Abstract
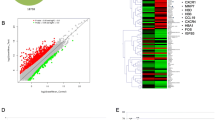
A number of previous studies have revealed the abnormal expression of various angiogenesis-related genes or products in brain arteriovenous malformation (AVM). To understand the molecular process of this disease, we analyzed gene expression profiles in brain AVM. Using a DNA microarray consisting of 17,086 genes, we identified differentially expressed genes in 5 brain AVMs from their draining veins, vessels retaining basic venous architecture. Not many genes were differentially expressed between the AVM nidus and the draining vein. When we applied an absolute cut-off value for normalized log2 (cy5/cy3 ratio) of 0.4, 19 genes were selected. Genes such as SOX8, TRIM2, FENA1 (ephrin A1), and AQP4 were upregulated, and genes such as I_1000105, KRT18, IGFBP7, EMILIN-2, and KRT14 were downregulated. Genes relating to angiogenesis, such as vascular endothelial growth factor and angiopoietin and other members of the ephrin family, were not differentiated. Among differentially expressed genes detected in this analysis, we focused on ephrin A1, a gene related to embryogenesis and angiogenesis. The expression of ephrin A1 was two and three to nine times higher than that of the draining vein and normal brain, respectively, using real-time reverse transcription-polymerase chain reaction. For the first time, here we report the increased expression of ephrin A1 in brain AVM, which may play an important role in the pathogenesis of AVM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Arteriovenous malformation (AVM) of the brain is defined as the direct communication between one or more arteries of the cerebral parenchyma and one or more draining veins without the intervention of a capillary bed [3, 15, 23]. Most brain AVM vessels are composed of large, thin-walled dilated channels displaying fibroblast proliferation and increased cellularity in the wall with focal deposition of elastic tissue layers. They are generally believed to arise when capillaries fail to develop in an area of the brain during early embryogenesis, resulting in abnormal communications between arteries and veins. The growth of blood vessels during embryogenesis and in adult organisms is tightly controlled, and this growth is thought to be mediated by soluble factors [13, 19, 27]. A number of previous studies revealed the abnormal expression of various angiogenesis-related genes or products in brain AVM [7–11, 14, 16, 27].
In order to understand the molecular process of this disease, we attempted to identify differentially expressed genes in brain AVMs as compared to draining veins, vessels retaining the basic venous architecture [3, 15]. We approached the problem by examining gene-expression profiles. DNA microarray technology nowadays makes it possible to simultaneously analyze the expression patterns of thousands of genes in a given tissue [12, 18, 22]. We here report the identification of ephrin A1, a gene involved in embryogenesis and angiogenesis, which is overexpressed in the nidus of brain AVM.
Materials and methods
Samples and clinical information
After institutional review and informed consent, we obtained brain AVM specimens after resection. One specimen of normal brain was obtained from a temporal lobectomy that was performed for epilepsy. The brain AVM nidus was cut into two pieces for histology and DNA microarray analysis. The nidus was dissected from the adjacent brain tissue in the operating theatre, and a representative portion of nidus tissue was stored at -80°C until analysis. Draining vein specimens were also obtained from the five brain AVM patients. DNA microarray analysis was performed to compare the expression levels of genes between the nidus and draining vein of each patient. The clinical information for patients with brain AVM is shown in Table 1. Three patients had a history of hemorrhage from the brain AVM. None of the patients underwent radiosurgery. Two patients had embolization treatment before surgical resection.
RNA preparation
Frozen sections were placed directly in Trizol reagent (Invitrogen), homogenized for 2 min on ice, and RNA was isolated according to the manufacturer’s protocol. RNA was quantified based on absorbance at 260 nm. Total RNA was assessed for quality with an Agilent Lab-on-a-Chip 2100 bioanalyzer. Possible traces of genomic DNA contaminating the RNA preparations were removed by DNase I (Promega) digestion.
Labeling and hybridization
The targets for DNA microarray analysis were prepared according to the manufacturer’s instructions. Briefly, cyanine 3- and cyanine 5-labeled complementary RNA (cRNA) was synthesized followed by the production of double-stranded cDNA from 50–500 ng of total RNA with a Human 1A oligoDNA microarray kit (Agilent). After the purification of labeled cRNA, arrays containing 17,086 genes were hybridized with the fragmented cRNA targets at 60°C for 17 h, and then washed and dried with air. We repeated the experiments using the same sample set, but substituting dyes.
Data analysis
Scans were performed using an Agilent DNA microarray scanner (model G2565BA), and the expression value for each gene was calculated by Feature Extraction software. Data were normalized by the LOWESS normalization method. Only the genes deemed present on all slides were further considered. When both cyanine5 and cyanine3 dyes produced signal intensities lower than the cut-off [2.6× standard deviation (SD) of background], we excluded these genes from further analysis. We excluded a sample set when the correlation coefficient for each pair of color swapping was positive. We calculated the mean and SD of log2 (r1 × r2, where r1 and r2 are cy5/cy3 ratios for color swapping) for each pair of sample sets in the entire array, and eliminated pairs of elements whose log2 (r1 × r2) was greater than the SD from the mean. We selected genes whose absolute value for normalized log2 (cy5/cy3 ratio) was higher than the cut-off value determined in all sample sets examined [1, 25].
Quantitative real time RT-PCR
To evaluate the expression level of the target genes, quantitative RT-PCR was performed using a Sequence Detection System, model 7700 (Perkin Elmer). Quantitative real-time RT-PCR primers were designed using Primer 3. A sense primer and an antisense primer were synthesized using sequences from a database search at NCBI for human ephrin A1 and GAPDH as follows: ephrin A1 sense primer: TGGAACAGTTCAAATCCCAAG; ephrin A1 antisense primer: CTCATGCTCCACCAGGTACA; GAPDH sense primer: CAGCCTCAAGATCATCAGCA; GAPDH antisense primer: ATCCACAGTCTTCTGGGTGG. Five serial dilutions of each total RNA sample (100, 50, 25, 12.5, and 6.25 ng total RNA) were analyzed for each target gene. Amplification reactions (25 μl) comprised the total RNA samples: 2× SYBR green PCR master mix, 40× MultiScribe reverse transcriptase and RNase inhibitor mix, and 200 nmol/l of each primer. The thermal cycling conditions were as follows: 30 min at 48°C for reverse transcription, 10 min at 95°C for the AmpliTaq Gold activation, 40 cycles of 15 s at 95°C for denaturation, and 1 min at 60°C for annealing and extension. CT values corresponded to the cycle number at which the fluorescent emission monitored in real time reached the threshold, set at 10 SD above the mean of the baseline emission calculated from cycles 5 to 15. The CT values decreased linearly with increasing target quantity and could be used as a quantitative measurement of the input target number. Total RNA concentrations from each sample were normalized by the quantity of GAPDH mRNA, and the expression levels of the target genes were evaluated by the ratio of the number of target mRNA to GAPDH mRNA.
Immunohistochemical study
Immunohistochemistry was performed using paraffin-embedded tissue. Heat-induced epitope retrieval was performed in 10 mmol/l citrate buffer. After blocking, the slides were then incubated with a primary antibody. The slides were incubated with a Histofine simple stain MAX-PO(Multi) (Nichirei) for 30 min at room temperature. The slides were washed again and incubated with detecting reagent. The slides were counterstained with hematoxylin. Primary antibodies were polyclonal rabbit anti-human ephrin A1 antibody (Zymed Laboratories). The sensitivity and specificity of primary antibodies were verified using a positive control (prostate) and a negative control (no primary antibody).
Results
One sample set (patient 1, Table 1) was excluded from this study among the five sample sets because the correlation coefficient of color swapping was positive. When we applied an absolute cut-off value for the normalized log2 (cy5/cy3 ratio) of 0.3, 28 genes were selected; when we applied 0.4, 19 genes were selected. Table 2 shows six upregulated genes in the nidus of brain AVM compared to the draining vein. Table 3 shows 13 genes downregulated in the nidus of brain AVM compared to the draining vein. We examined the signal-intensity ratios of housekeeping genes and genes previously reported to be related to the pathogenesis of brain AVM: ACTB (beta actin); GAPDH (glyceraldehydes-3-phosphate dehydrogenase); VEGF (vascular endothelial growth factor); FLT1 (Fms-related tyrosine kinase 1); ANGPT1 (angiopoietin 1); ANGPT2 (angiopoietin 2); TIE (tyrosine kinase with immunoglobulin and epidermal growth factor homology domains); ENG (endoglin); ACVRL1 (activin receptor-like kinase-1), MMP9 (matrix metalloproteinase 9) and HIF1A (hypoxia-inducible factor 1 alpha subunit). There were no significant differences in gene expression between the nidus of brain AVM and the draining vein (minimum ratio <1, maximum ratio >1) (Table 4). We also examined the signal-intensity ratios of ephrin family genes: EPHA 1–9 (ephrin A 1–9 receptor); EPHB1–6 (ephrin B 1–6 receptor); EFNA 1–6 (ephrin A 1–6); EFNB 1–3 (ephrin B 1–3). Ephrin A6, A9, B4, B5 receptors and ephrin A6 were not detected at all. There were no significant differences in gene expression between the nidus of brain AVM and the draining vein (minimum ratio <1, maximum ratio >1) in other genes of the ephrin family except for ephrin A1. Fig. 1 shows the results of real-time RT-PCR in the draining vein, nidus, and the specimen of normal brain (one sample). The experiment was repeated three times in samples of patients 1, 4, and 5 (see Table 1). The gene expression of ephrin A1 in the nidus was upregulated two and three to nine times higher than that of the draining vein and normal brain specimen. In an immunohistochemical study of eight brain AVMs, expression of ephrin A1 was detected in astrocytes and neurons of the perinidal parenchymal tissue, but not in the surrounding reactive granulomatous tissue nor the normal brain tissue nor in the AVM vessels (Fig. 2).
Gene expressions of ephrin A1 in the draining vein (V), nidus of arteriovenous malformation (N), and normal brain tissue (B). Quantitative real-time RT-PCR was repeated using samples from patients 1, 4, and 5 in Table 1. Expression levels are expressed as relative ratios in CT values for ephrin A1 to GAPDH
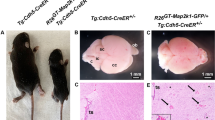
Localization of ephrin A1 in the nidus of brain AVM: immunohistochemical study. A: Ephrin A1-positive astrocytes in perinidal parenchymal tissue (original magnification 200); B: ephrin A1-positive neurons in the cortex of the perinidal region (×200); C: no ephrin A1-positive cells in the surrounding reactive granulomatous tissue of perinidal parenchyma. Depositions of hemosiderin are seen (×100); D: no ephrin A1-positive cells in AVM nidus (×100)
Discussion
The results of the DNA microarray analysis indicate that not many genes are differentially expressed between the AVM nidus and draining vein. When we applied an absolute cut-off value for the normalized log2 (cy5/cy3 ratio) of 0.3, only 28 genes were selected; when we applied 0.4, 19 genes were selected. Genes such as SOX8, TRIM2, FENA1 (ephrin A1), and AQP4 related to brain were upregulated. Among the downregulated genes, several genes were related to the extracellular matrix and keratin such as I_1000105, KRT18, IGFBP7, EMILIN-2, and KRT14. We focused on ephrin A1, a gene related to embryogenesis and angiogenesis, among differentially expressed genes detected in this analysis. We confirmed the upregulation of ephrin A1 in the nidus compared to the draining vein and the normal brain tissue using real-time RT-PCR. Our immunohistochemical study showed that ephrin A1 was highly expressed in astrocytes and neurons of the perinidal parenchymal tissue, but not in the endothelial or smooth muscle cells of the AVM vessels. Cells in normal brain tissue were not stained by ephrin A1 antibody.
Ephrin A1 was originally identified as a tumor necrosis factor (TNF)-α-inducible gene in human umbilical vein endothelial cells and is expressed in the developing vasculature during embryogenesis [18, 22]. Ephrin A1 induces endothelial cell migration and capillary assembly in vitro and angiogenesis in the corneal pocket assay in vivo [4, 18]. These studies indicate that ephrin A1 is critical for normal blood vessel development and suggest that the molecule may also play a role in angiogenesis for pathological processes in adults. The increasing size of AVMs over time has been documented in the literature [26]. Recurrent AVMs after negative angiograms have also been reported [13]. It is well known that new shunts between arteries and veins grow after embolization of feeding arteries.
We compared our DNA microarray data of angiogenesis-related genes with previously published data. We found no differences in these genes between the AVM nidus and draining vein. The decreased expression of vascular endothelial growth factor receptor (VEGFR), angiopoietin 1 (Ang1) and tyrosin kinase with immunoglobulin and epidermal growth factor homology domains 2 (Tie2), together with an increased expression of Ang2, matrix metalloproteinase (MMP)-9, and tissue inhibitors of metalloproteinases (TIMPs) was reported [7–9]. Hashimoto et al. confirmed a general correlation between many of the previously published findings on abnormal angiogenesis-related protein expression in brain AVM and gene microarray analysis [10]. Sure et al. recently indicated that in AVMs treated with embolization before surgery the expression of HIF1A and VEGF was significantly higher than in patients who did not have previous endovascular treatment [21]. In our data set there was no significant difference of HIF1A and VEGF between AVM patients with and without embolization before surgery. The difference may be due to the different tissues used for control samples as comparisons to brain AVM. Previous studies all used normal cortex as a control, whereas we selected the draining vein as control, because most of the AVM nidus consists of vessels with a dilated venule-like structure, and the draining vein retains that basic venous architecture. Moreover, our samples are heterogenous in clinical features: the size of the nidus varies, two have a history of bleeding, embolization was performed in two. Gene expressions of the nidus and the vein may be affected. Obviously, it would have been better and more informative if the gene expression levels in the vessel and the brain component of the nidus, the normal vein and the tissue of the normal brain were compared individually.
Among the ephrin family, ephrin A1 was upregulated only in the nidus of brain AVM as compared to the draining vein. The ephrin A1 ligand is expressed predominantly in tumor cells, and ephrin A2 receptors are expressed primarily in a complementary fashion in tumor-associated vasculature [17]. Treatment with soluble EphA2-Fc chimeric receptor results in decreased neovascularization in two different tumor types in vascular window assays, and EphA2-Fc of EphA3-Fc treatment results in decreased tumor vascular density and impaired tumor progression in vivo [2]. Although the pathological state of brain AVM is completely different from tumor neovascularization, a similar approach may elucidate the pathogenesis of brain AVM and help in finding a new treatment for this disease.
Ephrin B2 is expressed in developing arterial endothelial cells, whereas ephrin B4 receptor has a complementary expression and is confined to venous endothelial cells. The ephrin B4 receptor protein and ephrin B2 are therefore considered to define the boundaries between arterial and venous endothelial cells [5, 6]. It was recently reported that endoglin and activin receptor-like kinase-1 (ACVRL1) are necessary for the maintenance of distinct arterial-venous vascular beds and that attenuation of the ACVRL1 signaling pathway is the precipitating event in the etiology of hereditary hemorrhagic telangiectasia, a disease associated with AVM and characterized by the inappropriate fusion of arterioles with venules [20, 24]. We did not detect differences in the expressions of these genes between the AVM nidus and draining vein.
In conclusion, we here report for the first time the increased expression of ephrin A1 in the nidus of brain AVM, which may play important roles in the development and maintenance of AVM.
References
Arimoto T, Katagiri T, Oda K, Tsunoda T, Yasugi T, Osuga Y, Yoshikawa H, Nishii O, Yano T, Taketani Y, Nakamura Y (2003) Genome-wide cDNA microarray analysis of gene-expression profiles involved in ovarian endometriosis. Int J Oncol 22:551–560
Brantley DM, Cheng N, Thompson EJ, Lin Q, Brekken RA, Thorpe PE, Muraoka RS, Cerretti DP, Pozzi A, Jackson D, Lin C, Chen J (2002) Soluble Eph A receptors inhibit tumor angiogenesis and progression in vivo. Oncogene 21:7011–7026
Challa VR, Moody DM, Brown WR (1995) Vascular malformations of the central nervous system. J Neuropath Exp Neurol 54:609–621
Cheng N, Brantley DM, Liu H, Lin Q, Enriquez M, Gale N, Yancopoulos G, Cerretti DP, Daniel TO, Chen J (2002) Blockade of EphA receptor tyrosine kinase activation inhibits vascular endothelial cell growth factor-induced angiogenesis. Mol Cancer Res 1:2–11
Dodelet VC, Pasquale EB (2000) Eph receptors and ephrin ligands: embryogenesis to tumorigenesis. Oncogene 19:5614–5619
Gale NW, Yancopoulos GD (1999) Growth factors acting via endothelial cell-specific receptor tyrosine kinases: VEGFs, angiopoietins, and ephrins in vascular development. Genes Dev 13:1055–1066
Hashimoto T, Emala CW, Joshi S, Mesa-Tejada R, Quick CM, Feng L, Libow A, Marchuk DA, Young WL (2000) Abnormal pattern of Tie-2 and vascular endothelial growth factor receptor expression in human cerebral arteriovenous malformations. Neurosurgery 47:910–919
Hashimoto T, Lam T, Boundreau NJ, Bollen AW, Lawton MT, Young WL (2001) Abnormal balance in the angiopoietin-Tie2 system in human brain arteriovenous malformations. Circ Res 89:111–113
Hashimoto T, Wen G, Lawton MT, Boudreau NJ, Bollen AW, Yang GY, Barbaro NM, Higashida RT, Dowd CF, Halbach VV, Young WL (2003) Abnormal expression of matrix metalloproteinases and tissue inhibitors of metalloproteinases in brain arteirovenous malformations. Stroke 34:925–931
Hashimoto T, Lawton MT, Wen G, Yang GY, Chaly Jr T, Stewart CL, Dressman HK, Barbaro NM, Marchuk DA, Young WL (2004) Gene microarray analysis of human brain arteriovenous malformation. Neurosurgery 54:410–425
Hatva E, Jaaskelainen J, Hirvonen H, Alitalo K, Haltia M (1996) Tie endothelial cell-specific receptor tyrosine kinase is upregulated in the vasculature of arteriovenous malformations. J Neuropath Exp Neurol 55:1124–1133
Holzman LB, Marks RM, Dixit VM (1990) A novel immediate-early response gene of endothelium is induced by cytokines and encodes a secreted protein. Mol Cell Biol 10:5830–5838
Kader A, Goodrich JT, Sonstein WJ, Stein BM, Carmel PW, Michelsen WJ (1996) Recurrent cerebral arteriovenous malformations after negative postoperative angiograms. J Neurosurg 85:14–18
Koizumi T, Shiraishi T, Hagihara N, Tabuchi K, Hayashi T, Kawano T (2002) Expression of vascular endothelial growth factors and their receptors in and around intracranial arteriovenous malformations. Neurosurgery 50:117–126
Mandybur TI, Nazek M (1990) Cerebral arteriovenous malformations. A detailed morphological and immunohistochemical study using actin. Arch Pathol Lab Med 114:970–973
Matsubara S, Bourdeau A, terBrugge KG, Wallace C, Letarte M (2000) Analysis of endoglin expression in normal brain tissue and in cerebral arteriovenous malformations. Stroke 31:2653–2660
Ogawa K, Pasqualini R, Lindberg RA, Kain R, Freeman AL, Pasquale EB (2000) The ephrin-A1 ligand and its receptor, EphA2, are expressed during tumor neovascularization. Oncogene 19:6043–6052
Pandey A, Shao H, Marks RM, Polverini PJ, Dixit VM (1995) Roles of B61, the ligand for the Eck receptor tyrosine kinase, in YNF-α-induced angiogenesis. Science 268:567–569
Sonstein WJ, Kader A, Michelsen WJ, Llena JF, Hirano A, Casper D (1996) Expression of vascular endothelial growth factor in pediatric and adult cerebral arteriovenous malformations: an immunocytochemical study. J Neurosurg 85:838–845
Sorensen LK, Brooke BS, Li DY, Urness LD (2003) Loss of distinct arterial and venous boundaries in mice lacking endoglin, a vascular-specific TGFβ coreceptor. Dev Biol 261:235–250
Sure U, Battenberq E, Dempfle A, Tirakotai W, Bien S, Bertalanffy H (2004) Hypoxia-inducible factor and vascular endothelial growth factor are expressed more frequently in embolized than in nonembolized cerebral arteriovenous malformations. Neurosurgery 55:663–670
Takahashi H, Ikeda T (1995) Molecular cloning and expression of rat and mouse B61 gene: implications on organogenesis. Oncogene 11:879–883
The arteriovenous malformation study group (1999) Arteriovenous malformations of the brain in adults. New Eng J Med 340:1812–1818
Urness LD, Sorensen LK, Dean YL (2000) Arteriovenous malformations in mice lacking activin receptor-loke kinase-1. Nature Genet 26:328–331
Yang IV, Chen E, Hasseman JP, Liang W, Frank BC, Wang S, Sharov V, Saeed AI, White J, Li J, Lee NH, Yeatman TJ, Quackenbush J (2002) Within the fold: assessing differential expression measures and reproducibility in microarray assays. Genome Biology 3:research 0062.1–0062.12
Yasargil MG (1987) Microneurosurgery. Vol IIA. Thieme, New York, pp 138–160
Zadeh G, Guha A (2003) Angiogenesis in nervous system disorders. Neurosurgery 53:1362–1376
Acknowledgments
This work was supported in part by a grant-in-aid for scientific research (C) from the Japanese Ministry of Education, Culture, Sports, Science, and Technology (to H. K. and H.O.); the Japanese Ministry of Health, Labor, and Welfare (to H.K. and I.I.) and a Research for the Future Program Grant from the Japan Society for the Promotion of Science (I.I.). We thank Mr. Takashi Sakayori for his technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Comments
R. Loch Macdonald, Toronto, Canada
The authors used DNA microarrays, quantitative PCR and immunohistochemistry to determine if brain AVMs expressed different genes compared to the draining vein of the AVM. Several assumptions are inherent in the design, but the authors have done an excellent series of experiments to document that there is increased expression of ephrin A1 in brain AVMs. The immunohistochemistry suggests it is in the brain tissue around the AVM rather than nidal vessels. The utility of this kind of study is to now provide rationale for additional study of upregulated genes because at this point, one cannot know if the increase is a primary cause of AVM formation or pathogenesis or just a secondary reaction to hemorrhage or the presence of the AVM. However, that is what could be answered in detailed studies in an animal model such as the rat carotid-jugular fistula model [1–4].
References
1. Bederson JB, Wiestler OD, Brustle O, Roth P, Frick R, Yasargil MG (1991) Intracranial venous hypertension and the effects of venous outflow obstruction in a rat model of arteriovenous fistula. Neurosurgery 29:341–350
2. Lawton MT, Jacobowitz R, Spetzler RF (1997) Redefined role of angiogenesis in the pathogenesis of dural arteriovenous malformations. J Neurosurg 87:267–274
3. Morgan MK, Anderson RE, Sundt TM (1989) The effects of hyperventilation on cerebral blood flow in the rat with an open and closed carotid-jugular fistula. Neurosurgery 25:606–611
4. Yassari R, Sayama T, Jahromi BS, Aihara Y, Stoodley M, Macdonald RL (2004) Angiographic, hemodynamic and histological characterization of an arteriovenous fistula in rats. Acta Neurochir (Wien ) 146:495–504
Kiyohiro Houkin, Sapporo, Japan
This manuscript describes “Increased expression of ephrin A in brain AVM.” It is a novel and distinguished finding that ephrin A is expressed in neurons or glia in the perinidal parenchymal tissue. This work is one of the continuing studies by Tokyo Women’s Medical University group. Their studies have always been leading the study on gene expression of vascular lesions. This basic study has a chance to open the door to the translational research to the novel strategy for the intractable AVM.
I guess that further study will be appreciated to figure out whether this increased gene expression is the primary pathology or the secondary reaction in the pathological processes. Samples of nidus seem to contain parenchymal tissues, although those of draining veins do not. It would be more informative to study the gene expression profiles of only vascular components in nidus, drainer and normal brain. The detailed analysis of receptor expression for ephrin A in the vascular component would also be very helpful.
I hope that their continuous endeavor to reveal the gene expression in vascular lesion will disclose the unknown mechanism of the formation of AVM and other intracranial vascular lesions. I know that there is a long way in front of us to reach the novel treatment for the difficult AVM. However, without any doubt, I am confident that the authors are leading researchers to develop this difficult task.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License ( https://creativecommons.org/licenses/by-nc/2.0 ), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Sasahara, A., Kasuya, H., Akagawa, H. et al. Increased expression of ephrin A1 in brain arteriovenous malformation: DNA microarray analysis. Neurosurg Rev 30, 299–305 (2007). https://doi.org/10.1007/s10143-007-0087-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10143-007-0087-3