Abstract
Isolated oceanic islands constitute interesting model systems for the study of colonization processes, as several climatic and oceanographic phenomena have played an important role in the history of the marine ichthyofauna. The present study describes the presence of two morphotypes of Caranx lugubris, in the St. Peter and St. Paul Archipelago located in the mid-Atlantic. Morphotypes were compared in regard to their morphological and cytogenetic patterns, using C-banding, Ag-NORs, staining with CMA3/DAPI fluorochromes and chromosome mapping by dual-color FISH analysis with 5S rDNA and 18S rDNA probes. We found differences in chromosome patterns and marked divergence in body patterns which suggest that different populations of the Atlantic or other provinces can be found in the Archipelago of St. Peter and St. Paul.
Similar content being viewed by others
Introduction
Ichthyofauna on the St. Peter and St. Paul Archipelago (SPSPA) is of great biological interest, due to its degree of geographic isolation. The region is a remote point, far from the South American (≈1,100 km) and African (≈1,824 km) continents, with a high level of endemic fish species (Edwards and Lubbock 1983). This small archipelago is made up of four larger islands (Belmonte, St. Paul, St. Peter and Barão de Teffé), in addition to 11 smaller rocky points. The combined action of the South Equatorial Current and Pacific Equatorial Undercurrent provides a highly complex hydrological pattern that significantly influences the insular ecosystem (Becker 2001). These components ultimately determine a common ichthyofauna between Brazil, Africa and the Caribbean as a result of larval dispersion, colonization and settlement (Feitoza et al. 2003). In fact, fish fauna from SPSPA represents a small set of ichthyofauna from Brazil, with some contribution from Ascension Island (≈1,940 km) and East Africa (Edwards and Lubbock 1983).
Physical factors such as ocean currents, temperature and salinity contribute to the definition of biogeographic and ecological limits through training, maintenance and distribution of the fauna in particular (Molina 2007). In many cases, environmental changes may cause the capacity of a single species to generate a phenotypic response immediately play a key role in promoting different phenotypes (polyphenism) within populations and subsequently conducting genetic disruption between them (Pfennig et al. 2010). However, the vast aquatic systems complicate the estimation of biological parameters and the absence of obvious barriers seems to indicate marine ecosystems resilient to facilitate gene flow between extensive marine populations, which can result in darkening of the influence of historical factors of species and their non-detection shallow levels of divergence (Palumbi and Metz 1991). It is almost a rule that species have wide distribution morphological and/or behavioral problems as a first step in the process of speciation (Molina 2007; Rocha et al. 2005).
In recent decades, several ichthyofaunal surveys have been conducted on SPSPA (e.g., Lubbock and Edwards 1981, Feitoza et al. 2003; Vaske et al. 2005). Among species considered pelagic, Caranx lugubris (Perciformes: Carangidae; Poey 1860), the black trevally, which exhibits circumtropical distribution, is one of the most abundant species and typically forms large schools (Feitoza et al. 2003). The C. lugubris in this ocean region is considered unique and taxonomically cohesive. To date, there are no known descriptions of inter- or intrapopulational variations in body pattern for this species in the Atlantic. However, oral accounts from artisan fishermen working in this region suggest two different morphotypes coexisting in the waters of the St. Peter and St. Paul Archipelago.
The present study identifies and characterizes two morphotypes of the black trevally collected in sympatry during expeditions on SPSPA in August and October 2009. Cytogenetic aspects were analyzed by conventional Giemsa staining, identification of Ag-NORs sites, C-banding, staining with CMA3/DAPI fluorochromes and chromosome mapping using dual-color FISH with 18S and 5S rDNA probes. In addition, morphotypes were also compared with regard to body proportions through geometric morphometrics. The presence of two black trevally morphotypes in the mid-Atlantic represents a peculiar and previously unidentified condition, with important ecological, genetic and biogeographical implications.
Materials and methods
Caranx lugubris specimens were collected from areas surrounding the SPSPA islands (00°55′02″N; 29°20′42″W) (Fig. 1). There were notable differences among individuals with regard to body shape, primarily in relation to eye size and body height. Given the apparently distinct morphological patterns, specimens were denominated morphotype I and II and were henceforth analyzed separately. The sex of all individuals was defined based on macroscopic gonadal examination.
Morphometric and meristic analyses
Geometric morphometrics was applied to analyze the body patterns of 65 C. lugubris specimens (Fig. 2) from SPSPA, with sizes varying between 25.5 and 44.0 cm. Individuals were preliminarily classified as morphotype I (N = 37) and morphotype II (N = 28) through visual examination.
Schematic representation of anatomical points for the 12 landmarks a defined for morphometric analysis in C. lugubris; (1) tip of the mouth; (2) indentation of the front profile above the snout; (3) insertion of the first dorsal fin; (4) insertion of the second dorsal fin; (5) base of the last dorsal fin ray; (6) end of the anal fin; (7) insertion of the soft anal fin; (8) insertion of the first spine of anal fin; (9) base of the pelvic fin; (10) insertion of the pectoral fin; (11); posterior end of the ocular orbit; (12) anterior end of the ocular orbit. Thin-plate spline representation of significant differences between C. lugubris morphotypes I and II in regard to the combined mean of samples. b Morphotypes I and II of C. lugubris; bar = 5 cm. c Distribution of morphological canonical scores grouped for both morphotypes. d Individuals with negative and positive canonical scores represent morphotypes I and II, respectively. Grouping resulted in 100 % correct classifications
Specimens were photographed individually from a right lateral view with an 8.1 megapixel Sony DSC-H10 digital camera. Twelve landmarks were defined for comparative analysis of morphotypes (Fig. 2a) using tpsDig2 software, version 1.40 (Rohlf 2004). Landmarks were selected so as to provide adequate coverage of the body’s lateral profile. Body landmarks were superimposed based on generalized procrustes superimposition (Rohlf and Slice 1990; Dryden and Mardia 1998).
Discriminant function analysis was applied to identify possible effects of sexual dimorphism in each of the morphotypes and between them, providing information on the extent of morphological divergence. Hotelling’s T2 distribution and permutation testing (1,000 times) were used to evaluate the significance of intergroup differences. Since the quality of discriminant analysis can be influenced by sample size, cross-validation was performed where each individual is sequentially removed from a group and classified according to the discriminant function derived from the remaining data.
Morphological differences between morphotypes I and II were observed by thin-plate spline with wireframe plots. In order to assess the degree of similarity between C. lugubris morphotypes and some other species of the genus, such as Caranx and Carangoides, body comparisons were developed using geometric morphometrics for C. latus (N = 14), C. hippos (N = 7), C. lugubris (N = 21), Carangoides crysos (N = 8) and Carangoides bartholomaei (N = 6), based on canonical variate analysis. Data were grouped according to Mahalanobis distances in the form of a dendrogram, using the UPGMA cluster analysis. All analyses were conducted with MorphoJ 1.02 software (Klingenberg 2008). Meristic values of serial elements dorsal and anal fins were obtained from a sample of individuals each C. lugubris morphotype. Meristic data for comparison with other Carangidae (Table 1) were based on Honebrink (2000) and Smith-Vaniz and Carpenter (2007).
Cytogenetic analysis
Cytogenetic comparison of C. lugubris morphotypes from SPSPA was conducted on a sample of 17 individuals, eight from morphotype I and nine belonging to morphotype II.
Prior to chromosomal preparations, specimens were submitted to in vivo mitotic stimulation for 24 h by intramuscular and intraperitoneal inoculation of bacterial and fungal antigens (Molina 2001; Molina et al. 2010), then anesthetized with clove oil (Eugenol) and sacrificed to remove kidney tissue. Mitotic chromosomes were obtained from cell suspensions of anterior kidney fragments through short duration in vivo methodology (Gold et al. 1990). Cell suspensions were dripped onto slides and recoated with a film of distilled water heated to 60 °C. Chromosomal preparations were stained with 5 % Giemsa diluted in a phosphate buffer pH 6.8. Approximately, 30 metaphases were analyzed for each individual to define the chromosome number. Metaphases were photographed on an Olympus BX50 epifluorescent microscope, with an Olympus DP70 digital image capturing system.
Chromosome banding
Heterochromatic regions and ribosomal sites were identified by the Sumner (1972) and Howell and Black (1980) techniques, respectively. CMA3/DAPI staining was applied in accordance with Barros-e-Silva and Guerra (2009).
Probes for chromosome hybridization
Two tandem-arrayed DNA sequences isolated from the Hoplias malabaricus (Teleostei, Characiformes) genome were used. The first probe contained a 5S rDNA repeat copy and included 120 base pairs (bp) of the 5S rRNA encoding gene and 200 bp of the non-transcribed spacer (NTS) (Martins et al. 2006). The second probe corresponded to a 1,400-bp segment of the 18S rRNA gene obtained via PCR from nuclear DNA (Cioffi et al. 2009). The 18S rDNA probe was labeled by nick translation with DIG-11-dUTP, according to manufacturer specifications (Roche). The 5S rDNA probe was labeled with biotin-14-dATP by nick translation, as per manufacturer specifications (Bionick Labelling System, Invitrogen).
Chromosome hybridization and analysis
Fluorescent in situ hybridization (FISH) was performed on mitotic chromosome spreads (Pinkel et al. 1986). Metaphase chromosome slides were incubated with RNAse (40 μg/ml) for 1.5 h at 37 °C. The chromosomal DNA was denatured in 70 % formamide/0.6× SSC for 3 min at 72 °C. After denaturation of chromosomal DNA in 70 % formamide, spreads were incubated in 2× SSC for 4 min at 70 °C. Hybridization mixtures containing 100 ng of denatured probe, 10 mg/ml dextran sulfate, 2× SSC and 50 % formamide/1× SSC in a final volume of 30 μl were dripped onto the slides, and hybridization was performed overnight at 37 °C in a 2× SSC moist chamber. Post-hybridization washes were carried out at 37 °C in 2× SSC, 50 % formamide for 15 min, followed by a second wash in 2× SSC for 15 min and a final wash at room temperature in 4× SSC for 15 min. Signal detection was performed using avidin-FITC (Sigma) for the 5S rDNA probe and antidigoxigenin-rhodamine (Roche) for 18S. Two-color 5S and 18S rDNA were detected by dual-color FISH, post-hybridization washes were performed on a shaker (150 rpm) and the chromosomes were then counterstained with a mixture of DAPI (1.2 μg/ml) and antifading Vectashield mounting medium solution (Vector Laboratories). FISH images were captured with an epifluorescence microscope (Olympus BX50) equipped with CoolSNAP system software, Image Pro Plus, (Media Cybernetics).
Results
Morphological characteristics
Discriminant analysis identified the absence of sexual dimorphism in each morphotype (T 2 = 32.3; p = 0.9), which could be due to morphological heterogeneity. However, individuals assigned as morphotypes I and II showed significant differences (T 2 = 340, p < 0.001), with 100 % of specimens correctly classified. In fact, the most apparent differences between morphotypes, observed via the deformation grid (Fig. 2b), are primarily related to eye diameter and body height.
Reliability of specimen allocation into their respective groups was determined by cross-validation. This analysis was performed on 65 individuals, with 34 of 37 assigned to morphotype I and 26 of 28 classified as morphotype II.
The dendrogram based on morphological patterns of C. lugubris and other species of Caranx and Carangoides, compiled from Mahalanobis distances using the UPGMA technique, showed that both morphotypes exhibited greater similarities between one another than in relation to other species analyzed (Fig. 3). Morphologically, the genera Caranx and Carangoides are perfectly discriminated into two clusters. One is composed of C. lugubris, C. latus and C. hippos, displaying greater body height, while the other, consisting of Carangoides, C. crysos and C. bartholomaei, have more elongated bodies.
Dendrogram of morphological similarity, obtained by UPGMA grouping, for species of Caranx and Carangoides. Morphotypes of C. lugubris show similarities among them, but are different from morphologically closer species such as C. hippos and C. latus. A second cluster groups C. crysos and C. bartholomaei, which display elongated bodies
Cytogenetic patterns
Morphotypes I and II displayed 2n = 48 chromosomes and karyotype composed of 6sm + 42a (FN = 54) (Fig. 4). Ag-NORs sites are located on the terminal portion of the short arm of chromosome pair 1. This region was heterochromatic and GC rich. C-banding also identified notable constitutive heterochromatin (CH) blocks in centromeric and telomeric regions of chromosomal pairs, with CH-rich pattern equilocal to NORs, CMA3/DAPI sequential staining also found visible CMA3+ markings in all the heterochromatic, centromeric and telomeric regions among karyotypes.
Karyotypes for morphotypes I (left) and II (right) of C. lugubris present in SPSPA. Giemsa staining, highlighting chromosome pair 1, bearer of Ag-NOR sites a C-banding. b DAPI/CMA3 staining showing GC-rich sites scattered over centromeric and telomeric regions of most chromosome pairs. c Dual-color FISH showing the mapping of 18S rDNA (red) and 5S rDNA (green) sites (color figure online)
Dual-color FISH with 18S and 5S rDNA probes demonstrated a non-syntenic condition for these ribosomal subunits. The 18S rDNA sites coincide with Ag-NOR markings, showing no variation in frequency or position between karyotypes of morphotypes I and II. These sites corroborate Ag-NOR signals located exclusively on the short arm of submetracentric chromosome pair 1. On the other hand, 5S rDNA sites are different with regard to frequency between the two morphotypes. Whereas in morphotype I, these sites are situated in the terminal position of the long arm on acrocentric chromosome pair 9, in morphotype II specimens, in addition to this region, they are also located on the short arm of chromosome pair 15 (Fig. 4).
Discussion
Morphological divergences in C. lugubris from the St. Peter and St. Paul Archipelago
The set of results obtained indicates the coexistence of two significantly different morphotypes of C. lugubris in areas surrounding the St. Peter and St. Paul Archipelago in the mid-Atlantic.
Morphological attributes in groups with wide geographic distribution usually vary according to the degree of population isolation or adaptive patterns associated with different habitats (Wainwright and Reilly 1994; Motta et al. 1995). The higher body and smaller eyes exhibited by morphotype I, as well as the less deep body and larger eyes apparent in morphotype II, may indicate differential adaptive characteristics. These may be related to swimming, predation or defense mechanisms, among other ecological aspects, thereby linked to the division of spatial resources (Motta 1988; Gibran 2007; 2010).
In addition to the C. lugubris morphotypes, the waters of SPSPA are home to other species of Carangid including C. latus, Carangoides bartholomaei, C. crysos and Elegatis bipinnulata (Feitoza et al. 2003). The genus Caranx is composed of generalist predators of fish and small crustaceans (Sazima 1986; Silvano 2001). Behavioral studies conducted on Caranx and Carangoides species indicate prey are captured both in the water column and rocky substrate, showing potential for a wide range of prey types (Barroso 1965; Potts 1980; Sancho 2000; Silvano 2001). Such behavioral and feeding flexibility is particularly appropriate for species inhabiting densely populated areas such as SPSPA (Feitoza et al. 2003), since they may undergo niche displacement. In these regions, adaptation to rapid fluctuations in the availability of resources may cause severe population restrictions, primarily among predators with less variable diets or those incapable of changing their feeding habits over time (Edgar and Shaw 1995).
Despite the limitations of using only morphological data in phylogenetic inferences, the set of body data and meristic characteristics of the morphotypes regarding sympatric species of Carangidae are informative. Morphological similarities were established between species of the same genus, discarding the action of ecological convergences that may mask the establishment of phylogenetic relationships, as seen in several groups of Actinopterygii (see Winemiller 1992). Notwithstanding displaying similar morphology to cogeneric species C. latus (sympatric) and C. hippos, the morphotypes demonstrated more pronounced morphological similarity among themselves.
Cytogenetic comparison of C. lugubris morphotypes
Although they show less variation in chromosomal number (2n = 46–48), as do several groups of Perciformes (Galetti et al. 2006; Molina 2007), the family Carangidae shows relative diversity in chromosome structure (FN = 46–78). Pericentric inversions in acrocentric pairs constitute the main karyotype diversification mechanisms in this group, as observed in Seriola quinqueradiata (2n = 48; FN = 52) (Ida et al. 1978), C. equula and Carangoides sexfasciatus (2n = 48; FN = 50) (Murofushi and Yoshida 1979).
Karyotype analyses of C. lugubris morphotypes I and II corroborate the cytogenetic characteristics typical of Perciformes, a group in which around 60 % of species studied exhibit 2n = 48 chromosomes, mostly associated with single NOR sites and reduced heterochromatin (Sola et al. 1997; Galetti et al. 2000). Among themselves, morphotypes display marked karyotype similarities, both in regard to karyotype formula, heterochromatic distribution and composition, and in the position and number of primary ribosomal sites (18S rDNA signals), located only on chromosome pair 1.
The presence of GC-rich heterochromatic regions is an uncommon condition in fish. The scattered occurrence of these sequences in the karyotype of C. lugubris indicates possible homogenization processes of these sequences in the karyotype evolution of this species. Greater heterochromatic dynamism in very diverse groups of fish (Moreira-Filho and Bertollo 1991) suggests they may promote a higher number of chromosomal rearrangements and represent a first step toward establishing post-zygotic barriers (Souza et al. 1995; Caputo et al. 1996; Molina and Galetti, 2002). However, heterochromatinization processes have been identified as a secondary evolutionary mechanism in the karyotype evolution of Perciformes (Galetti et al. 2000; Molina 2007).
Chromosome mapping of 18S rDNA and 5S rDNA sequences has pinpointed interspecific variations and indications of differences between fish populations in the Atlantic separated by biogeographical barriers (Motta-Neto et al. 2011). Ribosomal sites may be effective cytotaxonomic markers or indicators of chromosomal rearrangements in the karyotype. In fact, the 5S sites reveal divergences in the number of chromosome bearers between morphotypes, resulting in two distinct karyotypes. In the karyotype of morphotype I, only chromosome pair 9 shows signs of hybridization in the terminal portion of the long arm, while in morphotype II, as well as signals on these chromosomes in the same position, additional sites are also present in the terminal region of the short arm on chromosome pair 15.
There are few examples of intra- or interpopulation chromosomal polymorphism among Carangidae. One of the few cases was described in specimens of Seriola dumerili from the coast of Sicily, which exhibited two karyotypes, 2n = 47 and 2n = 48 (NF = 50), resulting from Robertsonian fusion (Vitturi et al. 1986). This condition is probably rare, since it was not recorded in subsequent studies (Sola et al. 1997).
Frequency or positioning of 5S rRNA genes has provided cytotaxonomic markers used for species identification and in evolutionary research (e.g., Aguilar and Galetti 2008). During karyotype evolution of several fish species, these genes can acquire independent characteristics. In some cases, they may be conserved (e.g., Wasko et al. 2001), whereas in others they display transient polymorphisms or fixed evolutionary divergences (e.g., Hatanaka and Galetti, 2004; Garcia et al. 2010). Although the difference in the number of 5S sites alone does not definitively ensure that C. lugubris morphotypes constitute genetically different stock, its constancy between karyotypes suggests a fixed condition among them.
Origins of different C. lugubris stocks
Although it is not currently possible to accurately establish the origin of differences between morphotypes of C. lugubris, some hypotheses can be raised based on examples from other fish groups. Some of these can be analyzed in detail, such as ecological diversification, interspecific hybridization and the breaching of marine biogeographical barriers by allopatric populations.
Diversification through ecological specializations has been identified, among others, in the evolution of cichlids from large African lakes (e.g., Genner et al. 1999), leading to notable morphological adaptations among closely related species (Clabaut et al. 2007). Although possible, this seems very unlikely as the cause of morphological and cytogenetic variations present in morphotypes of C. lugubris on SPSPA. The substantial mobility of this species, gregarious habits (in the form of vast schools) and few environmental micro-partitions on SPSPA (Lubbock and Edwards 1981) hamper population fractionation and restrict gene flow in such a small area.
Another possibility is the occurrence, in this ocean region, of a hybrid swarm resulting from interspecific hybridization between C. lugubris and another phylogenetically close species. Although interspecific hybrids are considered rare in marine fish (Randall et al. 1977), some cases have been reported in species from the family Carangidae (e.g., Murakami et al. 2007; Santos et al. 2011). In fish, this can happen accidentally during aggregations for spawning (in the same period and same location) among closely related species, or due to the scarcity or lack of reproductive partners of the same species, leading to spawning with individuals from a more abundant-related species (Frisch and Van Herwerden 2006). In fact, there is evidence of increased capture of several species, among them C. lugubris, reducing the population size of various species on SPSPA (Oliveira et al. 1997). Among available examples of hybridization in fish, morphological, meristic and coloration aspects of hybrids are generally intermediate to those of the parents. The occurrence of such an event would be supported by the expectation of some level of morphological similarity between one of both C. lugubris morphotypes with another closely related phylogenetically sympatric species from the family Carangidae. However, geometric morphometrics, which has proved to be decisive in intergroup (Nacua et al. 2010), showed that morphotypes exhibit even more similarities among themselves than with any other species of local Carangidae. Therefore, the absence of an intermediate pattern, whether in regard to color (both morphotypes displaying similar color patterns to C. lugubris) or number of serial or body elements, with any other sympatric species of the genera Caranx and Carangoides hinders accepting the hypothesis of hybridization as an explanation for distinct characteristics between morphotypes. The lack of intermediate morphological and cytogenetic patterns suggests the absence of genetic flow between morphotypes I and II.
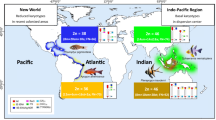
On the other hand, there are historical and contemporary examples of breached biogeographical barriers (e.g., Bowen et al. 2001; Luiz-Júnior et al. 2004) that shaped the distribution pattern of species in the Atlantic. These consist of the discharge from the Amazon/Orinoco rivers, which separate the Caribbean and Brazilian provinces; the vast ocean areas separating the East and West Atlantic (Banford et al. 1999); and the region of cold upwelling in southwest Africa that separates the Atlantic and Indian oceans (Briggs 1974). However, it would be possible that specimens of C. lugubris from the Caribbean, Western Atlantic, Eastern Atlantic or even the Indian Ocean (which can overstep cold waters of the Benguela Current) are meeting in ASPSP.
Exact understanding of the coexistence of morphotypes I and II of C. lugubris in SPSPA indicates the need for further studies. An intriguing question is whether these individuals occupy SPSPA through a contemporary event, as a result of a recent invasion, or live in sympatry, apparently reproductively isolated, for an extended period of time. Phylogeographic analysis of C. lugubris would be decisive in measuring its contemporary and historic distribution patterns, mainly in light of chromosomal and morphological variations found, as well as contribute to a better understanding of their occurrence. Data obtained highlight the evolutionary importance of isolated environments, such as SPSPA in the Atlantic, and their role in genetic differentiation, larval dispersion patterns and as a contact zone between different Atlantic regions.
References
Aguilar CT, Galetti PM Jr (2008) Chromosome mapping of 5S rRNA genes differentiates Brazilian populations of Leporellus vittatus (Anostomidae, Characiformes). Genet Mol Biol 31:188–194
Banford HM, Bermingham E, Collette BB, McCafferty SS (1999) Phylogenetic systematics of the Scomberomorus regalis (Teleostei: Scombridae) species group: molecules, morphology and biogeography of Spanish mackerels. Copeia 1999:596–613
Barros-e-Silva AE, Guerra M (2009) The meaning of DAPI bands observed after C-banding and FISH procedures. Biotech Histochem 85:115–125
Barroso LM (1965) Nota preliminar sobre a alimentação do xaréu-preto (Caranx lugubris, Poey 1860) no nordeste do Brasil. B de Est Pesca 5:7–11
Becker H (2001) Hidrologia dos bancos e ilhas oceânicas do nordeste brasileiro. Uma contribuição ao Programa Revizee. Doctoral Thesis. Universidade Federal de São Carlos
Bowen BW, Bass AL, Rocha LA, Grant WS, Robertson DR (2001) Phylogeography of the trumpetfish (Aulostomus spp.): ring species complex on a global scale. Evolution 55:1029–1039
Briggs JC (1974) Mar Zoo. McGraw-Hill, New York
Caputo V, Marchegftani F, Olmo E (1996) Karyotype differentiation between two species of carangid fishes, genus Trachurus (Perciformes: Carangidae). Mar Biol 127:193–199
Cioffi MB, Martins C, Centofante L, Jacobina U, Bertollo LAC (2009) Chromosomal variability among allopatric populations of Erythrinidae fish Hoplias malabaricus: mapping of three classes of repetitive DNAs. Cytogenet Genome Res 125:132–141
Clabaut C, Bunje PME, Salzburger W, Meyer A (2007) Geometric morphometric analyses provide evidence for the adaptive character of the Tanganyikan cichlid radiation. Evolution 61:560–578
Dryden IL, Mardia KV (1998) Statistical shape analysis. Wiley Press, New York
Edgar GJ, Shaw C (1995) The production and trophic ecology of shallow-water fish assemblages in southern Australia. II. Diets of fishes and trophic relationships between fishes and benthos at Western Port, Victoria. J Exp Mar Biol Ecol 194:83–106
Edwards AJ, Lubbock R (1983) Marine zoogeography of St. Paul’s Rocks. J Biogeogr 10:65–72
Feitoza B, Rocha LA, J-Luiz O Jr, Floeter SR, Gasparini JL (2003) Reef fishes of Saint Paul’s Rocks: new records and notes on biology and zoogeography. Aqua 7:61–82
Frisch A, Van Herwerden L (2006) Field and experimental studies of hybridization between coral trouts, Plectropomus leopardus and Plectropomus maculatus (Serranidae), on the Great Barrier Reef, Australia. J Fish Biol 68:1013–1025
Galetti PM Jr, Aguilar CT, Molina WF (2000) An overview of marine fish cytogenetics. Hydrobiologia 420:55–62
Galetti PM Jr, Molina WF, Affonso PRAM, Aguilar CT (2006) Assessing genetic diversity of Brazilian reef fishes by chromosomal and DNA markers. Genetica 126:161–177
Garcia C, Oliveira C, Almeida-Toledo LF (2010) Karyotypic evolution trends in Rhamdia quelen (Siluriformes, Heptapteridae) with considerations about the origin and differentiation of its supernumerary chromosomes. Genet Mol Res 9:365–384
Genner MJ, Turner GF, Barker S, Hawkins SJ (1999) Niche segregation among Lake Malawi cichlid fishes? Evidence from stable isotope signatures. Ecol Lett 2:185–190
Gibran FZ (2007) Activity, habitat use, feeding behavior, and diet of four sympatric species of Serranidae (Actinopterygii: Perciformes) in southeastern Brazil. Neotrop Ichthyol 5:387–398
Gibran FZ (2010) Habitat partitioning, habits and convergence among coastal nektonic fish species from the São Sebastião Channel, southeastern Brazil. Neotrop Ichthyol 8:299–310
Gold LC Jr, Shipley NS, Powers PK (1990) Improved methods for working with fish chromosomes with a review of metaphase chromosome banding. J Fish Biol 37:563–575
Hatanaka T, Galetti PM Jr (2004) Mapping of the 18S and 5S ribosomal RNA genes in the fish Prochilodus argenteus Agassiz, 1829 (Characiformes, Prochilodontidae). Genetica 122:239–244
Honebrink RR (2000) A review of the family Carangidae, with emphasis on species found in Hawaiian waters. DAR technical report 20-01. Division of Aquatic Resources, Department of Land & Natural Resources, Hawaii
Howell WM, Black DA (1980) Controller silver staining of nucleolus organizer region with protective colloidal developer: a 1-step method. Experientia 36:1014–1015
Ida H, Murofushi M, Fujiwawa S, Fujino K (1978) Preparation of fish chromosomes by in vitro colchicine treatment. Jpn J Ichthyol 24:281–284
Klingenberg CP (2008) MorphoJ. Faculty of Life Sciences, University of Manchester, Manchester. Available from: http://www.flywings.org.uk/MorphoJpage.htm
Lubbock R, Edwards A (1981) Fishes of Saint Paul’s Rocks. J Fish Biol 18:135–157
Luiz-Júnior OJ, Floeter SR, Gasparini JL, Ferreira CEL, Wirtz P (2004) The occurrence of Acanthurus monroviae (Perciformes: Acanthuridae) in the south-western Atlantic, with comments on other eastern Atlantic reef fishes occurring in Brazil. J Fish Biol 65:1173–1179
Martins C, Ferreira IA, Oliveira C, Foresti F, Galetti PM Jr (2006) A tandemly repetitive centromeric DNA sequence of the fish Hoplias malabaricus (Characiformes: Erythrinidae) is derived from 5S rDNA. Genetica 127:133–141
Molina WF (2001) An alternative method for mitotic stimulation in fish cytogenetics. Chromosom Sci 5:149–152
Molina WF (2007) Chromosomal changes and stasis in marine fish groups. In: Pisano E, Ozouf-Costaz C, Foresti F, Kapoor BG (eds) Fish cytogenetics. Science Publishers, Enfield, pp 69–110
Molina WF, Galetti PM Jr (2002) Robertsonian rearrangements in the reef fish Chromis (Perciformes, Pomacentridae) involving chromosomes bearing 5S rRNA genes. Genet Mol Biol 25:373–377
Molina WF, Alves DOE, Araújo WC, Martinez PA, Silva MFM, Costa GWWF (2010) Performance of human immunostimulant agents in the improvement of fish cytogenetics. Genet Mol Res 9:1807–1814
Moreira-Filho O, Bertollo LAC (1991) Astyanax scabripinnis (Pisces, Characidae): a species complex. Rev Bras Genet 14:331–357
Motta PJ (1988) Functional morphology of the feeding apparatus of ten species of Pacific butterflyfishes (Perciformes, Chaetodontidae): an ecomorphological approach. Environ Biol Fish 22:39–67
Motta PJ, Clifton KB, Hernandez P, Eggold BT (1995) Ecomorphological correlates in ten species of subtropical seagrass fishes: diet and microhabitat utilization. Environ Biol Fishes 44:37–60
Motta-Neto CC, Cioffi MB, Bertollo LAC, Molina WF (2011) Extensive chromosomal homologies and evidence of karyotypic stasis in Atlantic grunts of the genus Haemulon (Perciformes). J Exp Mar Biol Ecol 401:75–79
Murakami K, James SA, Randall JE, Suzumoto AY (2007) Two hybrids of carangid fishes of the genus Caranx, C. ignobilis × C. melampygus and C. melampygus × C. sexfasciatus, from the Hawaiian Islands. Zoo Stud 46:186–193
Murofushi M, Yoshida TH (1979) Cytogenetical studies on fishes. II. Karyotypes of four carangid fishes. Jpn J Genet 54:367–370
Nacua SS, Dorado EL, Torres MAJ, Demayo CG (2010) Body shape variation between two populations of the white goby, Glossogobius giuris (Hamilton and Buchanan). Res J Fish Hydrobiol 5:44–51
Oliveira GM, Evangelista JEV, Ferreira BP (1997) Considerações sobre a biologia e a pesca no arquipélago dos penedos de São Pedro e São Paulo. Boletim Técnico Cientifico, 5, CEPENE, Tamandaré-PE 16
Palumbi SR, Metz EC (1991) Strong reproductive isolation between closely related tropical sea urchins (genus Echinometra). Mol Biol Evol 8:227–239
Pfennig DW, Wund MA, Snell-Rood EC, Cruickshank T, Schlichting CD, Moczek AP (2010) Phenotypic plasticity’s impacts on diversification and speciation. Trends Ecol Evol 25:459–467
Pinkel D, Straume T, Gray J (1986) Cytogenetic analysis using quantitative, high sensitivity, fluorescence hybridization. Proc Natl Acad Sci USA 83:2934–2938
Potts GW (1980) The predatory behaviour of Caranx melampygus (Pisces) in the channel environment of Aldabra Atoll (Indian Ocean). J Zool Lond 192:323–350
Randall JE, Allen GR, Steene RC (1977) Five probable hybrid butterflies fishes of the genus Chaetodon from the central and western Pacific. Rec Aust Mus 6:3–26
Rocha LA, Robertson DR, Roman J, Bowen BW (2005) Ecological speciation in tropical reef fishes. Proc R Soc B 272:573–579
Rohlf FJ (2004) TpsDig. Department of Ecology and Evolution. Stony Brook, New York
Rohlf FJ, Slice DE (1990) Methods for comparison of sets of landmarks. Syst Zool 39:40–59
Sancho G (2000) Predatory behaviors of Caranx melampygus (Carangidae) feeding on spawning reef fishes: a novel ambushing strategy. Bull Mar Sci 66:487–496
Santos SR, Xiang Y, Tagawa AW (2011) Population structure and comparative phylogeography of jack species (Caranx ignobilis and C. melampygus) in the high Hawaiian Islands. J Hered 102:47–54
Sazima I (1986) Similarities in feeding behaviour between some marine and freshwater fishes in two tropical communities. J Fish Biol 29:53–65
Silvano RAM (2001) Feeding habits and interspecific feeding associations of Caranx latus (Carangidae) in a subtropical reef. Environ Biol Fishes 60:465–470
Smith-Vaniz WF, Carpenter KE (2007) Review of the crevalle jacks, Caranx hippos complex (Teleostei: Carangidae), with a description of a new species from West Africa. Fish Bull 105:207–233
Sola L, Cipelli O, Gornung E, Rossi AR, Andaloro F, Crosetti D (1997) Cytogenetic characterization of the greater amberjack, Seriola dumerili (Pisces: Carangidae), by different staining techniques and fluorescence in situ hybridization. Mar Biol 128:573–577
Souza IL, Moreira-Filho O, Bertollo LAC (1995) Cytogenetic diversity in the Astyanax scabripinnis (Pisces, Characidae) complex. II. Different cytotypes living in sympatry. Cytologia 60:273–281
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304–306
Vaske T Jr, Lessa RP, Nóbrega M, Montealegre-Quijano S, Marcante Santana F, Bezerra JL Jr (2005) A checklist of fishes from Saint Peter and Saint Paul Archipelago, Brazil. J App Ichthyol 21:75–79
Vitturi R, Mazzola A, Macaluso M, Catalano E (1986) Chromosomal polymorphism associated with Robertsonian fusion in Seriola dumerili (Risso, 1810) (Pisces: Carangidae). J Fish Biol 26:529–534
Wainwright PC, Reilly SM (1994) Ecological morphology: integrative organism biology. The University of Chicago Press, Chicago, p 367
Wasko AP, Martins C, Wright JM, Galetti PM Jr (2001) Molecular organization of 5S rDNA in fishes of the genus Brycon. Genome 44:893–902
Winemiller KO (1992) Ecomorphology of freshwater fishes. Ecological divergence and convergence in freshwater fishes. Nat Geo Res Expl 3:308–327
Acknowledgments
We would like to thank the National Council of Technological and Scientific Development—CNPq for their financial support (Process No. 556793/2009-9), IBAMA (Process No. 19135/1) for giving permission to collect specimens and SECIRM for providing us with the means to conduct this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H.-D. Franke.
Rights and permissions
About this article
Cite this article
Jacobina, U.P., Martinez, P.A., Cioffi, M.B. et al. Morphological and karyotypic differentiation in Caranx lugubris (Perciformes: Carangidae) in the St. Peter and St. Paul Archipelago, mid-Atlantic Ridge. Helgol Mar Res 68, 17–25 (2014). https://doi.org/10.1007/s10152-013-0365-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10152-013-0365-0