Abstract
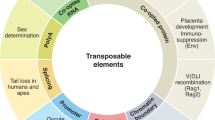
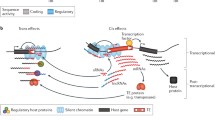
Transposable elements, often referred to as “jumping genes,” have long been recognized as genomic parasites due to their ability to integrate and disrupt normal gene function and induce extensive genomic alterations, thereby compromising the host's fitness. To counteract this, the host has evolved a plethora of mechanisms to suppress the activity of the transposons. Recent research has unveiled the host-transposon relationships to be nuanced and complex phenomena, resulting in the coevolution of both entities. Transposition increases the mutational rate in the host genome, often triggering physiological pathways such as immune and stress responses. Current gene transfer technologies utilizing transposable elements have potential drawbacks, including off-target integration, induction of mutations, and modifications of cellular machinery, which makes an in-depth understanding of the host-transposon relationship imperative. This review highlights the dynamic interplay between the host and transposable elements, encompassing various factors and components of the cellular machinery. We provide a comprehensive discussion of the strategies employed by transposable elements for their propagation, as well as the mechanisms utilized by the host to mitigate their parasitic effects. Additionally, we present an overview of recent research identifying host proteins that act as facilitators or inhibitors of transposition. We further discuss the evolutionary outcomes resulting from the genetic interactions between the host and the transposable elements. Finally, we pose open questions in this field and suggest potential avenues for future research.






Similar content being viewed by others
Data availability
Not applicable.
References
Aravin AA, Hannon GJ, Brennecke J (2007) The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318(5851):761–764. https://doi.org/10.1126/SCIENCE.1146484
Aravin AA, Sachidanandam R, Bourc’his D, Schaefer C, Pezic D, Toth KF, Bestor T, Hannon GJ (2008) A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell 31(6):785–799. https://doi.org/10.1016/J.MOLCEL.2008.09.003
Arnold BJ, Huang IT, Hanage WP (2022) Horizontal gene transfer and adaptive evolution in bacteria. Nat Rev Microbiol 20(4):206–218. https://doi.org/10.1038/s41579-021-00650-4
Asif-Laidin A, Conesa C, Bonnet A, Grison C, Adhya I, Menouni R, Fayol H, Palmic N, Acker J, Lesage P (2020) A small targeting domain in Ty1 integrase is sufficient to direct retrotransposon integration upstream of tRNA genes. EMBO J 39(17). https://doi.org/10.15252/embj.2019104337
Banks JA, Masson P, Fedoroff N (1988) Molecular mechanisms in the developmental regulation of the maize suppressor-mutator transposable element. Genes Dev 2(11):1364–1380. https://doi.org/10.1101/GAD.2.11.1364
Bao W, Kojima KK, Kohany O (2015) Repbase update, a database of repetitive elements in eukaryotic genomes. Mobile DNA 6(1). https://doi.org/10.1186/s13100-015-0041-9
Barrangou R, Marraffini LA (2014) CRISPR-Cas systems: prokaryotes upgrade to adaptive immunity. Mol Cell 54(2):234–244. https://doi.org/10.1016/J.MOLCEL.2014.03.011
Bartels D, Salamini F (2001) Desiccation tolerance in the resurrection plant Craterostigma plantagineum. A contribution to the study of drought tolerance at the molecular level. Plant Physiol 127(4):1346. https://doi.org/10.1104/pp.010765
Behrens R, Hayles J, Nurse P (2000) Fission yeast retrotransposon Tf1 integration is targeted to 5’ ends of open reading frames. Nucleic Acids Res 28(23):4709–4716. https://doi.org/10.1093/NAR/28.23.4709
Belancio VP, Hedges DJ, Deininger P (2006) LINE-1 RNA splicing and influences on mammalian gene expression. Nucleic Acids Res 34(5):1512–1521. https://doi.org/10.1093/NAR/GKL027
Bellen HJ, Levis RW, He Y, Carlson JW, Evans-Holm M, Bae E, Kim J, Metaxakis A, Savakis C, Schulze KL, Hoskins RA, Spradling AC (2011) The Drosophila gene disruption project: progress using transposons with distinctive site specificities. Genetics 188(3):731–743. https://doi.org/10.1534/GENETICS.111.126995
Bernatavichute YV, Zhang X, Cokus S, Pellegrini M, Jacobsen SE (2008) Genome-wide association of histone H3 lysine nine methylation with CHG DNA methylation in Arabidopsis thaliana. PloS One 3(9). https://doi.org/10.1371/JOURNAL.PONE.0003156
Berretta J, Pinskaya M, Morillon A (2008) A cryptic unstable transcript mediates transcriptional trans-silencing of the Ty1 retrotransposon in S. cerevisiae. Genes Dev 22(5). https://doi.org/10.1101/gad.458008
Bestor TH, Bourc’his D (2004) Transposon silencing and imprint establishment in mammalian germ cells. Cold Spring Harb Symp Quant Biol 69:381–387. https://doi.org/10.1101/SQB.2004.69.381
Biessmann H, Valgeirsdottir K, Lofsky A, Chin C, Ginther B, Levis RW, Pardue ML (1992) HeT-A, a transposable element specifically involved in “healing” broken chromosome ends in Drosophila melanogaster. Mol Cell Biol 12(9):3910–3918. https://doi.org/10.1128/MCB.12.9.3910-3918.1992
Bire S, Casteret S, Arnaoty A, Piégu B, Lecomte T, Bigot Y (2013) Transposase concentration controls transposition activity: myth or reality? Gene 530(2):165–171. https://doi.org/10.1016/j.gene.2013.08.039
Bire S, Casteret S, Piégu B, Beauclair L, Moiré N, Arensbuger P, Bigot Y (2016) mariner transposons contain a silencer: possible role of the polycomb repressive complex 2. PLoS Genet 12(3):e1005902. https://doi.org/10.1371/journal.pgen.1005902
Bonnet A, Lesage P (2021) Light and shadow on the mechanisms of integration site selection in yeast Ty retrotransposon families. Curr Genet 67(3):347–357. https://doi.org/10.1007/s00294-021-01154-7
Bostick M, Jong KK, Estève PO, Clark A, Pradhan S, Jacobsen SE (2007) UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science 317(5845):1760–1764. https://doi.org/10.1126/SCIENCE.1147939
Botelho J, Grosso F, Peixe L (2019) Antibiotic resistance in Pseudomonas aeruginosa - mechanisms, epidemiology and evolution. Drug Resist Updat 2019(44):100640. https://doi.org/10.1016/J.DRUP.2019.07.002
Bourque G, Burns KH, Gehring M, Gorbunova V, Seluanov A, Hammell M, Imbeault M, Izsvák Z, Levin HL, Macfarlan TS, Mager DL, Feschotte C (2018) Ten things you should know about transposable elements. Genome Biol 19(1). https://doi.org/10.1186/s13059-018-1577-z
Bouttier M, Laperriere D, Memari B, Mangiapane J, Fiore A, Mitchell E, Verway M, Behr MA, Sladek R, Barreiro LB, Mader S, White JH (2016) Alu repeats as transcriptional regulatory platforms in macrophage responses to M. tuberculosis infection. Nucleic Acids Res 44(22):10571–10587. https://doi.org/10.1093/NAR/GKW782
Bouuaert CC, Lipkow K, Andrews SS, Liu D, Chalmers R (2013) The autoregulation of a eukaryotic DNA transposon. ELife 2:e00668. https://doi.org/10.7554/eLife.00668
Bouuaert CC, Tellier M, Chalmers R (2014) One to rule them all. Mobil Genet Elements 4(1):e28807. https://doi.org/10.4161/mge.28807
Braam LAM, Goryshin IY, Reznikoff WS (1999) A mechanism for Tn5 inhibition. J Biol Chem 274:1. https://doi.org/10.1074/jbc.274.1.86
Brennecke J, Aravin AA, Stark A, Dus M, Kellis M, Sachidanandam R, Hannon GJ (2007) Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128(6):1089–1103. https://doi.org/10.1016/J.CELL.2007.01.043
Bridier-Nahmias A, Tchalikian-Cosson A, Baller JA, Menouni R, Fayol H, Flores A, Saïb A, Werner M, Voytas DF, Lesage P (2015) An RNA polymerase III subunit determines sites of retrotransposon integration. Science 348(6234). https://doi.org/10.1126/science.1259114
Broecker F, Moelling K (2019) Evolution of immune systems from viruses and transposable elements. Front Microbiol 10(JAN):51. https://doi.org/10.3389/FMICB.2019.00051/BIBTEX
Brutnell TP (2002) Transposon tagging in maize. Funct Integr Genomics 2:4–12. https://doi.org/10.1007/s10142-001-0044-0
Callinan PA, Batzer MA (2006) Retrotransposable elements and human disease. Genome Dyn 1:104–115. https://doi.org/10.1159/000092503
Campos-Sánchez R, Cremona MA, Pini A, Chiaromonte F, Makova KD (2016) Integration and fixation preferences of human and mouse endogenous retroviruses uncovered with functional data analysis. PLoS Comput Biol. 12(6). https://doi.org/10.1371/journal.pcbi.1004956
Cao J, Yu T, Xu B, Hu Z, Zhang X, Theurkauf WE, Weng Z (2023) Epigenetic and chromosomal features drive transposon insertion in Drosophila melanogaster. Nucleic Acids Res 51(5):2066–2086. https://doi.org/10.1093/nar/gkad054
Capy P (2021) Taming, domestication and exaptation: trajectories of transposable elements in genomes. Cells 10(12). https://doi.org/10.3390/CELLS10123590
Carmona LM, Schatz DG (2017) New insights into the evolutionary origins of the recombination-activating gene proteins and V(D)J recombination. FEBS J. 284(11):1590–1605. https://doi.org/10.1111/febs.13990
Chandler V, Alleman M (2008) Anecdotal, historical and critical commentaries on genetics: paramutation: epigenetic instructions passed across generations. Genetics 178(4):1839. https://doi.org/10.1093/GENETICS/178.4.1839
Chandler M, Mahillon J (2007) Insertion sequences revisited. Mobile DNA II 303–366. https://doi.org/10.1128/9781555817954.CH15
Chandler VL, Walbot V (1986) DNA modification of a maize transposable element correlates with loss of activity. Proc Natl Acad Sci USA 83(6):1767–1771. https://doi.org/10.1073/PNAS.83.6.1767
Chandler M, de La Cruz F, Dyda F, Hickman AB, Moncalian G, Ton-Hoang B (2013) Breaking and joining single-stranded DNA: the HUH endonuclease superfamily. Nat Rev Microbiol 11(8):525–538. https://doi.org/10.1038/NRMICRO3067
Chatterjee AG, Leem YE, Kelly FD, Levin HL (2009) The chromodomain of Tf1 integrase promotes binding to cDNA and mediates target site selection. J Virol 83(6). https://doi.org/10.1128/jvi.01588-08
Chen J, Greenblatt IM, Dellaporta SL (1992) Molecular analysis of Ac transposition and DNA replication. Genetics 130(3):665–676. https://doi.org/10.1093/GENETICS/130.3.665
Chen JM, Chuzhanova N, Stenson PD, Férec C, Cooper DN (2005) Meta-Analysis of gross insertions causing human genetic disease: novel mutational mechanisms and the role of replication slippage. Hum Mutat 25(2):207–221. https://doi.org/10.1002/HUMU.20133
Chénais B (2022) Transposable elements and human diseases: mechanisms and implication in the response to environmental pollutants. Int J Mol Sci 23(5):2551. https://doi.org/10.3390/IJMS23052551
Chiu YL, Greene WC (2008) The APOBEC3 cytidine deaminases: an innate defensive network opposing exogenous retroviruses and endogenous retroelements. Annu Rev Immunol 26:317–353. https://doi.org/10.1146/ANNUREV.IMMUNOL.26.021607.090350
Chomet PS, Wessler S, Dellaporta SL (1987) Inactivation of the maize transposable element activator (Ac) is associated with its DNA modification. EMBO J 6(2):295–302. https://doi.org/10.1002/J.1460-2075.1987.TB04753.X
Chou HH, Hayakawa T, Diaz S, Krings M, Indriati E, Leakey M, Paabo S, Satta Y, Takahata N, Varki A (2002) Inactivation of CMP-N-acetylneuraminic acid hydroxylase occurred prior to brain expansion during human evolution. Proc Natl Acad Sci USA 99(18):11736–11741. https://doi.org/10.1073/PNAS.182257399
Chung H, Bogwitz MR, McCart C, Andrianopoulos A, Ffrench-Constant RH, Batterham P, Daborn PJ (2007) Cis-regulatory elements in the Accord retrotransposon result in tissue-specific expression of the Drosophila melanogaster insecticide resistance gene Cyp6g1. Genetics 175(3):1071–1077. https://doi.org/10.1534/GENETICS.106.066597
Chuong EB, Elde NC, Feschotte C (2016) Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 351(6277):1083–1087. https://doi.org/10.1126/SCIENCE.AAD5497
Cordaux R, Batzer MA (2009) The impact of retrotransposons on human genome evolution. Nat Rev Genet 10(10):691. https://doi.org/10.1038/NRG2640
Cordaux R, Udit S, Batzer MA, Feschotte C (2006) Birth of a chimeric primate gene by capture of the transposase gene from a mobile element. Proc Natl Acad Sci USA 103(21):8101–8106. https://doi.org/10.1073/PNAS.0601161103
Cowley M, Oakey RJ (2013) Transposable elements re-wire and fine-tune the transcriptome. PLoS Genet 9(1). https://doi.org/10.1371/JOURNAL.PGEN.1003234
Curcio MJ, Derbyshire KM (2003) The outs and ins of transposition: from MU to kangaroo. In Nat Rev Mol Cell Biol 4(11):865–877. https://doi.org/10.1038/nrm1241
Czech B, Hannon GJ (2016) One loop to rule them all: the ping-pong cycle and piRNA-guided silencing. Trends Biochem Sci 41(4):324–337. https://doi.org/10.1016/J.TIBS.2015.12.008
Dai J, Xie W, Brady TL, Gao J, Voytas DF (2007) Phosphorylation regulates integration of the yeast Ty5 retrotransposon into heterochromatin. Mol Cell 27(2). https://doi.org/10.1016/j.molcel.2007.06.010
Debyser Z, Christ F, de Rijck J, Gijsbers R (2015) Host factors for retroviral integration site selection. Trends Biochem Sci 40(2):108–116. https://doi.org/10.1016/J.TIBS.2014.12.001
Deininger PL, Batzer MA (1999) Alu repeats and human disease. Mol Genet Metab 67(3):183–193. https://doi.org/10.1006/MGME.1999.2864
Deng W, Lin H (2002) miwi, a murine homolog of piwi, encodes a cytoplasmic protein essential for spermatogenesis. Dev Cell 2(6):819–830. https://doi.org/10.1016/S1534-5807(02)00165-X
Devine SE, Boeke JD (1996) Integration of the yeast retrotransposon Ty1 is targeted to regions upstream of genes transcribed by RNA polymerase III. Genes Dev 10(5):620–633. https://doi.org/10.1101/GAD.10.5.620
Díaz-Maldonado H, Gómez MJ, Moreno-Paz M, Martín-Úriz PS, Amils R, Parro V, de Saro FJL (2015) Transposase interaction with the β sliding clamp: effects on insertion sequence proliferation and transposition rate. Sci Rep 5. https://doi.org/10.1038/SREP13329
Dimitriu T, Szczelkun MD, Westra ER (2020) Evolutionary Ecology and Interplay of Prokaryotic Innate and Adaptive Immune Systems. Curr Biol. 30(19):R1189–R1202. https://doi.org/10.1016/j.cub.2020.08.028
Duval-Valentin G, Chandler M (2011) Cotranslational control of DNA transposition: a window of opportunity. Mol Cell 44(6):989–996. https://doi.org/10.1016/j.molcel.2011.09.027
Eickbush TH, Jamburuthugoda VK (2008) The diversity of retrotransposons and the properties of their reverse transcriptases. Virus Res 134(1–2):221–234. https://doi.org/10.1016/j.virusres.2007.12.010
Ellis MJ, Trussler RS, Haniford DB (2015) Hfq binds directly to the ribosome-binding site of IS10 transposase mRNA to inhibit translation. Mol Microbiol 96:633–650. https://doi.org/10.1111/mmi.12961
Feschotte C, Pritham EJ (2007) DNA transposons and the evolution of eukaryotic genomes. Annu Rev Genet 41:331–368. https://doi.org/10.1146/ANNUREV.GENET.40.110405.090448
Feschotte C, Wessler SR (2001) Treasures in the attic: rolling circle transposons discovered in eukaryotic genomes. Proc Natl Acad Sci USA 98(16):8923–8924. https://doi.org/10.1073/PNAS.171326198
Fineran PC, Charpentier E (2012) Memory of viral infections by CRISPR-Cas adaptive immune systems: acquisition of new information. Virology 434(2):202–209. https://doi.org/10.1016/J.VIROL.2012.10.003
Finnegan DJ (1989) Eukaryotic transposable elements and genome evolution. Trends Genet 5(4):103–107. https://doi.org/10.1016/0168-9525(89)90039-5
Fraser MJ, Ciszczon T, Elick T, Bauser C (1996) Precise excision of TTAA-specific lepidopteran transposons piggyBac (IFP2) and tagalong (TFP3) from the baculovirus genome in cell lines from two species of Lepidoptera. Insect Mol Biol 5(2):141–151. https://doi.org/10.1111/J.1365-2583.1996.TB00048.X
Fugmann SD (2010) The origins of the Rag genes–from transposition to V(D)J recombination. Semin Immunol 22(1):10–16. https://doi.org/10.1016/J.SMIM.2009.11.004
Fujiwara H, Osanai M, Matsumoto T, Kojima KK (2005) Telomere-specific non-LTR retrotransposons and telomere maintenance in the silkworm. Bombyx Mori Chromosome Res 13(5):455–467. https://doi.org/10.1007/S10577-005-0990-9
Ganko EW, Fielman KT, McDonald JF (2001) Evolutionary history of Cer elements and their impact on the C. elegans genome. Genome Res 11(12):2066–2074. https://doi.org/10.1101/gr.196201
Gao X, Hou Y, Ebina H, Levin HL, Voytas DF (2008) Chromodomains direct integration of retrotransposons to heterochromatin. Genome Res 18(3). https://doi.org/10.1101/gr.7146408
Garcillán-Barcia MP, Bernales I, Mendiola MV, de la Cruz F (2007) IS91 rolling-circle transposition. Mob DNA II 889–904. https://doi.org/10.1128/9781555817954.CH37
George JA, Traverse KL, DeBaryshe PG, Kelley KJ, Pardue ML (2010) Evolution of diverse mechanisms for protecting chromosome ends by Drosophila TART telomere retrotransposons. Proc Natl Acad Sci USA 107(49):21052–21057. https://doi.org/10.1073/PNAS.1015926107
Ghildiyal M, Zamore PD (2009) Small silencing RNAs: an expanding universe. Nat Rev Genet 10(2):94–108. https://doi.org/10.1038/NRG2504
Gilbert C, Cordaux R (2017) Viruses as vectors of horizontal transfer of genetic material in eukaryotes. Curr Opin Virol 25:16–22. https://doi.org/10.1016/j.coviro.2017.06.005
Girard A, Sachidanandam R, Hannon GJ, Carmell MA (2006) A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442(7099):199–202. https://doi.org/10.1038/NATURE04917
Gómez MJ, Díaz-Maldonado H, González-Tortuero E, De Saro FJL (2014) Chromosomal replication dynamics and interaction with the β sliding clamp determine orientation of bacterial transposable elements. Genome Biol Evol 6(3). https://doi.org/10.1093/gbe/evu052
Goodier JL, Cheung LE, Kazazian HH Jr (2013) Mapping the LINE1 ORF1 protein interactome reveals associated inhibitors of human retrotransposition. Nucleic Acids Res 41(15):7401–7419. https://doi.org/10.1093/nar/gkt512
Grabundzija I, Messing SA, Thomas J, Cosby RL, Bilic I, Miskey C, Gogol-Döring A, Kapitonov V, Diem T, Dalda A et al (2016) A Helitron transposon reconstructed from bats reveals a novel mechanism of genome shuffling in eukaryotes. Nat Commun 7:10716. https://doi.org/10.1038/ncomms10716
Grabundzija I, Hickman AB, Dyda F (2018) Helraiser intermediates provide insight into the mechanism of eukaryotic replicative transposition. Nat Commun 9:1278. https://doi.org/10.1038/s41467-018-03688-w
Grivna ST, Beyret E, Wang Z, Lin H (2006) A novel class of small RNAs in mouse spermatogenic cells. Genes Dev 20(13):1709–1714. https://doi.org/10.1101/GAD.1434406
Grognet P, Lalucque H, Malagnac F, Silar P (2014) Genes that bias Mendelian segregation. PLoS Genet 10(5):e1004387. https://doi.org/10.1371/journal.pgen.1004387
Guio L, Barrõn MG, González J (2014) The transposable element Bari-Jheh mediates oxidative stress response in Drosophila. Mol Ecol 23(8):2020–2030. https://doi.org/10.1111/MEC.12711
Gunawardane LS, Saito K, Nishida KM, Miyoshi K, Kawamura Y, Nagami T, Siomi H, Siomi MC (2007) A slicer-mediated mechanism for repeat-associated siRNA 5’ end formation in Drosophila. Science 315(5818):1587–1590. https://doi.org/10.1126/SCIENCE.1140494
Guo H, Chitiprolu M, Gagnon D, Meng L, Perez-Iratxeta C, Lagace D, Gibbings D (2014) Autophagy supports genomic stability by degrading retrotransposon RNA. Nat Commun 5(1):1–11. https://doi.org/10.1038/ncomms6276
Guo Z, Kuang Z, Tao Y et al (2022) Miniature inverted-repeat transposable elements drive rapid MicroRNA diversification in angiosperms. Mol Biol Evol. 39(11):224. https://doi.org/10.1093/molbev/msac224
Haase AD, Fenoglio S, Muerdter F, Guzzardo PM, Czech B, Pappin DJ, Chen C, Gordon A, Hannon GJ (2010) Probing the initiation and effector phases of the somatic piRNA pathway in Drosophila. Genes Dev 24(22):2499–2504. https://doi.org/10.1101/GAD.1968110
Hacein-Bey-Abina S, von Kalle C, Schmidt M, McCormack MP, Wulffraat N, Leboulch P, Lim A, Osborne CS, Pawliuk R, Morillon E, Sorensen R, Forster A, Fraser P, Cohen JI, de Saint Basile G, Alexander I, Wintergerst U, Frebourg T, Aurias A, … Cavazzana-Calvo M (2003) LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science 302(5644):415–419. https://doi.org/10.1126/SCIENCE.1088547
Halling KC, Lazzaro CR, Honchel R, Bufill JA, Powell SM, Arndt CAS, Lindor NM (1999) Hereditary desmoid disease in a family with a germline Alu I repeat mutation of the APC gene. Hum Hered 49(2):97–102. https://doi.org/10.1159/000022852
Han BW, Wang W, Li C, Weng Z, Zamore PD (2015) Noncoding RNA. piRNA-guided transposon cleavage initiates Zucchini-dependent, phased piRNA production. Science 348(6236):817–821. https://doi.org/10.1126/SCIENCE.AAA1264
Haudiquet M, de Sousa JM, Touchon M, Rocha EPC (2022) Selfish, promiscuous and sometimes useful: how mobile genetic elements drive horizontal gene transfer in microbial populations. Philos Trans R Soc Lond B Biol Sci 377(1861):20210234. https://doi.org/10.1098/rstb.2021.0234
Hedges DJ, Deininger PL (2007) Inviting instability: transposable elements, double-strand breaks, and the maintenance of genome integrity. Mutat Res 616(1–2):46–59. https://doi.org/10.1016/j.mrfmmm.2006.11.021
Heringer P, Kuhn GCS (2022) Pif1 helicases and the evidence for a prokaryotic origin of helitrons. Mol Biol Evol 39(1). https://doi.org/10.1093/molbev/msab334
Hickman AB, Dyda F (2014) CRISPR-Cas immunity and mobile DNA: a new superfamily of DNA transposons encoding a Cas1 endonuclease. Mob DNA 5(1):1–4. https://doi.org/10.1186/1759-8753-5-23/FIGURES/1
Hilbricht T, Varotto S, Sgaramella V, Bartels D, Salamini F, Furini A (2008) Retrotransposons and siRNA have a role in the evolution of desiccation tolerance leading to resurrection of the plant Craterostigma plantagineum. New Phytol 179(3):877–887. https://doi.org/10.1111/J.1469-8137.2008.02480.X
Hirochika H, Okamoto H, Kakutani T (2000) Silencing of retrotransposons in Arabidopsis and reactivation by the ddm1 mutation. Plant Cell 12(3):357–368. https://doi.org/10.1105/TPC.12.3.357
Ho KL, Ma L, Cheung S, Manhas S, Fang N, Wang K, Young B, Loewen C, Mayor T, Measday V (2015) A role for the budding yeast separase, Esp1, in Ty1 Element Retrotransposition. PLoS Genet 11(3). https://doi.org/10.1371/journal.pgen.1005109
Horie K, Yusa K, Yae K, Odajima J, Fischer SEJ, Keng VW, Hayakawa T, Mizuno S, Kondoh G, Ijiri T, Matsuda Y, Plasterk RHA, Takeda J (2003) Characterization of sleeping beauty transposition and its application to genetic screening in mice. Mol Cell Biol 23(24):9189–9207. https://doi.org/10.1128/MCB.23.24.9189-9207.2003
Huang L, Sun Q, Qin F, Li C, Zhao Y, Zhou DX (2007) Down-regulation of a silent information regulator2-related histone deacetylase gene, OsSRT1, induces DNA fragmentation and cell death in rice. Plant Physiol 144(3):1508–1519. https://doi.org/10.1104/PP.107.099473
Hummel B, Hansen EC, Yoveva A, Aprile-Garcia F, Hussong R, Sawarkar R (2017) The evolutionary capacitor HSP90 buffers the regulatory effects of mammalian endogenous retroviruses. Nat Struct Mol Biol 24(3):234–242. https://doi.org/10.1038/NSMB.3368
Inoue Y, Saga T, Aikawa T, Kumagai M, Shimada A, Kawaguchi Y, Naruse K, Morishita S, Koga A, Takeda H (2017) Complete fusion of a transposon and herpesvirus created the Teratorn mobile element in medaka fish. Nat Commun 8(1):551. https://doi.org/10.1038/s41467-017-00527-2
Inoue Y, Kumagai M, Zhang X, Saga T, Wang D, Koga A, Takeda H (2018) Fusion of piggyBac-like transposons and herpesviruses occurs frequently in teleosts. Zool Lett 4(1):6. https://doi.org/10.1186/s40851-018-0089-8
Irwin NAT, Pittis AA, Richards TA, Keeling PJ (2021) Systematic evaluation of horizontal gene transfer between eukaryotes and viruses. Nat Microbiol 7(2):327–336. https://doi.org/10.1038/s41564-021-01026-3
Jacob Y, Feng S, LeBlanc CA, Bernatavichute YV, Stroud H, Cokus S, Johnson LM, Pellegrini M, Jacobsen SE, Michaels SD (2009) ATXR5 and ATXR6 are H3K27 monomethyltransferases required for chromatin structure and gene silencing. Nat Struct Mol Biol 16(7):763–768. https://doi.org/10.1038/NSMB.1611
Jangam D, Feschotte C, Betrán E (2017) Transposable element domestication as an adaptation to evolutionary conflicts. Trends Genet: TIG 33(11):817–831. https://doi.org/10.1016/J.TIG.2017.07.011
Jin Y, Chen Y, Zhao S, Guan KL, Zhuang Y, Zhou W, Wu X, Xu T (2017) DNA-PK facilitates piggyBac transposition by promoting paired-end complex formation. Proc Natl Acad Sci USA 114(28):7408–7413. https://doi.org/10.1073/PNAS.1612980114
Jung D, Alt FW (2004) Unraveling V(D)J Recombination: insights into gene regulation. Cell 116(2):299–311. https://doi.org/10.1016/S0092-8674(04)00039-X
Jurka J, Kapitonov VV, Pavlicek A, Klonowski P, Kohany O, Walichiewicz J (2005) Repbase update, a database of eukaryotic repetitive elements. Cytogenet Genome Res 110(1–4). https://doi.org/10.1159/000084979
Kapitonov VV, Jurka J (2001) Rolling-circle transposons in eukaryotes. Proc Natl Acad Sci USA 98(15):8714–8719. https://doi.org/10.1073/PNAS.151269298
Kapitonov VV, Jurka J (2005) RAG1 core and V(D)J recombination signal sequences were derived from Transib transposons. PLoS Biol 3(6):0998–1011. https://doi.org/10.1371/JOURNAL.PBIO.0030181
Kapitonov VV, Jurka J (2006) Self-synthesizing DNA transposons in eukaryotes. Proc Natl Acad Sci USA 103(12):4540–4545. https://doi.org/10.1073/PNAS.0600833103/SUPPL_FILE/00833TABLE1.PDF
Kapitonov VV, Koonin EV (2015) Evolution of the RAG1-RAG2 locus: both proteins came from the same transposon. Biol Direct 10(1). https://doi.org/10.1186/S13062-015-0055-8
Kawakami K, Largaespada DA, Ivics Z (2017) Transposons as tools for functional genomics in vertebrate models. Trends Genet: TIG 33(11):784–801. https://doi.org/10.1016/j.tig.2017.07.006
Kazazian HH (2004) Mobile elements: drivers of genome evolution. Science 303(5664):1626–1632. https://doi.org/10.1126/science.1089670
Kent TV, Uzunović J, Wright SI (2017) Coevolution between transposable elements and recombination. Philos Trans R Soc Lond B Biol Sci 372(1736):20160458. https://doi.org/10.1098/rstb.2016.0458
Kleckner N (1990) Regulating TN10 and IS10 transposition. Genetics 124(3). https://doi.org/10.1093/genetics/124.3.449
Klompe SE, Vo PLH, Halpin-Healy TS, Sternberg SH (2019) Transposon-encoded CRISPR–Cas systems direct RNA-guided DNA integration. Nature 571(7764):219–225. https://doi.org/10.1038/s41586-019-1323-z
Kosek D, Grabundzija I, Lei H, Bilic I, Wang H, Jin Y, Peaslee GF, Hickman AB, Dyda F (2021) The large bat Helitron DNA transposase forms a compact monomeric assembly that buries and protects its covalently bound 5′-transposon end. Mol Cell 81(20):4271-4286.e4. https://doi.org/10.1016/j.molcel.2021.07.028
Krupovic M, Koonin EV (2015) Polintons: a hotbed of eukaryotic virus, transposon and plasmid evolution. Nat Rev Microbiol 13(2):105–115. https://doi.org/10.1038/nrmicro3389
Krupovic M, Koonin EV (2016) Self-synthesizing transposons: unexpected key players in the evolution of viruses and defense systems. Curr Opin Microbiol 31:25–33. https://doi.org/10.1016/J.MIB.2016.01.006
Krupovic M, Makarova KS, Forterre P, Prangishvili D, Koonin EV (2014) Casposons: a new superfamily of self-synthesizing DNA transposons at the origin of prokaryotic CRISPR-Cas immunity. BMC Biol 12. https://doi.org/10.1186/1741-7007-12-36
Ku HY, Lin H (2014) PIWI proteins and their interactors in piRNA biogenesis, germline development and gene expression. Natl Sci Rev 1(2):205–218. https://doi.org/10.1093/NSR/NWU014
Kuduvalli PN, Rao JE, Craig NL (2001) Target DNA structure plays a critical role in Tn7 transposition. EMBO J 20(4):924–932. https://doi.org/10.1093/EMBOJ/20.4.924
Labrie SJ, Samson JE, Moineau S (2010) Bacteriophage resistance mechanisms. Nat Rev Microbiol 8(5):317–327. https://doi.org/10.1038/NRMICRO2315
Langley CH, Montgomery E, Hudson R, Kaplan N, Charlesworth B (1988) On the role of unequal exchange in the containment of transposable element copy number. Genet Res 52(3):223–235. https://doi.org/10.1017/S0016672300027695
le Rouzic A, Capy P (2005) The first steps of transposable elements invasion: parasitic strategy vs. genetic drift. Genetics 169(2):1033–1043. https://doi.org/10.1534/GENETICS.104.031211
Lee YCG (2015) The role of piRNA-mediated epigenetic silencing in the population dynamics of transposable elements in Drosophila melanogaster. PLoS Genetics 11(6). https://doi.org/10.1371/JOURNAL.PGEN.1005269
Lee YCG, Ogiyama Y, Martins NMC, Beliveau BJ, Acevedo D, Wu CT, Cavalli G, Karpen GH (2020) Pericentromeric heterochromatin is hierarchically organized and spatially contacts H3K9me2 islands in euchromatin. PLOS Genet 16(3):e1008673. https://doi.org/10.1371/JOURNAL.PGEN.1008673
Leem YE, Ripmaster TL, Kelly FD, Ebina H, Heincelman ME, Zhang K, Grewal SIS, Hoffman CS, Levin HL (2008) Retrotransposon Tf1 is targeted to Pol II promoters by transcription activators. Mol Cell 30(1):98–107. https://doi.org/10.1016/J.MOLCEL.2008.02.016
Lesage P, Todeschini AL (2005) Happy together: the life and times of Ty retrotransposons and their hosts. Cytogenet Genome Res 110(1–4):70–90. https://doi.org/10.1159/000084940
Levin H, Moran J (2011) Dynamic interactions between transposable elements and their hosts. Nat Rev Genet 12(9):615–627. https://doi.org/10.1038/nrg3030
Levine B, Kroemer G (2008) Autophagy in the pathogenesis of disease. Cell 132(1):27–42. https://doi.org/10.1016/J.CELL.2007.12.018
Lindroth AM, Shultis D, Jasencakova Z, Fuchs J, Johnson L, Schubert D, Patnaik D, Pradhan S, Goodrich J, Schubert I, Jenuwein T, Khorasanizadeh S, Jacobsen SE (2004) Dual histone H3 methylation marks at lysines 9 and 27 required for interaction with CHROMOMETHYLASE3. EMBO J 23(21):4146–4155. https://doi.org/10.1038/SJ.EMBOJ.7600430
Lipkow K, Buisine N, Chalmers R (2004) Promiscuous target interactions in the mariner transposon himar1. J Biol Chem 279(47). https://doi.org/10.1074/jbc.M408759200
Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo QM, Edsall L, Antosiewicz-Bourget J, Stewart R, Ruotti V, Millar AH, Thomson JA, Ren B, Ecker JR (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462(7271):315–322. https://doi.org/10.1038/NATURE08514
Liu S, Yu Y, Ruan Y, Meyer D, Wolff M, Xu L, Wang N, Steinmetz A, Shen WH (2007) Plant SET- and RING-associated domain proteins in heterochromatinization. Plant J 52(5):914–926. https://doi.org/10.1111/J.1365-313X.2007.03286.X
Liu HN, Pei MS, Ampomah-Dwamena C et al (2023) Genome-wide characterization of long terminal repeat retrotransposons provides insights into trait evolution of four cucurbit species. Funct Integr Genomics 23:218. https://doi.org/10.1007/s10142-023-01128-7
Liu C, Yang Y, Schatz DG (2019) Structures of a RAG-like transposase during cut-and-paste transposition. Nature. 575(7783):540–544. https://doi.org/10.1038/s41586-019-1753-7
Locke J, Kotarski MA, Tartof KD (1988) Dosage-dependent modifiers of position effect variegation in Drosophila and a mass action model that explains their effect. Genetics 120(1):181–198. https://doi.org/10.1093/GENETICS/120.1.181
Lohe AR, Hartl DL (1996) Autoregulation of mariner transposase activity by overproduction and dominant-negative complementation. Mol Biol Evol 13(4):549–555. https://doi.org/10.1093/OXFORDJOURNALS.MOLBEV.A025615
Lu C, Chen J, Zhang Y, Hu Q, Su W, Kuang H (2012) miniature inverted–repeat transposable elements (MITEs) have been accumulated through amplification bursts and play important roles in gene expression and species diversity in Oryza sativa. Mol Biol Evol 29(3):1005. https://doi.org/10.1093/MOLBEV/MSR282
Mahmoud M, Zhou Z, Kaur R et al (2022) Toward the development of Ac/Ds transposon-mediated gene tagging system for functional genomics in oat (Avena sativa L.). Funct Integr Genomics 22:669–681. https://doi.org/10.1007/s10142-022-00861-9
Makarevitch I, Waters AJ, West PT, Stitzer M, Hirsch CN, Ross-Ibarra J, Springer NM (2015) Transposable elements contribute to activation of maize genes in response to abiotic stress. PLoS Genet 11(1). https://doi.org/10.1371/JOURNAL.PGEN.1004915
Malone CD, Hannon GJ (2009) Small RNAs as guardians of the genome. Cell 136(4):656–668. https://doi.org/10.1016/J.CELL.2009.01.045
Mao Z, Hine C, Tian X, van Meter M, Au M, Vaidya A, Seluanov A, Gorbunova V (2011) SIRT6 promotes DNA repair under stress by activating PARP1. Science (New York, N.Y.) 332(6036):1443–1446. https://doi.org/10.1126/SCIENCE.1202723
Mao H, Wang H, Liu S, Li Z, Yang X, Yan J, Li J, Tran LSP, Qin F (2015) A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat Commun 6. https://doi.org/10.1038/NCOMMS9326
Market E, Papavasiliou FN (2003) V(D)J recombination and the evolution of the adaptive immune system. PLoS Biol 1(1). https://doi.org/10.1371/JOURNAL.PBIO.0000016
Mateo L, Ullastres A, González J (2014) A transposable element insertion confers xenobiotic resistance in Drosophila. PLoS Genet 10(8):1. https://doi.org/10.1371/JOURNAL.PGEN.1004560
Mathieu O, Probst AV, Paszkowski J (2005) Distinct regulation of histone H3 methylation at lysines 27 and 9 by CpG methylation in Arabidopsis. EMBO J 24(15):2783–2791. https://doi.org/10.1038/SJ.EMBOJ.7600743
Matsuda E, Garfinkel DJ (2009) Posttranslational interference of Ty1 retrotransposition by antisense RNAs. Proc Natl Acad Sci USA 106(37):1. https://doi.org/10.1073/pnas.0908305106
Matsui T, Leung D, Miyashita H, Maksakova IA, Miyachi H, Kimura H, Tachibana M, Lorincz MC, Shinkai Y (2010) Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET. Nature 464(7290):927–931. https://doi.org/10.1038/NATURE08858
McCue AD, Nuthikattu S, Reeder SH, Slotkin RK (2012) Gene expression and stress response mediated by the epigenetic regulation of a transposable element small RNA. PLoS Genet 8(2):1. https://doi.org/10.1371/JOURNAL.PGEN.1002474
McVean G (2010) What drives recombination hotspots to repeat DNA in humans? Philos Trans R Soc B: Biol Sci 365:(Issue 1544). https://doi.org/10.1098/rstb.2009.0299
Meir YJJ, Weirauch MT, Yang HS, Chung PC, Yu RK, Wu SCY (2011) Genome-wide target profiling of piggyBac and Tol2 in HEK 293: pros and cons for gene discovery and gene therapy. BMC Biotechnol 11. https://doi.org/10.1186/1472-6750-11-28
Merenciano M, Ullastres A, de Cara MAR, Barrón MG, González J (2016) Multiple independent retroelement insertions in the promoter of a stress response gene have variable molecular and functional effects in drosophila. PLoS Genet 12(8):1. https://doi.org/10.1371/JOURNAL.PGEN.1006249
Michishita E, McCord RA, Berber E, Kioi M, Padilla-Nash H, Damian M, Cheung P, Kusumoto R, Kawahara TLA, Barrett JC, Chang HY, Bohr VA, Ried T, Gozani O, Chua KF (2008) SIRT6 is a histone H3 lysine 9 deacetylase that modulates telomeric chromatin. Nature 452(7186):492–496. https://doi.org/10.1038/NATURE06736
Modzelewski AJ, Gan Chong J, Wang T, He L (2022) Mammalian genome innovation through transposon domestication. Nat Cell Biol 24(9):1332–1340. https://doi.org/10.1038/s41556-022-00970-4
Mohn F, Handler D, Brennecke J (2015) Noncoding RNA. piRNA-guided slicing specifies transcripts for Zucchini-dependent, phased piRNA biogenesis. Science (New York, N.Y.) 348(6236):812–817. https://doi.org/10.1126/SCIENCE.AAA1039
Montgomery EA, Huang SM, Langley CH, Judd BH (1991) Chromosome rearrangement by ectopic recombination in Drosophila melanogaster: genome structure and evolution. Genetics 129(4):1085–1098. https://doi.org/10.1093/GENETICS/129.4.1085
Morrish TA, Garcia-Perez JL, Stamato TD, Taccioli GE, Sekiguchi JA, Moran JV (2007) Endonuclease-independent LINE-1 retrotransposition at mammalian telomeres. Nature 446(7132):208–212. https://doi.org/10.1038/NATURE05560
Munoz-Lopez M, Garcia-Perez J (2010) DNA transposons: nature and applications in genomics. Curr Genomics 11(2):115–128. https://doi.org/10.2174/138920210790886871
Naito K, Zhang F, Tsukiyama T, Saito H, Hancock CN, Richardson AO, Okumoto Y, Tanisaka T, Wessler SR (2009) Unexpected consequences of a sudden and massive transposon amplification on rice gene expression. Nature 461(7267):1130–1134. https://doi.org/10.1038/NATURE08479
Nguyen PQ, Huecas S, Asif-Laidin A, Plaza-Pegueroles A, Capuzzi B, Palmic N, Conesa C, Acker J, Reguera J, Lesage P, Fernández-Tornero C (2023) Structural basis of Ty1 integrase tethering to RNA polymerase III for targeted retrotransposon integration. Nat Commun 14(1):1729. https://doi.org/10.1038/s41467-023-37109-4
Nichuguti N, Fujiwara H (2020) Essential factors involved in the precise targeting and insertion of telomere-specific non-LTR retrotransposon, SART1Bm. Sci Rep 10(1):1–12. https://doi.org/10.1038/s41598-020-65925-x
Noel HR, Petrey JR, Palmer LD (2022) Mobile genetic elements in Acinetobacter antibiotic-resistance acquisition and dissemination. Ann N Y Acad Sci 1518(1):166–182. https://doi.org/10.1111/nyas.14918
Okazaki S, Tsuchida K, Maekawa H, Ishikawa H, Fujiwara H (1993) Identification of a pentanucleotide telomeric sequence, (TTAGG)n, in the silkworm Bombyx mori and in other insects. Mol Cell Biol 13(3):1424–1432. https://doi.org/10.1128/MCB.13.3.1424-1432.1993
Okazaki S, Ishikawa H, Fujiwara H (1995) Structural analysis of TRAS1, a novel family of telomeric repeat-associated retrotransposons in the silkworm. Bombyx Mori Mol Cell Biol 15(8):4545–4552. https://doi.org/10.1128/MCB.15.8.4545
Palazzo A, Caizzi R, Viggiano L, Marsano RM (2017) Does the promoter constitute a barrier in the horizontal transposon transfer process? Insight from Bari transposons. Genome Biol Evol 9(6):1637–1645. https://doi.org/10.1093/GBE/EVX122
Palazzo A, Lorusso P, Miskey C, Walisko O, Gerbino A, Marobbio CMT, Ivics Z, Marsano RM (2019) Transcriptionally promiscuous “blurry” promoters in Tc1/ mariner transposons allow transcription in distantly related genomes. Mobile DNA 10(1). https://doi.org/10.1186/S13100-019-0155-6
Pang Z, Raudonis R, Glick BR, Lin TJ, Cheng Z (2019) Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and alternative therapeutic strategies. Biotechnol Adv 37(1):177–192. https://doi.org/10.1016/J.BIOTECHADV.2018.11.013
Pardue ML, DeBaryshe PG (2011) Retrotransposons that maintain chromosome ends. Proc Natl Acad Sci USA 108(51):20317–20324. https://doi.org/10.1073/PNAS.1100278108
Park CW, Park J, Kren BT, Steer CJ (2006) Sleeping beauty transposition in the mouse genome is associated with changes in DNA methylation at the site of insertion. Genomics 88(2). https://doi.org/10.1016/j.ygeno.2006.04.007
Parks AR, Li Z, Shi Q, Owens RM, Jin MM, Peters JE (2009) Transposition into replicating DNA occurs through interaction with the processivity factor. Cell 138(4). https://doi.org/10.1016/j.cell.2009.06.011
Partridge SR, Kwong SM, Firth N, Jensen SO (2018) Mobile genetic elements associated with antimicrobial resistance. Clin Microbiol Rev 31(4):e00088-e117. https://doi.org/10.1128/CMR.00088-17
Pavlíček A, Jabbari K, Pačes J, Pačes V, Hejnar J, Bernardi G (2001) Similar integration but different stability of Alus and LINEs in the human genome. Gene 276(1–2):39–45. https://doi.org/10.1016/S0378-1119(01)00645-X
Payer LM, Burns KH (2019) Transposable elements in human genetic disease. Nat Rev Genet 20(12):760–772. https://doi.org/10.1038/s41576-019-0165-8
Pelisson A, Sun Song U, Prud’homme N, Smith PA, Bucheton A, Corces VG (1994) Gypsy transposition correlates with the production of a retroviral envelope-like protein under the tissue-specific control of the Drosophila flamenco gene. EMBO J 13(18):4401–4411. https://doi.org/10.1002/J.1460-2075.1994.TB06760.X
Perepelitsa-Belancio V, Deininger P (2003) RNA truncation by premature polyadenylation attenuates human mobile element activity. Nat Genet 35(4):363–366. https://doi.org/10.1038/NG1269
Peters JE, Craig NL (2000) Tn7 transposes proximal to DNA double-strand breaks and into regions where chromosomal DNA replication terminates. Mol Cell 6(3):573–582. https://doi.org/10.1016/S1097-2765(00)00056-3
Peters JE, Craig NL (2001) Tn7: smarter than we thought. Nat Rev Mol Cell Biol 2(11):806–814. https://doi.org/10.1038/35099006
Petrov DA, Aminetzach YT, Davis JC, Bensasson D, Hirsh AE (2003) Size matters: Non-LTR retrotransposable elements and ectopic recombination in Drosophila. Mol Biol Evol 20(6):880–892. https://doi.org/10.1093/MOLBEV/MSG102
Piégu B, Bire S, Arensburger P, Bigot Y (2015) A survey of transposable element classification systems - a call for a fundamental update to meet the challenge of their diversity and complexity. Mol Phylogenet Evol 86:90–109. https://doi.org/10.1016/j.ympev.2015.03.009
Poeschla EM (2008) Integrase, LEDGF/p75 and HIV replication. Cell Mol Life Sci 75(14):2491–2507. https://doi.org/10.1007/S00018-008-7540-5
Prud’homme N, Gans M, Masson M, Terzian C, Bucheton A (1995) Flamenco, a gene controlling the gypsy retrovirus of Drosophila melanogaster. Genetics 139(2):697–711. https://doi.org/10.1093/GENETICS/139.2.697
Puzakov MV, Puzakova LV, Shi S et al (2023) maT and mosquito transposons in cnidarians: evolutionary history and intraspecific differences. Funct Integr Genomics 23:244. https://doi.org/10.1007/s10142-023-01175-0
Rai SK, Sangesland M, Lee M, Esnault C, Cui Y, Chatterjee AG, Levin HL (2017) Host factors that promote retrotransposon integration are similar in distantly related eukaryotes. PLoS Genet 13(12). https://doi.org/10.1371/journal.pgen.1006775
Ren Y (2022) Regulatory mechanism and biological function of UHRF1–DNMT1-mediated DNA methylation. Funct Integr Genomics 22:1113–1126. https://doi.org/10.1007/s10142-022-00918-9
Reuter M, Berninger P, Chuma S, Shah H, Hosokawa M, Funaya C, Antony C, Sachidanandam R, Pillai RS (2011) Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature 480(7376):264–267. https://doi.org/10.1038/nature10672
Reznikoff WS (2008) Transposon Tn5. Annu Rev Genet 42:269–286. https://doi.org/10.1146/annurev.genet.42.110807.091656
Ros F, Kunze R (2001) Regulation of activator/dissociation transposition by replication and DNA methylation. Genetics 157(4):1723–1733. https://doi.org/10.1093/GENETICS/157.4.1723
Ross JA, Wardle SJ, Haniford DB (2010) Tn10/IS10 transposition is downregulated at the level of transposase expression by the RNA-binding protein Hfq. Mol Microbiol 78(3):607–621. https://doi.org/10.1111/j.1365-2958.2010.07359.x
Ross JA, Ellis MJ, Hossain S, Haniford DB (2013) Hfq restructures RNA-IN and RNA-OUT and facilitates antisense pairing in the Tn10/IS10 system. RNA 19(5):670–684. https://doi.org/10.1261/rna.037747.112
Rowe HM, Jakobsson J, Mesnard D, Rougemont J, Reynard S, Aktas T, Maillard PV, Layard-Liesching H, Verp S, Marquis J, Spitz F, Constam DB, Trono D (2010) KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463(7278):237–240. https://doi.org/10.1038/NATURE08674
Saint-Leandre B, Nguyen SC, Levine MT (2019) Diversification and collapse of a telomere elongation mechanism. Genome Res 29(6):920–931. https://doi.org/10.1101/GR.245001.118
Samelak-Czajka A, Wojciechowski P, Marszalek-Zenczak M et al (2023) Differences in the intraspecies copy number variation of Arabidopsis thaliana conserved and nonconserved miRNA genes. Funct Integr Genomics 23:120. https://doi.org/10.1007/s10142-023-01043-x
Sandmeyer S (2003) Integration by design. Proc Natl Acad Sci USA 100(10):5586–5588. https://doi.org/10.1073/PNAS.1031802100
Schaack S, Gilbert C, Feschotte C (2010) Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution. Trends Ecol Evol 25(9):537–546. https://doi.org/10.1016/j.tree.2010.06.001
Schultz DC, Ayyanathan K, Negorev D, Maul GG, Rauscher FJ (2002) SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev 16(8):919–932. https://doi.org/10.1101/GAD.973302
Sebastián C, Zwaans BMM, Silberman DM, Gymrek M, Goren A, Zhong L, Ram O, Truelove J, Guimaraes AR, Toiber D, Cosentino C, Greenson JK, MacDonald AI, McGlynn L, Maxwell F, Edwards J, Giacosa S, Guccione E, Weissleder R, … Mostoslavsky R (2012) The histone deacetylase SIRT6 is a tumor suppressor that controls cancer metabolism. Cell:151(6):1185–1199. https://doi.org/10.1016/J.CELL.2012.10.047
Seed KD, Lazinski DW, Calderwood SB, Camilli A (2013) A bacteriophage encodes its own CRISPR/Cas adaptive response to evade host innate immunity. Nature 494(7438):489–491. https://doi.org/10.1038/NATURE11927
Senti KA, Jurczak D, Sachidanandam R, Brennecke J (2015) piRNA-guided slicing of transposon transcripts enforces their transcriptional silencing via specifying the nuclear piRNA repertoire. Genes Dev 29(16):1747–1762. https://doi.org/10.1101/GAD.267252.115
Sheng Y, Wang H, Ou Y et al (2023) Insertion sequence transposition inactivates CRISPR-Cas immunity. Nat Commun 14:4366. https://doi.org/10.1038/s41467-023-39964-7
Shpiz S, Ryazansky S, Olovnikov I, Abramov Y, Kalmykova A (2014) Euchromatic transposon insertions trigger production of novel Pi- and endo-siRNAs at the target sites in the Drosophila germline. PLoS Genet 10(2):e1004138. https://doi.org/10.1371/JOURNAL.PGEN.1004138
Siguier P, Gourbeyre E, Varani A, Ton-Hoang B, Chandler M (2015) Everyman’s Guide to Bacterial Insertion Sequences. Microbiol Spectr 3(2). https://doi.org/10.1128/microbiolspec.mdna3-0030-2014
Sijen T, Plasterk RHA (2003) Transposon silencing in the Caenorhabditis elegans germ line by natural RNAi. Nature 426(6964):310–314. https://doi.org/10.1038/NATURE02107
Singh PK, Plumb MR, Ferris AL, Iben JR, Wu X, Fadel HJ, Luke BT, Esnault C, Poeschla EM, Hughes SH, Kvaratskhelia M, Levin HL (2015) LEDGF/p75 interacts with mRNA splicing factors and targets HIV-1 integration to highly spliced genes. Genes Dev 29(21):2287–2297. https://doi.org/10.1101/GAD.267609.115
Skipper KA, Andersen PR, Sharma N, Mikkelsen JG (2013) DNA transposon-based gene vehicles - scenes from an evolutionary drive. J Biomed Sci 20(1):92. https://doi.org/10.1186/1423-0127-20-92
Smaruj P, Kieliszek M (2023) Casposons - silent heroes of the CRISPR-Cas systems evolutionary history. EXCLI J 22:70–83. https://doi.org/10.17179/excli2022-5581
Sorek R, Ast G, Graur D (2002) Alu-containing exons are alternatively spliced. Genome Res 12(7):1060–1067. https://doi.org/10.1101/GR.229302
Sorek R, Lev-Maor G, Reznik M, Dagan T, Belinky F, Graur D, Ast G (2004) Minimal conditions for exonization of intronic sequences: 5’ splice site formation in alu exons. Mol Cell 14(2):221–231. https://doi.org/10.1016/S1097-2765(04)00181-9
Soucy SM, Huang J, Gogarten JP (2015) Horizontal gene transfer: building the web of life. Nat Rev Genet 16(8):472–482. https://doi.org/10.1038/nrg3962
Sridhar VV, Kapoor A, Zhang K, Zhu J, Zhou T, Hasegawa PM, Bressan RA, Zhu JK (2007) Control of DNA methylation and heterochromatic silencing by histone H2B deubiquitination. Nature 447(7145):735–738. https://doi.org/10.1038/NATURE05864
Sultana T, Zamborlini A, Cristofari G, Lesage P (2017) Integration site selection by retroviruses and transposable elements in eukaryotes. Nat Rev Genet 18(5):292–308. https://doi.org/10.1038/nrg.2017.7
Sultana T, van Essen D, Siol O, Bailly-Bechet M, Philippe C, Zine El Aabidine A, Pioger L, Nigumann P, Saccani S, Andrau JC, Gilbert N, Cristofari G (2019) The landscape of L1 retrotransposons in the human genome is shaped by pre-insertion sequence biases and post-insertion selection. Mol Cell. 74(3). https://doi.org/10.1016/j.molcel.2019.02.036
Sun ZW, Allis CD (2002) Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature 418(6893):104–108. https://doi.org/10.1038/NATURE00883
Takahashi H, Okazaki S, Fujiwara H (1997) A new family of site-specific retrotransposons, SART1, is inserted into telomeric repeats of the silkworm, Bombyx mori. Nucleic Acids Res 25(8):1. https://doi.org/10.1093/nar/25.8.1578
Taylor MS, LaCava J, Mita P, Molloy KR, Huang CRL, Li D, Adney EM, Jiang H, Burns KH, Chait BT, Rout MP, Boeke JD, Dai L (2013) Affinity proteomics reveals human host factors implicated in discrete stages of LINE-1 retrotransposition. Cell 155(5). https://doi.org/10.1016/j.cell.2013.10.021
Thomas J, Pritham EJ (2015) Helitrons, the eukaryotic rolling-circle transposable elements. Microbiol Spectr 3(4):10.1128. https://doi.org/10.1128/microbiolspec.mdna3-0049-2014
Tsukahara S, Kobayashi A, Kawabe A, Mathieu O, Miura A, Kakutani T (2009) Bursts of retrotransposition reproduced in Arabidopsis. Nature 461(7262):423–426. https://doi.org/10.1038/NATURE08351
Vaistij FE, Jones L, Baulcombe DC (2002) Spreading of RNA targeting and DNA methylation in rna silencing requires transcription of the target gene and a putative RNA-dependent RNA polymerase. Plant Cell 14(4):857–867. https://doi.org/10.1105/TPC.010480
Van Etten J, Bhattacharya D (2020) Horizontal gene transfer in eukaryotes: not if, but how much? Trends Genet 36(12):915–925. https://doi.org/10.1016/j.tig.2020.08.006
van Rij RP, Berezikov E (2009) Small RNAs and the control of transposons and viruses in Drosophila. Trends Microbiol 17(4):163–171. https://doi.org/10.1016/J.TIM.2009.01.003
van Houdt H, Bleys A, Depicker A (2003) RNA target sequences promote spreading of RNA silencing. Plant Physiol 131(1):245–253. https://doi.org/10.1104/PP.009407
van Meter M, Gorbunova V, Seluanov A (2014a) SIRT6: a promising target for cancer prevention and therapy. Adv Exp Med Biol 818:181–196. https://doi.org/10.1007/978-1-4471-6458-6_9
van Meter M, Kashyap M, Rezazadeh S, Geneva AJ, Morello TD, Seluanov A, Gorbunova V (2014b) SIRT6 represses LINE1 retrotransposons by ribosylating KAP1 but this repression fails with stress and age. Nat Commun 5(1):1–10. https://doi.org/10.1038/ncomms6011
Vigdal TJ, Kaufman CD, Izsvák Z, Voytas DF, Ivics Z (2002) Common physical properties of DNA affecting target site selection of Sleeping Beauty and other Tc1/mariner transposable elements. J Mol Biol 323(3):441–452. https://doi.org/10.1016/S0022-2836(02)00991-9
Vogan AA, Ament-Velásquez SL, Granger-Farbos A, Svedberg J, Bastiaans E, Debets AJ, Coustou V, Yvanne H, Clavé C, Saupe SJ, Johannesson H (2019) Combinations of Spok genes create multiple meiotic drivers in Podospora. ELife 8:e46454. https://doi.org/10.7554/eLife.46454
Vogan AA, Ament-Velásquez SL, Bastiaans E, Wallerman O, Saupe SJ, Suh A, Johannesson H (2021) The Enterprise, a massive transposon carrying Spok meiotic drive genes. Genome Res 31(5):789–798. https://doi.org/10.1101/GR.267609.120/-/DC1
Wang W, Han BW, Tipping C, Ge DT, Zhang Z, Weng Z, Zamore PD (2015) Slicing and binding by Ago3 or Aub trigger piwi-bound piRNA production by distinct mechanisms. Mol Cell 59(5):819–830. https://doi.org/10.1016/J.MOLCEL.2015.08.007
Warbrick E, Heatherington W, Lane DP, Glover DM (1998) PCNA binding proteins in Drosophila melanogaster: the analysis of a conserved PCNA binding domain. Nucleic Acids Res 26(17):1. https://doi.org/10.1093/nar/26.17.3925
Wheelan SJ, Aizawa Y, Han JS, Boeke JD (2005) Gene-breaking: a new paradigm for human retrotransposon-mediated gene evolution. Genome Res 15(8):1073–1078. https://doi.org/10.1101/GR.3688905
Wicker T, Sabot F, Hua-Van A, Bennetzen JL, Capy P, Chalhoub B, Flavell A, Leroy P, Morgante M, Panaud O, Paux E, SanMiguel P, Schulman AH (2007) A unified classification system for eukaryotic transposable elements. Nat Rev Genet 8(12):973–982. https://doi.org/10.1038/nrg2165
Widen SA, Bes IC, Koreshova A, Pliota P, Krogull D, Burga A (2023) Virus-like transposons cross the species barrier and drive the evolution of genetic incompatibilities. Science 380(6652). https://doi.org/10.1126/science.ade0705
Wolf D, Goff SP (2007) TRIM28 mediates primer binding site-targeted silencing of murine leukemia virus in embryonic cells. Cell 131(1):46–57. https://doi.org/10.1016/J.CELL.2007.07.026
Wolkow CA, DeBoy RT, Craig NL (1996) Conjugating plasmids are preferred targets for Tn7. Genes Dev 10(17):2145–2157. https://doi.org/10.1101/GAD.10.17.2145
Woo HR, Dittmer TA, Richards EJ (2008) Three SRA-domain methylcytosine-binding proteins cooperate to maintain global CpG methylation and epigenetic silencing in Arabidopsis. PLoS Genet 4(8):e1000156. https://doi.org/10.1371/JOURNAL.PGEN.1000156
Woodhouse MR, Freeling M, Lisch D (2006) The mop1 (mediator of paramutation1) mutant progressively reactivates one of the two genes encoded by the MuDR transposon in maize. Genetics 172(1):579–592. https://doi.org/10.1534/GENETICS.105.051383
Xie W, Gai X, Zhu Y, Zappulla DC, Sternglanz R, Voytas DF (2001) Targeting of the yeast Ty5 retrotransposon to silent chromatin is mediated by interactions between integrase and Sir4p. Mol Cell Biol 21(19):6606–6614. https://doi.org/10.1128/MCB.21.19.6606-6614.2001
Ying H, Hayward DC, Klimovich A, Bosch TCG, Baldassarre L, Neeman T, Forêt S, Huttley G, Reitzel AM, Fraune S, Ball EE, Miller DJ (2022) The role of DNA methylation in genome defense in Cnidaria and other invertebrates. Mol Biol Evol 39(2):msac018. https://doi.org/10.1093/molbev/msac018
Yuan YW, Wessler SR (2011) The catalytic domain of all eukaryotic cut-and-paste transposase superfamilies. Proc Natl Acad Sci USA 108(19):7884–7889. https://doi.org/10.1073/PNAS.1104208108
Yutin N, Shevchenko S, Kapitonov V, Krupovic M, Koonin EV (2015) A novel group of diverse Polinton-like viruses discovered by metagenome analysis. BMC Biol 13(1):95. https://doi.org/10.1186/s12915-015-0207-4
Zhang Y (2003) Transcriptional regulation by histone ubiquitination and deubiquitination. Genes Dev 17(22):2733–2740. https://doi.org/10.1101/GAD.1156403
Zhang X, Sun S, Xiang Y, Wong J, Klose T, Raoult D, Rossmann MG (2012) Structure of Sputnik, a virophage, at 3.5-Å resolution. Proc Natl Acad Sci U S A 109(45):18431–18436. https://doi.org/10.1073/pnas.1211702109
Zou S, Voytas DF (1997) Silent chromatin determines target preference of the Saccharomyces retrotransposon Ty5. Proc Natl Acad Sci USA 94(14):7412–7416. https://doi.org/10.1073/PNAS.94.14.7412
Zovoilis A, Cifuentes-Rojas C, Chu HP, Hernandez AJ, Lee JT (2016) Destabilization of B2 RNA by EZH2 activates the stress response. Cell 167(7):1788-1802.e13. https://doi.org/10.1016/J.CELL.2016.11.041
Author information
Authors and Affiliations
Contributions
P.C, R.S, and S.S conceived of the presented idea. P.C and R.S performed a literature survey to gather information. S.S analyzed the data. P.C and R.S wrote the main manuscript with inputs from all the authors. P.C. prepared the figures. S.S provided critical feedback and helped shape the analysis and the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chakrabarty, P., Sen, R. & Sengupta, S. From parasites to partners: exploring the intricacies of host-transposon dynamics and coevolution. Funct Integr Genomics 23, 278 (2023). https://doi.org/10.1007/s10142-023-01206-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-023-01206-w