Abstract
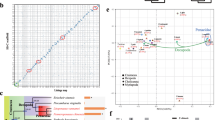
Tandem simple sequence repeats (SSRs) are one of the most popular molecular markers in genetic analysis owing to their ubiquitous occurrence,high reproducibility, multiallelic nature, and codominant mode. High mutability makes SSRs play a role in genome evolution and correspondingly show different patterns. Comparative analysis of genomic SSRs in different taxonomic groups usually focuses on land species, while marine animals have been neglected. This study examined the abundance of genomic SSRs with repeated unit lengths of 1–6 bp in 30 marine animals including nine taxonomic groups and further compared with the land species. More than thousands of SSRs were discovered in every organism which provided a huge resource for the development of molecular markers. Thirty marine animals showed profound differences in SSR characteristics, but some group-specific trends were also found. Both similarities and differences of repeat patterns were discovered between the land and marine species. Two taxon-specific SSR types were discovered: the pentanucleotides motif AGAGG in Euteleostei and the hexanucleotide repeats of ATGTAC in Porifera and Echinodermata. Gene ontology (GO) enrichment analysis of two representative species (Amphimedon queenslandica for Porifera and Strongylocentrotus purpuratus for Echinodermata) revealed functional preference of the ATGTAC motif associated genes, and this might hint at evolutionary significance.



Similar content being viewed by others
References
Amos W (1999) Microsatellites: evolution and applications. Oxford University Press, Oxford
Arora V, Iquebal MA, Rai A, Kumar D (2013) In silico mining of putative microsatellite markers from whole genome sequence of water buffalo (Bubalus bubalis) and development of first BuffSatDB. BMC Genomics 14:43
Bachtrog D, Weiss S, Zangerl B, Brem G, Schlötterer C (1999) Distribution of dinucleotide microsatellites in the Drosophila melanogaster genome. Mol Biol Evol 16:602–610
Brandt B, Gebhardt F, Bürger H (2000) Modulation of EGFR gene transcription by secondary structures, a polymorphic repetitive sequence and mutations-a link between genetics and epigenetics. Histol Histopathol 15:929–936
Cohen JB, Effron K, Rechavi G, Ben-Neriah Y, Zakut R, Givol D (1982) Simple DNA sequences in homologous flanking regions near immunoglobulin VH genes: a role in gene interaction? Nucleic Acids Res 10:3353–3370
Dybvig K, Clark CD, Aliperti G, Schlesinger MJ (1983) A chicken repetitive DNA sequence that is highly sensitive to single-strand specific endonucleases. Nucleic Acids Res 11:8495–8508
Field D, Wills C (1998) Abundant microsatellite polymorphism in Saccharomyces cerevisiae, and the different distributions of microsatellites in eight prokaryotes and S. cerevisiae, result from strong mutation pressures and a variety of selective forces. PNAS 95:1647–1652
Gebhardt F, Zänker KS, Brandt B (1999) Modulation of epidermal growth factor receptor gene transcription by a polymorphic dinucleotide repeat in intron 1. J Biol Chem 274:13176–13180
Hancock JM (2002) Genome size and the accumulation of simple sequence repeats: implications of new data from genome sequencing projects. Genetica 115:93–103
Hollenbeck CM, Portnoy DS, Gold JR (2012) Use of comparative genomics to develop EST-SSRs for red drum (Sciaenops ocellatus). Mar Biotechnol 14:672–680
Jurka J, Pethiyagoda C (1995) Simple repetitive DNA sequences from primates: compilation and analysis. J Mol Evol 40:120–126
Karaoglu H, Lee CMY, Meyer W (2005) Survey of simple sequence repeats in completed fungal genomes. Mol Biol Evol 22:639–649
Karlin S, Mrazek J, Campbell AM (1997) Compositional biases of bacterial genomes and evolutionary implications. J Bacteriol 179:3899–3913
Katti MV, Ranjekar PK, Gupta VS (2001) Differential distribution of simple sequence repeats in eukaryotic genome sequences. Mol Biol Evol 18:1161–1167
Kofler R, Schlötterer C, Lelley T (2007) SciRoKo: a new tool for whole genome microsatellite search and investigation. Bioinformatics 23:1683–1685
La Rota M, Kantety RV, Yu J-K, Sorrells ME (2005) Nonrandom distribution and frequencies of genomic and EST-derived microsatellite markers in rice, wheat, and barley. BMC Genomics 6:23
Lawson MJ, Zhang L (2006) Distinct patterns of SSR distribution in the Arabidopsis thaliana and rice genomes. Genome Biol 7:R14
Li YC, Korol AB, Fahima T, Beiles A, Nevo E (2002) Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review. Mol Ecol 11:2453–2465
Li Y-C, Korol AB, Fahima T, Nevo E (2004) Microsatellites within genes: structure, function, and evolution. Mol Biol Evol 21:991–1007
Liu S, Rexroad CE III, Couch CR, Cordes JF, Reece KS, Sullivan CV (2012) A microsatellite linkage map of striped bass (Morone saxatilis) reveals conserved synteny with the three-spined stickleback (Gasterosteus aculeatus). Mar Biotechnol 14:237–244
Mayer C, Leese F, Tollrian R (2010) Genome-wide analysis of tandem repeats in Daphnia pulex-a comparative approach. BMC Genomics 11:277
Meloni R, Albanèse V, Ravassard P, Treilhou F, Mallet J (1998) A tetranucleotide polymorphic microsatellite, located in the first intron of the tyrosine hydroxylase gene, acts as a transcription regulatory element in vitro. Hum Mol Genet 7:423–428
Mock T, Kirkham A (2012) What can we learn from genomics approaches in marine ecology? From sequences to eco-systems biology! Mar Ecol 33:131–148
Morgante M, Hanafey M, Powell W (2002) Microsatellites are preferentially associated with nonrepetitive DNA in plant genomes. Nat Genet 30:194–200
Moxon ER, Rainey PB, Nowak MA, Lenski RE (1994) Adaptive evolution of highly mutable loci in pathogenic bacteria. Curr Biol 4:24–33
MüllER WE, Wiens M, Adell T, Gamulin V, Schröder HC, Müller IM (2004) Bauplan of urmetazoa: basis for genetic complexity of metazoa. Int Rev Cytol 235:53–92
Pérez F, Ortiz J, Zhinaula M, Gonzabay C, Calderón J, Volckaert FA (2005) Development of EST-SSR markers by data mining in three species of shrimp: Litopenaeus vannamei, Litopenaeus stylirostris, and Trachypenaeus birdy. Mar Biotechnol 7:554–569
Rubinsztein DC, Amos W, Leggo J, Goodburn S, Jain S, Li S-H, Margolis RL, Ross CA, Ferguson-Smith MA (1995) Microsatellite evolution-evidence for directionality and variation in rate between species. Nat Genet 10:337–343
Schlötterer C (2000) Evolutionary dynamics of microsatellite DNA. Chromosoma 109:365–371
Schug MD, Mackay TF, Aquadro CF (1997) Low mutation rates of microsatellite loci in Drosophila melanogaster. Nat Genet 15:99–102
Srivastava M, Simakov O, Chapman J, Fahey B, Gauthier ME, Mitros T, Richards GS, Conaco C, Dacre M, Hellsten U (2010) The Amphimedon queenslandica genome and the evolution of animal complexity. Nature 466:720–726
Subramanian S, Mishra RK, Singh L (2003) Genome-wide analysis of microsatellite repeats in humans: their abundance and density in specific genomic regions. Genome Biol 4:R13
Suzuki M, Iwashima A, Kimura M, Kogure T, Nagasawa H (2013) The molecular evolution of the Pif family proteins in various species of mollusks. Mar Biotechnol 15:145–158
Tautz D, Trick M, Dover GA (1986) Cryptic simplicity in DNA is a major source of genetic variation. Nature 322:652–656
Tóth G, Gáspári Z, Jurka J (2000) Microsatellites in different eukaryotic genomes: survey and analysis. Genome Res 10:967–981
van Belkum A, Scherer S, van Alphen L, Verbrugh H (1998) Short-sequence DNA repeats in prokaryotic genomes. Microbiol Mol Biol Rev 62:275–293
von Stackelberg M, Rensing SA, Reski R (2006) Identification of genic moss SSR markers and a comparative analysis of twenty-four algal and plant gene indices reveal species-specific rather than group-specific characteristics of microsatellites. BMC Plant Biol 6:9
Wang S, Zhang L, Meyer E, Bao Z (2010) Genome-wide analysis of transposable elements and tandem repeats in the compact placozoan genome. Biol Direct 5:18–26
Weber JL, Wong C (1993) Mutation of human short tandem repeats. Hum Mol Genet 2:1123–1128
Webster MT, Smith NG, Ellegren H (2002) Microsatellite evolution inferred from human–chimpanzee genomic sequence alignments. PNAS 99:8748–8753
Werner D, Neuer-Nitsche B (1989) Site-specific location of covalent DNA-polypeptide complexes in the chicken genome. Nucleic Acids Res 17:6005–6015
Xu X, Peng M, Fang Z, Xu X (2000) The direction of microsatellite mutations is dependent upon allele length. Nat Genet 24:396–399
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:W293–W297
Young ET, Sloan JS, Van Riper K (2000) Trinucleotide repeats are clustered in regulatory genes in Saccharomyces cerevisiae. Genetics 154:1053–1068
Acknowledgments
This study was supported by the grants from 973 Program (2010CB126406), National High Technology Research and Development Program (2012AA10A405-6), and National Natural Science Foundation of China (31372524).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
Distribution of SSRs in the genomic sequence of thirty marine animals (XLSX 465122 kb)
Rights and permissions
About this article
Cite this article
Jiang, Q., Li, Q., Yu, H. et al. Genome-Wide Analysis of Simple Sequence Repeats in Marine Animals—a Comparative Approach. Mar Biotechnol 16, 604–619 (2014). https://doi.org/10.1007/s10126-014-9580-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-014-9580-1