Abstract
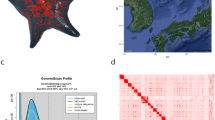
Penelope, originally found as a key element responsible for the hybrid dysgenesis in Drosophila virilis, has been widely conserved throughout eukaryotic genomes. In other organisms, they are often referred to as Penelope-like elements or PLEs. In this study, we found two types of PLEs, designated MjPLE01 and MjPLE02, from kuruma shrimp, Marsupenaeus japonicus. There was no observed nucleotide similarity between MjPLE01 and 02, and both elements differed from each other in terms of their structure; MjPLE02 has a distinctive endonuclease (EN) domain at the C-terminus while MjPLE01 do not. A phylogenetic tree that includes publicly available PLEs and TERTs showed that MjPLE01 and 02 were closely related to Coprina elements, which have been reported as an EN-deficient PLE, and to Penelope–Poseidon group, which possess an EN domain, respectively. Genomic Southern blot analysis using MjPLE01 as a probe showed several multiple bands that differ among individual shrimps. On the other hand, two major identical bands were observed when MjPLE02 was used. Colony hybridization showed co-localization of MjPLE01 and GGTTA repeats, suggesting that MjPLE01 might be prevalent in subtelomeric regions of kuruma shrimp genome. These results suggest that the kuruma shrimp genome has at least two types of PLEs with different domain compositions, phylogenetic positions, and probably chromosomeal localization. Such distinctive types of PLEs in an organism have never been described and hence could be a potential source to understand how multiple PLE types evolved.




Similar content being viewed by others
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21(9):2104–2105
Anderson JA, Song YS, Langley CH (2008) Molecular population genetics of Drosophila subtelomeric DNA. Genetics 178:477–487
Arkhipova IR (2006) Distribution and phylogeny of Penelope-like elements in eukaryotes. Syst Biol 55:875–885
Arkhipova IR, Pyatkov KI, Meselson M, Evgen'ev MB (2003) Retroelements containing introns in diverse invertebrate taxa. Nat Genet 33:123–124
Caipang CM, Haraguchi I, Ohira T, Hirono I, Aoki T (2004) Rapid detection of a fish iridovirus using loop-mediated isothermal amplification (LAMP). J Virol Methods 121:155–161
Dalle Nogare DE, Clark MS, Elgar G, Frame IG, Poulter RT (2002) Xena, a full-length basal retroelement from tetraodontid fish. Mol Biol Evol 19:247–255
Eickbush TH, Jamburuthugoda VK (2008) The diversity of retrotransposons and the properties of their reverse transcriptases. Virus Res 134:221–234
Evgen'ev MB, Arkhipova IR (2005) Penelope-like elements—a new class of retroelements: distribution, function and possible evolutionary significance. Cytogenet Genome Res 110:510–521
Evgen'ev MB, Zelentsova H, Shostak N, Kozitsina M, Barskyi V, Lankenau DH, Corces VG (1997) Penelope, a new family of transposable elements and its possible role in hybrid dysgenesis in Drosophila virilis. Proc Natl Acad Sci USA 94:196–201
Evgen'ev M, Zelentsova H, Mnjoian L, Poluectova H, Kidwell MG (2000a) Invasion of Drosophila virilis by the Penelope transposable element. Chromosoma 109:350–357
Evgen'ev MB, Zelentsova H, Poluectova H, Lyozin GT, Veleikodvorskaja V, Pyatkov KI, Zhivotovsky LA, Kidwell MG (2000b) Mobile elements and chromosomal evolution in the virilis group of Drosophila. Proc Natl Acad Sci USA 97:11337–11342
Gladyshev EA, Arkhipova IR (2007) Telomere-associated endonuclease-deficient Penelope-like retroelements in diverse eukaryotes. Proc Natl Acad Sci USA 104:9352–9357
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Kazazian HH Jr (2004) Mobile elements: drivers of genome evolution. Science 303:1626–1632
Kazazian HH Jr, Scott AF (1993) “Copy and paste” transposable elements in the human genome. J Clin Invest 91:1859–1860
Koyama T, Asakawa S, Katagiri T, Shimizu A, Fagutao FF, Mavichak R, Santos MD, Fuji K, Sakamoto T, Kitakado T, Kondo H, Shimizu N, Aoki T, Hirono I (2010) Hyper-expansion of large DNA segments in the genome of kuruma shrimp, Marsupenaeus japonicus. BMC Genomics 11:141
Linardopoulou EV, Williams EM, Fan Y, Friedman C, Young JM, Trask BJ (2005) Human subtelomeres are hot spots of interchromosomal recombination and segmental duplication. Nature 437:94–100
Lyozin GT, Makarova KS, Velikodvorskaja VV, Zelentsova HS, Khechumian RR, Kidwell MG, Koonin EV, Evgen'ev MB (2001) The structure and evolution of Penelope in the virilis species group of Drosophila: an ancient lineage of retroelements. J Mol Evol 52:445–456
SanMiguel P, Tikhonov A, Jin YK, Motchoulskaia N, Zakharov D, Melake-Berhan A, Springer PS, Edwards KJ, Lee M, Avramova Z, Bennetzen JL (1996) Nested retrotransposons in the intergenic regions of the maize genome. Science 274:765–768
Volff JN, Hornung U, Schartl M (2001) Fish retroposons related to the Penelope element of Drosophila virilis define a new group of retrotransposable elements. Mol Genet Genomics 265:711–720
Xiong Y, Eickbush TH (1990) Origin and evolution of retroelements based upon their reverse transcriptase sequences. EMBO J 9:3353–3362
Zelentsova H, Poluectova H, Mnjoian L, Lyozin G, Veleikodvorskaja V, Zhivotovsky L, Kidwell MG, Evgen'ev MB (1999) Distribution and evolution of mobile elements in the virilis species group of Drosophila. Chromosoma 108:443–456
Acknowledgment
This work was supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Koyama, T., Kondo, H., Aoki, T. et al. Identification of Two Penelope-Like Elements with Different Structures and Chromosome Localization in Kuruma Shrimp Genome. Mar Biotechnol 15, 115–123 (2013). https://doi.org/10.1007/s10126-012-9474-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-012-9474-z