg cells in contexts other than autoimmune diseases. One study suggested that FOXP3 SNPs (rs2232365, rs3761549, and rs3761548) may have an immunomodulatory influence in patients with chronic viral hepatitis. The results indicate that variations in the FOXP3 gene may play a role in histopathological sequelae observed in the liver following viral infection. Furthermore, elevated levels of liver enzymes and a reduction in viral count were detected in subjects with the GG genotype of rs2232365 FOXP3 [33]. In a recent study, the contribution of Treg cells in the inflammation induced by viral hepatitis has revealed a noteworthy implication of the FOXP3 SNP rs3761547 T > C. It has displayed a significant influence on the progression of hepatic fibrosis induced by viral hepatitis [32]. An additional investigation reported that the elevation of FOXP3 expression is linked to the polymorphic variant A allele in FOXP3 rs2232365 which potentially leads to an increase in the risk of tuberculosis [72]. This may explain the observed relationship between FOXP3 serum levels and the DAAs treatment responsiveness, which could be clinically significant for managing HCV patients. Further research is needed to confirm these findings.
The molecular interaction between IL-28B and FOXP3 in HCV in the era of DAAs therapy is still under investigation. Nevertheless, IL-28B has been anticipated to decrease the appearance of CD4+CD25+FOXP3 +Treg cells during DNA vaccination, thereby modulating adaptive immune responses [73]. Additionally, in murine models, IL-28B treatment has been shown to reduce Treg cells and improve antitumor immunity [73]. In the current study, we investigated a weak association between levels of IL-28B and FOXP3 among HCV patients. Further research is required to explore the effect of DAAs on these variables and the relationship between IL-28B and FOXP3 in HCV infection.
The current prediction model observed that being male, increased age, high levels of ALT, higher AFP, a high HCV viral load, and the presence of the T risk allele of IL-28B rs12979860 were associated with DAAs resistance in multivariate logistic regression analysis, with 76.2% sensitivity and 91.9% specificity. In other words, the ROC curve analysis demonstrated the utility of the IL-28B rs12979860 polymorphism, with a cut-off value of 5, as a predictor of the responsiveness to DAAs therapy in HCV-infected Egyptian patients.
Conclusion
The present study concludes that rs12979860 C˃T IL-28B and rs2232365 G˃A FOXP3 polymorphisms are regarded as favorable alleles determining the response to SOF/DAC therapy for HCV in Egyptian patients. Furthermore, we have revealed for the first time that the response to DAA combination therapy is correlated with the serum level of IL-28B and FOXP3. Hence,they can be utilized as a predictive measure to assessthe response to therapy. Until an approved HCV vaccine is developed, it is crucial to continue comprehensive research efforts aimed at reducing the risks associated with HCV treatment resistance. Therefore, our study enhances our understanding of the relationship between genotype and phenotype in the variability of response to DAAs among HCV patients.
It is important to consider that the current study was conducted in an Egyptian HCV-infected population receiving one regimen of DAAs. Hence, future research is needed to assess the generalizability of our findings in different contexts. Also, the present study is subject to limitations primarily due to the investigation of only two SNPs among the genes attributed to Treg functions. Regarding the complexities observed in immune responses, it is recommended that other SNPs of this gene be investigated to clarify the interactions between the pathogen and the host.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
World Health Organization: WHO [Internet]. 2023 [cited 2023 Aug 28]. Hepatitis C. Available from: https://www.who.int/news-room/fact-sheets/detail/hepatitis-c
Manns MP, Maasoumy B, Rice CM. Breakthroughs in hepatitis C research: from discovery to cure. Nat Rev Gastroenterol Hepatol. 2022;19:533–50.
Frank C, Mohamed MK, Strickland GT, Lavanchy D, Arthur RR, Magder LS, et al. The role of parenteral antischistosomal therapy in the spread of hepatitis C virus in Egypt. Lancet. 2000;355(9207):887–91.
Esmat G, El-Sayed MH, Hassany M, Doss W, Waked I. One step closer to elimination of hepatitis C in Egypt. Vol. 3, The Lancet Gastroenterology and Hepatology. Elsevier Ltd; 2018. p. 665.
Hajarizadeh B, Grebely J, Dore GJ. Epidemiology and natural history of HCV infection. Nat Rev Gastroenterol Hepatol. 2013;10(9):553–62.
Borgia SM, Hedskog C, Parhy B, Hyland RH, Stamm LM, Brainard DM, et al. Identification of a novel hepatitis C virus genotype from Punjab, India: expanding classification of hepatitis C virus into 8 genotypes. J Infect Dis. 2018;218(11):1722–9.
Wantuck JM, Ahmed A, Nguyen MH. Review article: The epidemiology and therapy of chronic hepatitis C genotypes 4, 5 and 6. Aliment Pharmacol Ther. 2014;39(2):137–47.
Abdel-Samiee M, Youssef MI, Elghamry F, Bazeed M, Al-Shorbagy M, Shalaby H, et al. A multicentric and nationwide predictive study role of T cell sub-population in the prevalence and prognosis of cryoglobulinemia among genotype 4 chronic hepatitis C patients. J Med Virol. 2023;95(12):e29248.
Götte M, Feld JJ. Direct-acting antiviral agents for hepatitis C: structural and mechanistic insights. Nat Rev Gastroenterol Hepatol. 2016;13(6):338–51. https://doi.org/10.1038/nrgastro.2016.60.
Abozeid M, Alsebaey A, Abdelsameea E, Othman W, Elhelbawy M, Rgab A, et al. High efficacy of generic and brand direct acting antivirals in treatment of chronic hepatitis C. Int J Infect Dis IJID Off Publ Int Soc Infect Dis. 2018;75:109–14.
Liver EA for the S of the. EASL recommendations on treatment of hepatitis C: Final update of the series(☆). Vol. 73, Journal of hepatology. Netherlands; 2020. p. 1170–218.
Hoofnagle JH, Sherker AH. Therapy for hepatitis C — The costs of success. N Engl J Med. 2014;370(16):1552–3.
Essa M, Sabry A, Abdelsameea E, Tharwa ES, Salama M. Impact of new direct-acting antiviral drugs on hepatitis C virus-related decompensated liver cirrhosis. Eur J Gastroenterol Hepatol. 2019;31(1):53–8.
Schwander B, Feldstein J, Sulo S, Gonzalez L. Pursuing elimination of hepatitis C in Egypt : cost- effectiveness and economic evaluation of a country- wide program. Infect Dis Ther. 2022;11(3):1193–203. https://doi.org/10.1007/s40121-022-00631-x.
Nahon P, Cobat A. Human genetics of HCV infection phenotypes in the era of direct-acting antivirals. Hum Genet. 2020;139(6–7):855–63. https://doi.org/10.1007/s00439-020-02136-4.
Suppiah V, Moldovan M, Ahlenstiel G, Berg T, Weltman M, Abate ML, et al. IL28B is associated with response to chronic hepatitis C interferon-α and ribavirin therapy. Nat Genet. 2009;41(10):1100–4.
Ge D, Fellay J, Thompson AJ, Simon JS, Shianna KV, Urban TJ, et al. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature. 2009;461(7262):399–401.
America N, Tanaka Y, Nishida N, Sugiyama M, Kurosaki M, Matsuura K, et al. Genome-wide association of IL28B with response to pegylated interferon- α and ribavirin therapy for chronic hepatitis C. Nat Genet. 2009;41(10):1105–9.
Khan AJ, Saraswat VA, Ranjan P, Parmar D, Negi TS, Mohindra S. Polymorphism in interferon λ3/interleukin-28B gene and risk to noncirrhotic chronic hepatitis C genotype 3 virus infection and its effect on the response to combined daclatasvir and sofosbuvir therapy. J Med Virol. 2019;91(4):659–67. https://doi.org/10.1002/jmv.25359.
El-Garawani I, Hassab El-Nabi S, Gadallah M, Abdelsameea E. Association between IFN-λ 3 gene polymorphisms and outcome of treatment with direct acting antivirals in chronic HCV-infected Egyptian psatients. Immunol Invest. 2020;50(1):1–11. https://doi.org/10.1080/08820139.2020.1722158.
Ghanem SE, Elsabaawy M, shebl N, Abdelsameea E, Othman W, EL-Bassal FI, et al. Value of IFNL3 genetic polymorphism in the prediction of HCV treatment response to direct-acting antiviral drugs versus interferon therapy. Expert Rev Anti Infect Ther. 2020;18(9):947–54.
Kotenko SV, Gallagher G, Baurin VV, Lewis-Antes A, Shen M, Shah NK, et al. IFN-λs mediate antiviral protection through a distinct class II cytokine receptor complex. Nat Immunol. 2003;4:69–77.
Sheppard P, Kindsvogel W, Xu W, Henderson K, Schlutsmeyer S, Whitmore TE, et al. IL-28, IL-29 and their class II cytokine receptor IL-28R. Nat Immunol. 2003;4:63–8.
Wack A, Terczyn E, Hartmann R. Guarding the frontiers : the biology of type III interferons. Nat Immunol. 2015;16(8):802–9.
Al-Qahtani A, Al-Anazi M, Abdo AA, Sanai FM, Al-Hamoudi W, Alswat KA, et al. Correlation between genetic variations and serum level of interleukin 28B with virus genotypes and disease progression in chronic hepatitis C virus infection. J Immunol Res. 2015;2015:768470.
Moghimi M, Tavakoli F, Doosti M, Ahmadi-Vasmehjani A, Akhondi-Meybodi M. Correlation between interleukin-28 gene polymorphism with interleukin-28 cytokine levels and viral genotypes among HCV patients in Yazd. Iran BMC Res Notes. 2019;12(1):10–4. https://doi.org/10.1186/s13104-019-4651-z.
Oda JMM, Hirata BKB, Guembarovski RL, Watanabe MAE. Genetic polymorphism in FOXP3 gene: Imbalance in regulatory T-cell role and development of human diseases. J Genet. 2013;92(1):163–71.
Wu KJ, Qian QF, Zhou JR, Sun DL, Duan YF, Zhu X, Lu YJ. Regulatory T cells (Tregs) in liver fibrosis. Cell death discovery. 2023;9(1):53.
Belkaid Y, Rouse BT. Natural regulatory T cells in infectious disease. Nat Immunol. 2005;6:353–60.
Zhao H, Liao X, Kang Y. Tregs: Where we are and what comes next? Front Immunol. 2017;8:1578.
Hoogendoorn B, Coleman SL, Guy CA, Smith K, Bowen T, Buckland PR, et al. Functional analysis of human promoter polymorphisms. Hum Mol Genet. 2003;12(18):2249–54.
Kao HH, Yu RL, Chuang WL, Huang JF, Dai CY, Tan CH. Genetic polymorphisms of regulatory T cell-related genes modulate systemic inflammation induced by viral hepatitis. Kaohsiung J Med Sci. 2021;37(11):1000–9.
Pereira LMS, da Amoras E, SG, Conde SRS d. S, Demachki S, Monteiro JC, Martins-Feitosa RN, et al. The -3279C > A and -924A > G polymorphisms in the FOXP3 gene are associated with viral load and liver enzyme levels in patients with chronic viral liver diseases. Front Immunol. 2018;9:1–16.
Weng CL, Yazid H, Appalasamy S, Geng BJ, Nasir WMNWM, Muhammad NMN, et al. Optimization of binding washing and elution buffer for development of DNA isolation kit. IOP Conf Ser Earth Environ Sci. 2020;596(1):012008.
Zakaria ZA, Knapp S, Hashem M, Zaghla H, Thursz M, Waked I, et al. Interleukin 28A rs12980602 and interleukin 28B rs8103142 genotypes could be protective against HCV infection among Egyptians. Immunol Res. 2019;67(1):123–33.
Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21(2):263–5.
Flower B, Nguyen Thi Ngoc P, McCabe L, Le Ngoc C, Vo Thi T, Thi Kim HV, et al. Rise in alanine aminotransferase after HCV treatment is a highly sensitive screen for treatment failure. Clin Liver Dis. 2023;21(5):138–42.
Ayoub HH, Mahmud S, Chemaitelly H, Abu-Raddad LJ. Treatment as prevention for hepatitis C virus in the middle East and North Africa: a modeling study. Front Public Heal. 2023;11:1–13.
Thomas DL, Thio CL, Martin MP, Qi Y, Ge D, Ohuigin C, et al. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature. 2009;461(7265):798–801.
Tanaka Y, Nishida N, Sugiyama M, Kurosaki M, Matsuura K, Sakamoto N, et al. Genome-wide association of IL28B with response to pegylated interferon-α and ribavirin therapy for chronic hepatitis C. Nat Genet. 2009;41(10):1105–9. https://doi.org/10.1038/ng.449.
Thompson AJ, Muir AJ, Sulkowski MS, Ge D, Fellay J, Shianna KV, et al. Interleukin-28B polymorphism improves viral kinetics and is the strongest pretreatment predictor of sustained virologic response in genotype 1 hepatitis C virus. Gastroenterology. 2010;139(1):120–9.
Abe H, Ochi H, Maekawa T, Hayes CN, Tsuge M, Miki D, et al. Common variation of IL28 affects gamma-GTP levels and inflammation of the liver in chronically infected hepatitis C virus patients. J Hepatol. 2010;53(3):439–43. https://doi.org/10.1016/j.jhep.2010.03.022.
Montes-Cano MA, García-Lozano JR, Abad-Molina C, Romero-Gómez M, Barroso N, Aguilar-Reina J, et al. Interleukin-28B genetic variants and hepatitis virus infection by different viral genotypes. Hepatology. 2010;52(1):33–7.
Mangia A, Thompson AJ, Santoro R, Piazzolla V, Tillmann HL, Patel K, et al. An IL28B polymorphism determines treatment response of hepatitis C virus genotype 2 or 3 patients who do not achieve a rapid virologic response. Gastroenterology. 2010;139(3):821–7.
Salum GM, Dawood RM, Abdel-Meguid M, Ibrahim NE, Aziz AOA, Awady MKEl. Correlation between IL28B/TLR4 genetic variants and HCC development with/without DAAs treatment in chronic HCV patients. Genes Dis. 2020;7(3):392–400.
Sghaier I, Mouelhi L, Gazouani E, Morel V, Besma YL, Brochot E. Roles of ITPA and IL28B genotypes in chronic hepatitis C patients treated with peginterferon plus ribavirin in Tunisian population. J Clin Virol. 2015;69:184–9.
Stättermayer AF, Ferenci P. Effect of IL28B genotype on hepatitis B and C virus infection. Curr Opin Virol. 2015;1(14):50–5.
Zaki SM, Ahmed HS, Yousif MM, Awad EM. Interleukin 28B polymorphism as a predictor of sustained virological response to sofosbuvir-based therapy for hepatitis C virus patients. Trop Med Infect Dis. 2022;7(9):230.
El- KSH, El- SM, Zakarya ZM, Helal GK. Association between interleukin 28B polymorphism and sustained virological response to sofosbuvir plus daclatasvir in chronic hepatitis C genotype 4 Egyptian patients. J Clin Pharm Ther. 2021;46:942–9.
Zidan HE, Talaat RM, Ammar AAA, Sakr MA. Interleukin 28B polymorphism as a predictor of response to treatment of Egyptian HCV patients working in nuclear material authority. Egypt J Hosp Med. 2019;77(1):4742–7.
Kuralay Z, Tuğ E, Fidan I. Investigation of the relationship between IL28B polymorphisms and plasma IL28B levels in patients with chronic hepatitis B or C. Mikrobiyol Bul. 2021;55(3):374–88.
Ragheb MM, Nemr NA, Kishk RM, Mandour MF, Abdou MM, Matsuura K, et al. Strong prediction of virological response to combination therapy by IL28B gene variants rs12979860 and rs8099917 in chronic hepatitis C genotype 4. Liver Int. 2014;34(6):890–5.
Duggal P, Thio CL, Wojcik GL, Goedert JJ, Mangia A, Latanich R, et al. Genome-wide association study of spontaneous resolution of hepatitis C virus infection: data from multiple cohorts. Ann Intern Med. 2013;158(4):235–45.
El-Fattah MA. Predictive power of Interleukin–28B gene variants for outcome of Hepatitis C virus genotype 4 in Egyptians: a systematic review and meta-analysis. Clin Res Hepatol Gastroenterol. 2021;45(2):101480. https://doi.org/10.1016/j.clinre.2020.06.006.
Akuta N, Suzuki F, Sezaki H, Suzuki Y, Hosaka T, Someya T, et al. Predictive factors of virological non-response to interferon-ribavirin combination therapy for patients infected with hepatitis C virus of genotype1b and high viral load. J Med Virol. 2006;78(1):83–90.
Chadha N, Turner A, Sterling RK. Prevalence and predictors of abnormal alanine aminotransferase in patients with HCV who have achieved SVR. J Viral Hepat. 2023;30(1):73–8.
Pachiadakis I, Pollara G, Chain BM, Naoumov NV. Is hepatitis C virus infection of dendritic cells a mechanism facilitating viral persistence? Lancet Infectious Diseases. 2005;5:296–304.
Rehermann B. Hepatitis C virus versus innate and adaptive immune responses: a tale of coevolution and coexistence. J Clin Invest. 2009;119(7):1745–54.
Longman RS, Talal AH, Jacobson IM, Albert ML, Rice CM. Presence of functional dendritic cells in patients chronically infected with hepatitis C virus. Blood. 2004;103(3):1026–9.
Langhans B, Kupfer B, Braunschweiger I, Arndt S, Schulte W, Nischalke HD, et al. Interferon-lambda serum levels in hepatitis C. J Hepatol. 2011;54(5):859–65. https://doi.org/10.1016/j.jhep.2010.08.020.
Nasab SDM, Vasmehjani AA, Kaghazian H, Mardani R, Zali F, Ahmadi N, et al. Association of IL28B (IFNL3) rs12979860 mRNA levels, viral load, and liver function among HCV genotype 1a patients. Gastroenterol Hepatol from Bed to Bench. 2018;2019(12):S156–62.
Alborzi A, Hashempour T, Moayedi J, Musavi Z, Pouladfar G, Merat S. Role of serum level and genetic variation of IL-28B in interferon responsiveness and advanced liver disease in chronic hepatitis C patients. Med Microbiol Immunol. 2017;206(2):165–74.
Wherry EJ, Blattman JN, Murali-Krishna K, van der Most R, Ahmed R. Viral persistence alters CD8 T-Cell immunodominance and tissue distribution and results in distinct stages of functional impairment. J Virol. 2003;77(8):4911–27.
Kalantar K, Khansalar S, Eshkevar Vakili M, Ghasemi D, Dabbaghmanesh MH, Amirghofran Z. Association of Foxp3 gene variants with risk of Hashimoto’s thyroiditis and correlation with anti-tpo antibody levels. Acta Endocrinol (Copenh). 2019;15(4):423–9.
An I, Harman M, Ibiloglu I. Topical ciclopiroxoOlamine 1%: revisiting a unique antifungal. Indian Dermatol Online J. 2017;10(4):481–5.
Song P, Wang XW, Li HX, Li K, Liu L, Wei C, et al. Association between FOXP3 polymorphisms and vitiligo in a Han Chinese population. Br J Dermatol. 2013;169(3):571–8. https://doi.org/10.1111/bjd.12377.
Claassen MAA, De KRJ, Tilanus HW, Janssen HLA, Boonstra A. Abundant numbers of regulatory T cells localize to the liver of chronic hepatitis C infected patients and limit the extent of fibrosis q. J Hepatol. 2010;52(3):315–21. https://doi.org/10.1016/j.jhep.2009.12.013.
Ward SM, Fox BC, Brown PJ, Worthington J, Fox SB, Chapman RW, et al. Quantification and localisation of FOXP3+ T lymphocytes and relation to hepatic inflammation during chronic HCV infection. J Hepatol. 2007;47(3):316–24.
Sulkowski MS, Gardiner DF, Rodriguez-Torres M, Reddy KR, Hassanein T, Jacobson I, et al. Daclatasvir plus sofosbuvir for previously treated or untreated chronic HCV infection. N Engl J Med. 2014;370(3):211–21.
Pereira LMS, Gomes STM, Ishak R, Vallinoto ACR. Regulatory T cell and forkhead box protein 3 as modulators of immune homeostasis. Front Immunol. 2017;8:1–24.
Beiranvand E, Abediankenari S, Khani S, Hosseini HM, Zeinali S, Beiranvand B, et al. G allele at -924 A > G position of FoxP3 gene promoter as a risk factor for tuberculosis. BMC Infect Dis. 2017;17(1):4–9.
Morrow MP, Pankhong P, Laddy DJ, Schoenly KA, Yan J, Cisper N, et al. Comparative ability of IL-12 and IL-28B to regulate treg populations and enhance adaptive cellular immunity. Blood, J Am Soc Hematol. 2009;113(23):5868–77.
Cd CD, Regulatory F, In TC, Chen X, Zhu B, Luo ÞY, et al. Interleukin-28B plays a therapeutic role on mouse U14 cervical cancer cells by down-regulating. Int J Gynecol Cancer. 2015;25(8):1369–76.
Acknowledgements
The authors would like to acknowledge Prof. Manar Mohamed Moneer, professor in Cancer Epidemiology and biostatistics at National Cancer Institute, Cairo University for statistical analysis of univariate and multivariate logistic regression and correlation analysis.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
Conceptualization and study design:AA, EA, SW, DE, ZZ and SA.
Patients enrollments, resources and data curation: MA.
Clinical data collection, Blood and serum collection: SG.
Follow up of clinical data curation and samples’ collection: EA and MA.
PCR and ELISA samples’ analysis: AA and ZZ.
Data Statistical analysis: AA and ZZ.
Data presentation and drafting of manuscript: AA and SA.
Data interpretation: AA, EA, SW, DE, ZZ and SA.
Revision and approval of the final manuscript: AA, EA, MA, SG, SW, DE, ZZ and SA.
Corresponding author
Ethics declarations
Conflict of interest
The authors have read the journal’s policy on disclosure of potential conflicts of interest, and they all wish to confirm that there are no known conflicts of interest associated with this publication and there has been no significant financial support for this work that could have influenced its outcome.
Consent for publication
All authors have read the journal’s authorship statement and agree to it. By your journal’s policy, we confirm that the material contained in the manuscript is original and has not been published and is not being submitted elsewhere. The authors qualify for authorship and have no financial or personal relationships that might lead to a conflict of interest. This manuscript is being submitted online in accordance with your policies for this type of submission.
The Authors hereby grant the Journal full and exclusive rights to the manuscript, all revisions, and the full copyright. The Journal rights include but are not limited to the following: (1) to reproduce, publish, sell, and distribute copies of the manuscript, selections of the manuscript, and translations and other derivative works based upon the manuscript, in print, audio-visual, electronic, or by any media now or hereafter known or devised; (2) to license reprints of the manuscript to third persons for educational photocopying; (3) to license others to create abstracts of the manuscript and to index the manuscript; (4) to license secondary publishers to reproduce the manuscript in print, microform, or any computer-readable form, including electronic on-line databases; and (5) to license the manuscript for document delivery. These exclusive rights run the full term of the copyright, and all renewals and extensions thereof.
Ethics approval
Ethical approvals are provided with submission documents.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abdelaziz, A.I., Abdelsameea, E., Abdel-Samiee, M. et al. Effect of immunogenetics polymorphism and expression on direct-acting antiviral drug response in chronic hepatitis C. Clin Exp Med 24, 184 (2024). https://doi.org/10.1007/s10238-024-01432-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10238-024-01432-x