Abstract
Background
Glioblastoma (GBM) is an aggressive and unstoppable malignancy. Natural killer T (NKT) cells, characterized by specific markers, play pivotal roles in many tumor-associated pathophysiological processes. Therefore, investigating the functions and complex interactions of NKT cells is great interest for exploring GBM.
Methods
We acquired a single-cell RNA-sequencing (scRNA-seq) dataset of GBM from Gene Expression Omnibus (GEO) database. The weighted correlation network analysis (WGCNA) was employed to further screen genes subpopulations. Subsequently, we integrated the GBM cohorts from The Cancer Genome Atlas (TCGA) and Chinese Glioma Genome Atlas (CGGA) databases to describe different subtypes by consensus clustering and developed a prognostic model by least absolute selection and shrinkage operator (LASSO) and multivariate Cox regression analysis. We further investigated differences in survival rates and clinical characteristics among different risk groups. Furthermore, a nomogram was developed by combining riskscore with the clinical characteristics. We investigated the abundance of immune cells in the tumor microenvironment (TME) by CIBERSORT and single sample gene set enrichment analysis (ssGSEA) algorithms. Immunotherapy efficacy assessment was done with the assistance of Tumor Immune Dysfunction and Exclusion (TIDE) and The Cancer Immunome Atlas (TCIA) databases. Real-time quantitative polymerase chain reaction (RT-qPCR) experiments and immunohistochemical profiles of tissues were utilized to validate model genes.
Results
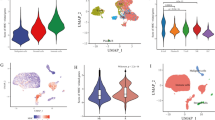
We identified 945 NKT cells marker genes from scRNA-seq data. Through further screening, 107 genes were accurately identified, of which 15 were significantly correlated with prognosis. We distinguished GBM samples into two distinct subtypes and successfully developed a robust prognostic prediction model. Survival analysis indicated that high expression of NKT cell marker genes was significantly associated with poor prognosis in GBM patients. Riskscore can be used as an independent prognostic factor. The nomogram was demonstrated remarkable utility in aiding clinical decision making. Tumor immune microenvironment analysis revealed significant differences of immune infiltration characteristics between different risk groups. In addition, the expression levels of immune checkpoint-associated genes were consistently elevated in the high-risk group, suggesting more prominent immune escape but also a stronger response to immune checkpoint inhibitors.
Conclusions
By integrating scRNA-seq and bulk RNA-seq data analysis, we successfully developed a prognostic prediction model that incorporates two pivotal NKT cells marker genes, namely, CD44 and TNFSF14. This model has exhibited outstanding performance in assessing the prognosis of GBM patients. Furthermore, we conducted a preliminary investigation into the immune microenvironment across various risk groups that contributes to uncover promising immunotherapeutic targets specific to GBM.










Similar content being viewed by others
Data availability
Publicly available datasets were analyzed in this study. This data can be found here: The datasets analyzed in the current study are available in the GEO database (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE163108, GSE163108, n = 5), TCGA database (https://portal.gdc.cancer.gov/, TCGA-GBM cohort, n = 170), CGGA database (http://www.cgga.org.cn/, CGGA.mRNAseq_325, n = 139; CGGA.mRNAseq_693, n = 249), UCSC database (https://xena.ucsc.edu/, GTEX-cohort, Brain-Frontal Cortex (BA9), Brain-Cortex, Brain-Anterior cingulate cortex (BA24), n = 290), TIDE database (http://tide.dfci.harvard.edu/login/), TCIA database (https://www.tcia.at/home), HPA database (https://www.proteinatlas.org/), and CellMarker database (http://xteam.xbio.top/CellMarker/).
References
An Z, Aksoy O, Zheng T, Fan QW, Weiss WA (2018) Epidermal growth factor receptor and EGFRvIII in glioblastoma: signaling pathways and targeted therapies. Oncogene 37(12):1561–1575
Bendelac A, Savage PB, Teyton L (2007) The biology of NKT cells. Annu Rev Immunol 25:297–336
Brettschneider EES, Terabe M (2021) The role of NKT cells in glioblastoma. Cells 10(7):1641
Chai RC, Zhang KN, Chang YZ, Wu F, Liu YQ, Zhao Z, Wang KY, Chang YH, Jiang T, Wang YZ (2019) Systematically characterize the clinical and biological significances of 1p19q genes in 1p/19q non-codeletion glioma. Carcinogenesis 40(10):1229–1239
Chen B, Khodadoust MS, Liu CL, Newman AM, Alizadeh AA (2018) Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol Biol 1711:243–259
Chongsathidkiet P, Jackson C, Koyama S, Loebel F, Cui X, Farber SH, Woroniecka K, Elsamadicy AA, Dechant CA, Kemeny HR et al (2018) Sequestration of T cells in bone marrow in the setting of glioblastoma and other intracranial tumors. Nat Med 24(9):1459–1468
Dashtsoodol N, Shigeura T, Tashiro T, Aihara M, Chikanishi T, Okada H, Hanada K, Sano H, Kurogi A, Taniguchi M (2017) Natural killer T cell-targeted immunotherapy mediating long-term memory responses and strong antitumor activity. Front Immunol 8:1206
Dirven L, Aaronson NK, Heimans JJ, Taphoorn MJ (2014) Health-related quality of life in high-grade glioma patients. Chin J Cancer 33(1):40–45
Dunn GP, Rinne ML, Wykosky J, Genovese G, Quayle SN, Dunn IF, Agarwalla PK, Chheda MG, Campos B, Wang A et al (2012) Emerging insights into the molecular and cellular basis of glioblastoma. Genes Dev 26(8):756–784
Galvez NMS, Bohmwald K, Pacheco GA, Andrade CA, Carreno LJ, Kalergis AM (2021) Type I natural killer T cells as key regulators of the immune response to infectious diseases. Clin Microbiol Rev 34(2):e00232–e00220
Giles DA, Zahner S, Krause P, Van Der Gracht E, Riffelmacher T, Morris V, Tumanov A, Kronenberg M (2018) The tumor necrosis factor superfamily members TNFSF14 (LIGHT), lymphotoxin β and lymphotoxin β receptor interact to regulate intestinal inflammation. Front Immunol 9:2585
Han S, Liu Y, Cai SJ, Qian M, Ding J, Larion M, Gilbert MR, Yang C (2020) IDH mutation in glioma: molecular mechanisms and potential therapeutic targets. Br J Cancer 122(11):1580–1589
Hara A, Koyama-Nasu R, Takami M, Toyoda T, Aoki T, Ihara F, Kobayashi M, Hirono S, Matsutani T, Nakayama T et al (2021) CD1d expression in glioblastoma is a promising target for NKT cell-based cancer immunotherapy. Cancer Immunol Immunother 70(5):1239–1254
Hunn MK, Hermans IF (2013) Exploiting invariant NKT cells to promote T cell responses to cancer vaccines. Oncoimmunology 2(4):e23789
Iser IC, Pereira MB, Lenz G, Wink MR (2017) The epithelial-to-mesenchymal transition-like process in glioblastoma: an updated systematic review and in silico investigation. Med Res Rev 37(2):271–313
Kim M, Yoo J, Chang JH, Kim SH (2022) Association of MGMT gene promoter methylation with clinicopathological parameters in patients with wild-type IDH glioblastoma. Anticancer Res 42(1):335–341
Kim S, Kim A, Shin JY, Seo JS (2020) The tumor immune microenvironmental analysis of 2,033 transcriptomes across 7 cancer types. Sci Rep 10(1):9536
Kim YW, Koul D, Kim SH, Lucio-Eterovic AK, Freire PR, Yao J, Wang J, Almeida JS, Aldape K, Yung WK (2013) Identification of prognostic gene signatures of glioblastoma: a study based on TCGA data analysis. Neuro Oncol 15(7):829–839
Korsunsky I, Millard N, Fan J, Slowikowski K, Zhang F, Wei K, Baglaenko Y, Brenner M, Loh PR, Raychaudhuri S (2019) Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods 16(12):1289–1296
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinf 9:559
Lindau D, Gielen P, Kroesen M, Wesseling P, Adema GJ (2013) The immunosuppressive tumour network: myeloid-derived suppressor cells, regulatory T cells and natural killer T cells. Immunology 138(2):105–115
Litak J, Mazurek M, Grochowski C, Kamieniak P, Roliński J (2019) PD-L1/PD-1 axis in glioblastoma multiforme. Int J Mol Sci 20(21):5347
Liu D, Song L, Brawley VS, Robison N, Wei J, Gao X, Tian G, Margol A, Ahmed N, Asgharzadeh S et al (2013) Medulloblastoma expresses CD1d and can be targeted for immunotherapy with NKT cells. Clin Immunol 149(1):55–64
Liu F, Huang J, Liu X, Cheng Q, Luo C, Liu Z (2020) CTLA-4 correlates with immune and clinical characteristics of glioma. Cancer Cell Int 20:7
Liu X, Li L, Si F, Huang L, Zhao Y, Zhang C, Hoft DF, Peng G (2021) NK and NKT cells have distinct properties and functions in cancer. Oncogene 40(27):4521–4537
Mackay A, Burford A, Molinari V, Jones DTW, Izquierdo E, Brouwer-Visser J, Giangaspero F, Haberler C, Pietsch T, Jacques TS et al (2018) Molecular, pathological, radiological, and immune profiling of non-brainstem pediatric high-grade glioma from the HERBY phase II randomized trial. Cancer Cell 33(5):829–842.e5
Marcus A, Gowen BG, Thompson TW, Iannello A, Ardolino M, Deng W, Wang L, Shifrin N, Raulet DH (2014) Recognition of tumors by the innate immune system and natural killer cells. Adv Immunol 122:91–128
Nair S, Dhodapkar MV (2017) Natural killer T cells in cancer immunotherapy. Front Immunol 8:1178
Nelson A, Lukacs JD, Johnston B (2021) The current landscape of NKT cell immunotherapy and the hills ahead. Cancers 13(20):5174
Ostrom QT, Gittleman H, Truitt G, Boscia A, Kruchko C, Barnholtz-Sloan JS (2018) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the united states in 2011-2015. Neuro Oncol 20(suppl_4):iv1–iv86
Qiao G, Kone LB, Phillips EH, Lee SS, Brown GE, Khetani SR, Thakur A, Lum LG, Prabhakar BS, Maker AV (2022) LIGHT enhanced bispecific antibody armed T-cells to treat immunotherapy resistant colon cancer. Oncogene 41(14):2054–2068
Safarians G, Sohrabi A, Solomon I, Xiao W, Bastola S, Rajput BW, Epperson M, Rosenzweig I, Tamura K, Singer B et al (2023) Glioblastoma spheroid invasion through soft, brain-like matrices depends on hyaluronic acid-CD44 interactions. Adv Healthc Mater 12(14):e2203143
Shu C, Li Q (2020) Current advances in PD-1/PD-L1 axis-related tumour-infiltrating immune cells and therapeutic regimens in glioblastoma. Crit Rev Oncol Hematol 151:102965
Skeate JG, Otsmaa ME, Prins R, Fernandez DJ, Da Silva DM, Kast WM (2020) TNFSF14: LIGHTing the way for effective cancer immunotherapy. Front Immunol 11:922
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn U et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352(10):987–996
Tang B, Wu W, Wei X, Li Y, Ren G, Fan W (2014) Activation of glioma cells generates immune tolerant NKT cells. J Biol Chem 289(50):34595–34600
Terabe M, Berzofsky JA (2018) Tissue-specific roles of NKT cells in tumor immunity. Front Immunol 9:1838
Tibshirani R (1997) The lasso method for variable selection in the Cox model. Stat Med 16(4):385–395
Tupin E, Kinjo Y, Kronenberg M (2007) The unique role of natural killer T cells in the response to microorganisms. Nat Rev Microbiol 5(6):405–417
Wang R, Zhang X, Huang J, Feng K, Zhang Y, Wu J, Ma L, Zhu A, Di L (2023) Bio-fabricated nanodrugs with chemo-immunotherapy to inhibit glioma proliferation and recurrence. J Control Release 354:572–587
Wei SC, Anang NAS, Sharma R, Andrews MC, Reuben A, Levine JH, Cogdill AP, Mancuso JJ, Wargo JA, Pe'er D et al (2019) Combination anti–CTLA-4 plus anti–PD-1 checkpoint blockade utilizes cellular mechanisms partially distinct from monotherapies. Proc Natl Acad Sci U S A 116(45):22699–22709
Wen PY, Huse JT (2017) 2016 World health organization classification of central nervous system tumors. Continuum 23(6, Neuro-oncology):1531–1547
Woroniecka KI, Rhodin KE, Chongsathidkiet P, Keith KA, Fecci PE (2018) T cell dysfunction in glioblastoma: applying a new framework. Clin Cancer Res 24(16):3792–3802
Xu H, Niu M, Yuan X, Wu K, Liu A (2020) CD44 as a tumor biomarker and therapeutic target. Exp Hematol Oncol 9(1):36
Acknowledgements
We sincerely appreciate the great work of GEO, TCGA, and CGGA, as well as other sites such as UCSC, CellMarker, TIDE, TCIA, and HPA for collating the data such that they are easily and quickly available to us.
Funding
This work was funded by the Zhejiang Provincial People’s Hospital Talent Introduction Project (No. C-2021-QDJJ03-01).
Author information
Authors and Affiliations
Contributions
JH and RX conceived and designed the study. LX performed the experiment. JH and WF were responsible for data analysis and checking. YS, NW, and JZ collated the data. CY and XZ were responsible for the resources. YZ, RW, and HZ designed computer programs. RM, XD, and XL developed the methodology. JH wrote the original draft of the manuscript, which was reviewed and revised by RX and SH. RX and SH supervised the study. All authors listed have made a substantial, direct, and intellectual contribution to the work and approved it for publication.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors have agreed to publish this manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 7384 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, J., Xu, L., Fu, W. et al. Development and validation a prognostic model based on natural killer T cells marker genes for predicting prognosis and characterizing immune status in glioblastoma through integrated analysis of single-cell and bulk RNA sequencing. Funct Integr Genomics 23, 286 (2023). https://doi.org/10.1007/s10142-023-01217-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01217-7