Abstract
Introduction
Sulfur-oxidizing bacteria (SOB) play a key role in the biogeochemical cycling of sulfur.
Objectives
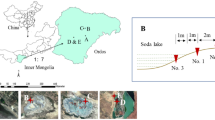
To explore SOB diversity, distribution, and physicochemical drivers in five volcanic lakes and two springs in the Wudalianchi volcanic field, China.
Methods
This study analyzed microbial communities in samples via high-throughput sequencing of the soxB gene. Physical-chemical parameters were measured, and QIIME 2 (v2019.4), R, Vsearch, MEGA7, and Mothur processed the data. Alpha diversity indices and UPGMA clustering assessed community differences, while heat maps visualized intra-sample variations. Canoco 5.0 analyzed community-environment correlations, and NMDS, Adonis, and PcoA explored sample dissimilarities and environmental factor correlations. SPSS v.18.0 tested for statistical significance.
Results
The diversity of SOB in surface water was higher than in springs (more than 7.27 times). We detected SOB affiliated to β-proteobacteria (72.3 %), α-proteobacteria (22.8 %), and γ-proteobacteria (4.2 %) distributed widely in these lakes and springs. Rhodoferax and Cupriavidus were most frequent in all water samples, while Rhodoferax and Bradyrhizobium are dominant in surface waters but rare in springs. SOB genera in both habitats were positively correlated. Co-occurrence analysis identified Bradyrhizobium, Blastochloris, Methylibium, and Metyhlobacterium as potential keystone taxa. Redundancy analysis (RDA) revealed positive correlations between SOB diversity and total carbon (TC), Fe2+, and total nitrogen (TN) in all water samples.
Conclusion
The diversity and community structure of SOB in volcanic lakes and springs in the Wudalianchi volcanic group were clarified. Moreover, the diversity and abundance of SOB decreased with the variation of water openness, from open lakes to semi-enclosed lakes and enclosed lakes.







Similar content being viewed by others
Data availability
The sequence data supporting our study findings have been deposited in the China National Microbiology Data Center (NMDC) with accession numbers NMDC40054990 to NMDC40054996 (soxB). Additionally, the paired-end forward and reverse sequences, labeled as SUB1713833866551, have been submitted for inclusion in the NMDC repository ( https://nmdc.cn/submit/metagenome/overview/SUB1713833866551).
References
Agarwalab L, Prakasha A, Purohitb HJ (2019) Expression of autotrophic genes under CO2 environment and genome mining of desert bacterium Cupriavidus sp. HPC(L). Bioresource Technol Rep 7:100258. https://doi.org/10.1016/j.biteb.2019.100258
Akan JC, Abbagambo MT, Chellube ZM et al (2012) Assessment of pollutants in water and sediment samples in Lake Chad, Baga North Eastern Nigeria. J Environ Prot 3(11):1428–1441. https://doi.org/10.4236/jep.2012.311161
Alessa O, Ogura Y, Fujitani Y et al (2021) Comprehensive comparative genomics and phenotyping of Methylobacterium species. Front Microbiol 12:740610. https://doi.org/10.3389/fmicb.2021.740610
Altschul SF, Madden TL, Schäffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402. https://doi.org/10.1093/nar/25.17.3389
Behera P, Mohapatra M, Kim JY et al (2019) Spatial and temporal heterogeneity in the structure and function of sediment bacterial communities of a tropical mangrove forest. Environ Sci Pollut Res Int 26(4):3893–3908. https://doi.org/10.1007/s11356-018-3927-5
Boada E, Santos-Clotas E, Bertran S et al (2020) Potential use of Methylibium sp. as a biodegradation tool in organosilicon and volatile compounds removal for biogas upgrading. Chemosphere 240:124908. https://doi.org/10.1016/j.chemosphere.2019.124908
Boës X, Prat S, Arrighi V et al (2019) Lake-Level changes and hominin occupations in the arid Turkana basin during volcanic closure of the Omo River outflows to the Indian Ocean. Quat Res: An Interdiscip J 91(2):892–909. https://doi.org/10.1017/qua.2018.118
Bolyen E, Rideout JR, Dillon MR et al (2018) QIIME 2: Reproducible, interactive, scalable, and extensible microbiome data science. Peer J Preprints. https://doi.org/10.7287/peerj.preprints.27295v2
Bouhnik O, Chaddad Z, Alami S et al (2024) Symbiotic efficiency of Bradyrhizobium symbiovars on Chamaecytisus albidus plants grown under water stress and acidic pH. Symbiosis 1–11. https://doi.org/10.1007/s13199-024-00989-1
Braak CT, Smilauer P (2012) Canoco reference manual and user’s guide: software for ordination, version 5.0. Microcomputer Power, Ithaca, NY, USA
Bukin SV, Pavlova ON, Kalmychkov GV et al (2018) Substrate specificity of methanogenic communities from Lake Baikal bottom sediments associated with hydrocarbon gas discharge. Microbiology 87(4):549–558. https://doi.org/10.1134/S0026261718040045
Chang W, Sun J, Pang Y et al (2020) Effects of different habitats on the bacterial community composition in the water and sediments of Lake Taihu. China Environ Sci Pollut Res 27(36):449483–444944. https://doi.org/10.1007/s11356-020-10376-0
Chaudhary S, Dhanker R, Singh K et al (2022) Characterization of sulfur-oxidizing bacteria isolated from mustard (Brassica juncea L.) rhizosphere having the capability of improving sulfur and nitrogen uptake. J Appl Microbiol 133(5):2814–2825. https://doi.org/10.1111/jam.15742
Chen M, Jiao Y, Zhang Y et al (2020) Succession of sulfur bacteria during decomposition of cyanobacterial bloom biomass in the shallow Lake Nanhu: an ex situ mesocosm study. Chemosphere 256:127101. https://doi.org/10.1016/j.chemosphere.2020.127101
Chen JS, Tsai HC, Hsu YL et al (2022) Comprehensive assessment of bacterial communities and their functional profiles in the Huang Gang Creek in the Tatun Volcano Group basin, Taiwan using 16S rRNA amplicon sequencing. Ecotoxicol Environ 234:113375. https://doi.org/10.1016/j.ecoenv.2022.113375
Cleary DFR, Ferreira MRS, Bat NK et al (2019) Bacterial composition of sponges, sediment and seawater in enclosed and open marine lakes in Ha Long Bay Vietnam. Mar Biol Res 16(12):1–14. https://doi.org/10.1080/17451000.2019.1702214
Deja-Sikora E, Gołębiewski M, Kalwasińska A et al (2019) Comamonadaceae OTU as a remnant of an ancient microbial community in sulfidic waters. Microb Ecol 78(1):85–101. https://doi.org/10.1007/s00248-018-1270-5
Findlay AJ, AndréPellerin KatjaLaufer et al (2020) Quantification of sulphide oxidation rates in marine sediment. Geochim Cosmochim Acta 280:441–452. https://doi.org/10.1016/j.gca.2020.04.007
Gałązka A, Grządziel J, Gałązka R et al (2018) Genetic and functional diversity of bacterial microbiome in soils with long term impacts of petroleum hydrocarbons. Front Microbiol 9:1923. https://doi.org/10.3389/fmicb.2018.01923
Gao S, Wen Y, Zhang W et al (2019) Microbial characteristics and eco-health implication of mineral spring water in Wudalianchi, Northeast China. Chin J Appl Ecol 30(8):2865–2874. https://doi.org/10.13287/j.1001-9332.201908.036
Ghosh W, Dam B (2009) Biochemistry and molecular biology of lithotrophic sulfur oxidation by taxonomically and ecologically diverse bacteria and archaea. FEMS Microbiol Rev 33(6):999–1043. https://doi.org/10.1111/j.1574-6976.2009.00187.x
González-Paz JR, Becerril-Varela K, Guerrero-Barajas C (2022) Iron reducing sludge as a source of electroactive bacteria: assessing iron reduction in biofilm bacteria, planktonic cells and isolates from a microbial fuel cell. Arch Microbiol 204(10):632. https://doi.org/10.1007/s00203-022-03253-6
Grattieri M (2020) Purple bacteria photo-bioelectrochemistry: enthralling challenges and opportunities. Photochem Photobiol Sci 19(4):424–435. https://doi.org/10.1039/c9pp00470j
Guo Q, Zhou Z, Zhang L et al (2020) Skermanella pratensis sp. nov., isolated from meadow soil, and emended description of the genus Skermanella. Int J Syst Evol Microbiol 70(3):1605–1609. https://doi.org/10.1099/ijsem.0.003944
Gupta D, Guzman MS, Rengasamy K et al (2021) Photoferrotrophy and phototrophic extracellular electron uptake is common in the marine anoxygenic phototroph Rhodovulum sulfidophilum. ISME J 15(11):3384–3398. https://doi.org/10.1038/s41396-021-01015-8
Hassan SHA, Ginkel SWV, Oh SE (2012) Detection of Cr6+ by the sulfur oxidizing bacteria biosensor: effect of different physical factors. Environ Sci Technol 46(14):7844–7848. https://doi.org/10.1021/es301360a
Hassan SHA, Ginkel SWV, Oh SE (2013) Effect of organics and alkalinity on the sulfur oxidizing bacteria (SOB) biosensor. Chemosphere 90(3):965–970. https://doi.org/10.1016/j.chemosphere.2012.06.040
He R, Wang J, Pohlman JW et al (2022) Metabolic flexibility of aerobic methanotrophs under anoxic conditions in Arctic lake sediments. ISME J 16(1):78–90
Heydari S, Pirzad A (2021) Improvement of the yield-related response of mycorrhized Lallemantia iberica to salinity through sulfur-oxidizing bacteria. J Sci Food Agric 101(9):3758–3766. https://doi.org/10.1002/jsfa.11007
Hiraishi A, Nagao N, Yonekawa C et al (2020) Distribution of phototrophic purple nonsulfur bacteria in massive blooms in coastal and wastewater ditch environments. Microorganisms 8(2):150. https://doi.org/10.3390/microorganisms8020150
Hossain MJ, AftabUddin S, Akhterc F et al (2022) Surface water, sediment, and biota: the first multi-compartment analysis of microplastics in the Karnafully river. Bangladesh Mar Pollut Bull 180:113820. https://doi.org/10.1016/j.marpolbul.2022.113820
Hu Q, Wang S, Lai Q et al (2021) Sulfurimonas indica sp. nov., a hydrogen- and sulfur-oxidizing chemolithoautotroph isolated from a hydrothermal sulfide chimney in the Northwest Indian Ocean. Int J Syst Evol Microbiol 71:1. https://doi.org/10.1099/ijsem.0.004575
Irfan T, Khalid S, Taneez M et al (2020) Plastic driven pollution in Pakistan: the first evidence of environmental exposure to microplastic in sediments and water of Rawal Lake. Environ Sci Pollut Res Int 27(13):15083–15092. https://doi.org/10.1007/s11356-020-07833-1
Jaffer YD, Purushothaman CS, Kumar HS et al (2019) A combined approach of 16S rRNA and a functional marker gene, soxB to reveal the diversity of sulphur-oxidising bacteria in thermal springs. Arch Microbiol 201(7):951–967. https://doi.org/10.1007/s00203-019-01666-4
Johnson DB, Smith SL, Santos AL (2021) Bioleaching of transition metals from limonitic laterite deposits and reassessment of the multiple roles of sulfur-oxidizing Acidophiles in the Process. Front Microbiol 12:703177. https://doi.org/10.3389/fmicb.2021.703177
Jung J, Yoo K-C, Rosenheim BE et al (2019) Microbial Fe(III) reduction as a potential iron source from Holocene sediments beneath Larsen Ice Shelf. Nat Commun 10(1):5786. https://doi.org/10.1038/s41467-019-13741-x
Klotz M, Bryant D, Hanson T (2011) The Microbial Sulfur Cycle. Front Microbiol 2:241. https://doi.org/10.3389/fmicb.2011.00241
Kojima H, Fukui M (2010) Sulfuricella denitrificans gen. nov., sp. nov., a sulfur-oxidizing autotroph isolated from a freshwater lake. Int J Syst Evol Microbiol 60(Pt 12):2862–2866. https://doi.org/10.1099/ijs.0.016980-0
Kojima H, Watanabe T, Iwata T et al (2014) Identification of major planktonic sulfur oxidizers in stratified freshwater lake. PLoS ONE 9(4):e93877. https://doi.org/10.1371/journal.pone.0093877
Kojima H, Watanabe M, Miyata N et al (2022) Sulfuricystis multivorans gen. nov., sp. nov. and Sulfuricystis thermophila sp. nov., facultatively autotropic sulfur-oxidizing bacteria isolated from a hot spring, and emended description of the genus Rugosibacter. Arch Microbiol 204(9):595. https://doi.org/10.1007/s00203-022-03186-0
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Kumar U, Panneerselvam P, Gupta VVSR et al (2018) Diversity of sulfur-oxidizing and sulfur-reducing microbes in diverse ecosystems. Adv Soil Microbiol Recent Trends Future Prospects (3):65–89. https://doi.org/10.1007/978-981-10-6178-3_4
Li B, Wang X, Liu G et al (2022) Microbial diversity response to geochemical gradient characteristics on AMD from abandoned Dashu pyrite mine in Southwest China. Environ Sci Pollut Res Int 29(49):74983–74997. https://doi.org/10.1007/s11356-022-21031-1
Liang Y, Pan Z, Feng H et al (2022) Biofilm coupled micro-electrolysis of waste iron shavings enhanced iron and hydrogen autotrophic denitrification and phosphate accumulation for wastewater treatment. J Environ Chem Eng 10(6):108959. https://doi.org/10.1016/j.jece.2022.108959
Liu H, Dai L, Yao J et al (2021) Efficient biotransformation of sulfide in anaerobic sequencing batch reactor by composite microbial agent: performance optimization and microbial community analysis. Environ Sci Pollut Res Int 28(35):48718–48727. https://doi.org/10.1007/s11356-021-12717-z
Liu T, Da H, Zhang S et al (2022) Magnetotactic bacteria in vertical sediments of volcanic lakes in NE China appear Alphaproteobacteria dominated distribution regardless of waterbody types. World J Microbiol Biotechnol 38(5):76. https://doi.org/10.1007/s11274-022-03262-z
Loy A, Duller S, Baranyi C et al (2009) Reverse dissimilatory sulfite reductase as phylogenetic marker for a subgroup of sulfur-oxidizing prokaryotes. Environ Microbiol 11(2):289–299. https://doi.org/10.1111/j.1462-2920.2008.01760.x
Ma T, Jiang Y, Elbehery AHA et al (2020) Resilience of planktonic bacterial community structure in response to short-term weather deterioration during the growing season in an alpine lake. Hydrobiologia 847:535–548. https://doi.org/10.1007/s10750-019-04118-8
Maeda I (2021) Potential of phototrophic purple nonsulfurbacteria to fix nitrogen in rice fields. Microorganisms 10(1):28. https://doi.org/10.3390/microorganisms10010028
Mao X, Wang Y, Chudaev OV et al (2009) Geochemical evidence of gas sources of CO2-rich cold springs from Wudalianchi, Northeast China. J Earth Sci 20(006):020. https://doi.org/10.1007/s12583-009-0081-5
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J 17(1):10–12. https://doi.org/10.14806/EJ.17.1.200
Meyer B, Imhoff JF, Kuever J (2007) Molecular analysis of the distribution and phylogeny of the soxB gene among sulfur-oxidizing bacteria - evolution of the Sox sulfur oxidation enzyme system. Environ Microbiol 9(12):2957–2977. https://doi.org/10.1111/j.1462-2920.2007.01407.x
Nagar S, Talwar C, Motelica-Heino M et al (2022) Microbial ecology of sulfur biogeochemical cycling at a mesothermic hot spring atop Northern Himalayas. India Cold Spring Harbor Lab 13:848010. https://doi.org/10.3389/fmicb.2022.848010
Ni H, Lu F, Luo X et al (2008) Assessment of sampling designs to measure riverine fluxes from the Pearl River Delta, China to the South China Sea. Environ Monit Assess 143:291–301. https://doi.org/10.1007/s10661-007-9982-x
Ni Z, Wang S, Zhang B-T et al (2019) Response of sediment organic phosphorus composition to lake trophic status in China. Sci Total Environ 652:495–504. https://doi.org/10.1016/j.scitotenv.2018.10.233
Nilsson RH, Ryberg M, Kristiansson E et al (2006) Taxonomic reliability of DNA sequences in public sequence databases: a fungal perspective. PLoS ONE 1(1):e59. https://doi.org/10.1371/journal.pone.0000059
Nosalova L, Piknova M, Bonova K et al (2022) Deep subsurface hypersaline environment as a source of novel species of halophilic sulfur-oxidizing bacteria. Microorganisms 10(5):995. https://doi.org/10.3390/microorganisms10050995
Okwir G, Kumar SP, Gao H et al (2022) Multi-variate regression analysis of lake level variability: a case of semi-closed, shallow rift valley lake in Northern Tanzania. Environ Challenges 7:100533. https://doi.org/10.1016/j.envc.2022.100533
Overholt WA, Trumbore S, Xu X et al (2022) Carbon fixation rates in groundwater similar to those in oligotrophic marine systems. Nat Geosci 15(7):561–567. https://doi.org/10.1038/s41561-022-00968-5
Pang Y, Wang J (2021) Inhibition of ferrous iron (Fe2+) to sulfur-driven autotrophic denitrification: insight into microbial community and functional genes. Biores Technol 342:125960. https://doi.org/10.1016/j.biortech.2021.125960
Pang Y, Wang J, Li S et al (2021) Long-term sulfide input enhances chemoautotrophic denitrification rather than DNRA in freshwater lake sediments. Environ Pollut 270:116201. https://doi.org/10.1016/j.envpol.2020.116201
Peng R, Shen J, Li S et al (2023) Sediment-isolated Comamonas terrigena strain HJ-2: a novel nitrate-dependent ferrous-oxidizing bacterium with multifunction on pollutant transformation. Lett Appl Microbiol 76(1):ovac022. https://doi.org/10.1093/lambio/ovac022
Pham VH, Yong JJ, Park SJ et al (2008) Molecular analysis of the diversity of the sulfide: quinone reductase (sqr) gene in sediment environments. Microbiology 154(Pt 10):3112–3121. https://doi.org/10.1099/mic.0.2008/018580-0
Qiu X, Peng W, Sun L et al (2022) Use of hydrogen and oxygen isotopes to understand evaporation from enclosed waterbodies. J Environ Eng Landsc Manag 30(1):220–225. https://doi.org/10.3846/jeelm.2022.16299
Quijano G, Valenzuela EI, Cantero D et al (2021) Impact of an anoxic desulfurization process on methane content of the purified biogas. Fuel 303:121256. https://doi.org/10.1016/j.fuel.2021.121256
Rasskazov S, Sun Y, Chuvashova I et al (2020) Trace-element and Pb isotope evidence on extracting sulfides from potassic melts beneath Longmenshan and Molabushan volcanoes, Wudalianchi. Northeast China Minerals 10(4):319. https://doi.org/10.3390/min10040319
Ratananikom K, Peungtim P, Phuinthiang P et al (2022) Development of bio-electrochemical reactor for groundwater denitrification: effect of electric current and water hardness. Sustainability 14(15):9454. https://doi.org/10.3390/su14159454
Reza MS, Mizusawa N, Kumano A et al (2018) Metagenomic analysis using 16S ribosomal RNA genes of a bacterial community in an urban stream, the Tama River, Tokyo. Fish Sci 84:563–577. https://doi.org/10.1007/s12562-018-1193-6
Roberts TL, Bettany JR (1985) The influence of topography on the nature and distribution of soil sulfur across a narrow environmental gradient. Canadian J Soil Sci 65(3):419–434. https://doi.org/10.4141/cjss85-046
Rodríguez AA, Puente-Sánchez F, Avendaño R et al (2019) Thermoplasmatales and sulfur-oxidizing bacteria dominate the microbial community at the surface water of a CO2-rich hydrothermal spring located in Tenorio Volcano National Park. Costa Rica Extremophiles 23(2):177–187. https://doi.org/10.1007/s00792-018-01072-6
Roost JV, Daae FL, Steen IH et al (2018) Distribution patterns of Iron-Oxidizing Zeta- and Beta-Proteobacteria from different environmental settings at the Jan Mayen Vent Fields. Front Microbiol 9:3008. https://doi.org/10.3389/fmicb.2018.03008
Roy S, Roy M (2019) Characterization of plant growth promoting feature of a neutromesophilic, facultatively chemolithoautotrophic, sulphur oxidizing bacterium Delftia sp. strain SR4 isolated from coal mine spoil. Int J Phytoremediation 21(6):531–540. https://doi.org/10.1080/15226514.2018.1537238
SamKamaleson A, Gonsalves MJ (2019) Role of sulfur-oxidizing bacteria on the ecology in tropical mangrove sediments. Reg Stud Mar Sci 28:100574. https://doi.org/10.1016/j.rsma.2019.100574
Sang S, Zhang X, Dai H et al (2018) Diversity and predictive metabolic pathways of the prokaryotic microbial community along a groundwater salinity gradient of the Pearl River Delta. China Sci Rep 8(1):17317. https://doi.org/10.1038/s41598-018-35350-2
See-Too WS, Ambrose M, Malley R et al (2019) Pandoraea fibrosis sp. nov., a novel Pandoraea species isolated from clinical respiratory samples. Int J Syst Evo Microbiol 69(3):645–651. https://doi.org/10.1099/ijsem.0.003147
Siddiqui FA, Trudel S, Goen AE et al (2022) Draft Genome Sequence of Xanthobacter aminoxidans ATCC BAA-299T. Microbiol Resour Announc 11(8):e00548-e522. https://doi.org/10.1128/mra.00548-22
Speck JJ, James EK, Sugawara M et al (2019) An alkane sulfonate monooxygenase Is required for symbiotic nitrogen fixation by Bradyrhizobium diazoefficiens (syn. Bradyrhizobium japonicum) USDA110T. Appl Environ Microbiol 85(24):e01552-01519. https://doi.org/10.1128/aem.01552-19
Su J, Wang Z, Huang T et al (2020) Simultaneous removal of nitrate, phosphorous and cadmium using a novel multifunctional biomaterial immobilized aerobic strain Proteobacteria Cupriavidus H29. Biores Technol 307:123196. https://doi.org/10.1016/j.biortech.2020.123196
Sugawara M, Shah GR, Sadowsky MJ et al (2011) Expression and functional roles of Bradyrhizobium japonicum genes involved in the utilization of inorganic and organic sulfur compounds in free-living and symbiotic conditions. Mol Plant-Microbe Interact: MPMI 24(4):451–457. https://doi.org/10.1094/mpmi-08-10-0184
Tai X, Li R, Zhang B et al (2020) Pollution gradients altered the bacterial community composition and stochastic process of rural polluted ponds. Microorganisms 8(2):311. https://doi.org/10.3390/microorganisms8020311
P TT, Hatamoto M, Aoki M et al (2022) Effect of inoculum sources on autotrophic nitrogen removal in anaerobic hollow fiber membrane reactors. Environ Technol Innov, 26, 102375. https://doi.org/10.1016/j.eti.2022.102375
Tian T, Yu H (2019) Denitrification with non-organic electron donor for treating low C/N ratio wastewaters. Biores Technol 299:122686. https://doi.org/10.1016/j.biortech.2019.122686
Tian H, Gao P, Chen Z et al (2017) Compositions and abundances of sulfate-reducing and sulfur-oxidizing microorganisms in water-flooded petroleum reservoirs with different temperatures in China. Front Microbiol 8:143. https://doi.org/10.3389/fmicb.2017.00143
Tourna M, Maclean P, Condron L et al (2014) Links between sulphur oxidation and sulphur-oxidising bacteria abundance and diversity in soil microcosms based on soxB functional gene analysis. FEMS Microbiol Ecol 88(3):538–549. https://doi.org/10.1111/1574-6941.12323
Tourova TP, Slobodova NV, Bumazhkin BK et al (2013) Analysis of community composition of sulfur-oxidizing bacteria in hypersaline and soda lakes using soxB as a functional molecular marker. FEMS Microbiol Ecol 84(2):280–289. https://doi.org/10.1111/1574-6941.12056
Tyc O, Kulkarni P, Ossowicki A et al (2023) Exploring the interspecific interactions and the metabolome of the soil isolate Hylemonella gracilis. Msystems 8(1):e00574-e522. https://doi.org/10.1128/msystems.00574-22
Vavourakis CD, Andrei AS, Mehrshad M et al (2018) A metagenomics roadmap to the uncultured genome diversity in hypersaline soda lake sediments. Microbiome 6(1):168. https://doi.org/10.1186/s40168-018-0548-7
Wang P, Chen B, Yuan R et al (2016) Characteristics of aquatic bacterial community and the influencing factors in an urban river. Sci Total Environ 569–570:382–389. https://doi.org/10.1016/j.scitotenv.2016.06.130
Wei Y, Yuan X (2015) Studies on silica-scaled chrysophytes from the Daxinganling Mountains and Wudalianchi Lake Regions. China Nova Hedwigia 101(3):299–312. https://doi.org/10.1127/novahedwigia/2015/0271
Williams P, Whitfield M, Biggs J et al (2004) Comparative biodiversity of rivers, streams, ditches and ponds in an agricultural landscape in Southern England. Biol Cons 115(2):329–341. https://doi.org/10.1016/S0006-3207(03)00153-8
Won M, Heo J, Lee D et al (2023) Melaminivora suipulveris sp. nov., isolated from pigpen dust. Int J Syst Evol Microbiol 73(1):005701. https://doi.org/10.1099/ijsem.0.005701
Xia Y, Lü C, Hou N et al (2017) Sulfide production and oxidation by heterotrophic bacteria under aerobic conditions. ISME J 11(12):2754–2766. https://doi.org/10.1038/ismej.2017.125
Xing W, Hu H, Zhang Y et al (2020) Magnetotactic bacteria diversity of and magnetism contribution to sediment in Wudalianchi volcanic barrier lakes, NE China. Sci Total Environ 718:137348. https://doi.org/10.1016/j.scitotenv.2020.137348
Xu S, Zheng G, Nakai S, i, et al (2013) Hydrothermal He and CO2 at Wudalianchi intra-plate volcano, NE China. J Asian Earth Sci 62:526–530. https://doi.org/10.1016/j.jseaes.2012.11.001
Yang Z, Liu Z, Sklodowska A et al (2021) Microbiological sulfide removal-from microorganism isolation to treatment of industrial effluent. Microorganisms 9(3):611. https://doi.org/10.3390/microorganisms9030611
Yusof HM, Halimi MS, Samsudin AA (2019) Reduction of hydrogen sulphide in chicken manure by immobilized sulphur oxidising bacteria isolated from hot spring. Korean J Microbiol Biotechnol 47(1):116–124. https://doi.org/10.4014/mbl.1801.01005
Zeng Y, Yang C (2018) Vertical distribution of total carbon, nitrogen and phosphorus in sediments of Drug Spring Lake, Wudalianchi. IOP Conf Ser Earth Environ Sci 113:012196. https://doi.org/10.1088/1755-1315/113/1/012196
Zhang Y, Wang X, Zhen Y et al (2017) Microbial diversity and cStructure of ulfate-reducing and sulfur-oxidizing bacteria in sediment cores from the East China Sea. Front Microbiol 8:2133. https://doi.org/10.3389/fmicb.2017.02133
Zhang H, Yang Y, Qi H et al (2018a) Hydrochemical evolution of rare cold mineral waters in the Wudalianchi UNESCO Global Geopark. China Environ Earth Sci 77(10):360. https://doi.org/10.1007/s12665-018-7543-y
Zhang H, Yang Y, Zou J et al (2018b) The sources and dispersal of nitrate in multiple waters, constrained by multiple isotopes, in the Wudalianchi region, northeast China. Environ Sci Pollut Res Int 25(24):24348–24361. https://doi.org/10.1007/s11356-018-2490-4
Zhao W, Guo Z, Lei M et al (2019) Volcanogenic CO2 degassing in the songliao Continental rift system. NE China Geofluids 2019(6):1–14. https://doi.org/10.1155/2019/8053579
Zou J, Yang Y, Jia S et al (2019) The sources and biogeochemical cycling of carbon in the Wudalianchi UNESCO Geopark volcanic system in Northeast China. Environ Sci Pollut Res Int 26(3):2918–2928. https://doi.org/10.1007/s11356-018-3840-y
Zou J, Song Z, Yamin D (2021) Water-source contributions to barrier lakes and water-rock interactions in the Wudalianchi volcanic area. Northeast China Water Supply 21(8):4276–4286. https://doi.org/10.2166/ws.2021.177
Funding
This work was supported by the Heilongjiang Provincial Key Research and Development Program Guidance Projects (GZ20220051), Heilongjiang Provincial Natural Science Foundation of China (LH2020C079), Talent Training Program under Special Funds Supporting the Development of Local Universities from the Central Finance (HFBE[2019]465), Heilongjiang Bayi Agricultural University Support Program for San Heng San Zong (ZRCQC202008).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Lei Yang, Tao Liu, Hong Pan, Shuang Zhang, Xindi Sun, Weidong Wang1and Lei Yan. The first draft of the manuscript was written by Lirong Geng and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This is an observational study. The Heilongjiang Bayi Agricultural University Research Ethics Committee has confirmed that no ethical approval is required.
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Geng, L., Yang, L., Liu, T. et al. Higher diversity of sulfur-oxidizing bacteria based on soxB gene sequencing in surface water than in spring in Wudalianchi volcanic group, NE China. Int Microbiol (2024). https://doi.org/10.1007/s10123-024-00526-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10123-024-00526-6