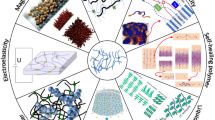
Abstract
The development of biopolymers for biomedical applications has traditionally been based on new chemistries. However, there is growing recognition that the biological responses can be regulated by the physical as well as the chemical properties of biomaterials. Understanding the biophysicochemical principles regarding biopolymers is thereby of great importance in the generation of advanced biomaterials. Herein, this review article seeks to provide a conceptual framework demonstrating how the approaches of tailored computer simulations and theoretical analysis are harnessed to explore the physicochemical principles of biopolymer cellular interactions. We briefly introduce the theoretical and simulation methods used in this field, summarize the typical findings based on these approaches, and describe the correlations between theoretical results and experiments. Finally, the future prospects for the theoretical aspect of biopolymers and their biophysicochemical interactions are discussed. The knowledge might be critical from the perspective of advantageous and safe use of designer biomaterials.
Similar content being viewed by others
References
Yadav, P.; Yadav, H.; Shah, V. G.; Shah, G.; Dhaka, G. Biomedical biopolymers, their origin and evolution in biomedical sciences: a systematic review. J. Clin. Diagn. Res. 2015, 9, ZE21–ZE25.
Jadoun, S.; Riaz, U.; Budhiraja, V. Biodegradable conducting polymeric materials for biomedical applications: a review. Med. Devices Sens. 2021, 4, e10141.
Torchilin, V. P. Multifunctional nanocarriers. Adv. Drug Del. Rev. 2006, 58, 1532–1555.
Gref, R.; Minamitake, Y.; Peracchia, M. T.; Trubetskoy, V.; Torchilin, V.; Langer, R. Biodegradable long-circulating polymeric nanospheres. Science 1994, 263, 1600–1603.
Langer, R.; Tirrell, D. A. Designing materials for biology and medicine. Nature 2004, 428, 487–492.
Mitragotri, S.; Lahann, J. Physical approaches to biomaterial design. Nat. Mater. 2009, 8, 15–23.
Rolland, J. P.; Maynor, B. W.; Euliss, L. E.; Exner, A. E.; Denison, G. M.; DeSimone, J. M. Direct fabrication and harvesting of monodisperse, shape-specific nanobiomaterials. J. Am. Chem. Soc. 2005, 127, 10096–10100.
Champion, J. A.; Mitragotri, S. Role of target geometry in phagocytosis. Proc. Natl. Acad. Sci. U.S.A. 2006, 103, 4930–4934.
Champion, J. A.; Katare, Y. K.; Mitragotri, S. Making polymeric micro- and nanoparticles of complex shapes. Proc. Natl. Acad. Sci. U.S.A. 2007, 104, 11901–11904.
Geng, Y.; Dalhaimer, P.; Cai, S.; Tsai, R.; Tewari, M.; Minko, T.; Discher, D. E. Shape effects of filaments versus spherical particles in flow and drug delivery. Nat. Nanotechnol. 2007, 2, 249–255.
Dalhaimer, P.; Bates, F. S.; Discher, D. E. Single molecule visualization of stable, stiffness-tunable, flow-conforming worm micelles. Macromolecules 2003, 36, 6873–6877.
Ding, H. M.; Ma, Y. Q. Theoretical and computational investigations of nanoparticle-biomembrane interactions in cellular delivery. Small 2015, 11, 1055–71.
Chithrani, B. D.; Chan, W. C. W. Elucidating the mechanism of cellular uptake and removal of protein-coated gold nanoparticles of different sizes and shapes. Nano Lett. 2007, 7, 1542–1550.
Gratton, S. E.; Ropp, P. A.; Pohlhaus, P. D.; Luft, J. C.; Madden, V. J.; Napier, M. E.; DeSimone, J. M. The effect of particle design on cellular internalization pathways. Proc. Natl. Acad. Sci. U.S.A. 2008, 105, 11613–8.
Qu, Z. G.; He, X. C.; Lin, M.; Sha, B. Y.; Shi, X. H.; Lu, T. J.; Xu, F. Advances in the understanding of nanomaterial-biomembrane interactions and their mathematical and numerical modeling. Nanomedicine 2013, 8, 995–1011.
Tian, W. D.; Ma, Y. Q. Theoretical and computational studies of dendrimers as delivery vectors. Chem. Soc. Rev. 2013, 42, 705–727.
Tachikawa, M.; Mochizuki, A. Golgi apparatus self-organizes into the characteristic shape via postmitotic reassembly dynamics. Proc. Natl. Acad. Sci. U.S.A. 2017, 114, 5177–5182.
Huang, C.; Zhang, Y.; Yuan, H.; Gao, H.; Zhang, S. Role of nanoparticle geometry in endocytosis: laying down to stand up. Nano Lett. 2013, 13, 4546–50.
Chen, P.; Yue, H.; Zhai, X.; Huang, Z.; Ma, G. H.; Wei, W.; Yan, L. T. Transport of a graphene nanosheet sandwiched inside cell membranes. Sci. Adv. 2019, 5, eaaw3192.
Escriba, P. V.; Gonzalez-Ros, J. M.; Goni, F. M.; Kinnunen, P. K.; Vigh, L.; Sanchez-Magraner, L.; Fernandez, A. M.; Busquets, X.; Horvath, I.; Barcelo-Coblijn, G. Membranes: a meeting point for lipids, proteins and therapies. J. Cell. Mol. Med. 2008, 12, 829–75.
Ge, Z.; Li, Q.; Wang, Y. Free energy calculation of nanodiamond-membrane association—the effect of shape and surface functionalization. J. Chem. Theory Comput. 2014, 10, 2751–2758.
Tu, Y.; Lv, M.; Xiu, P.; Huynh, T.; Zhang, M.; Castelli, M.; Liu, Z.; Huang, Q.; Fan, C.; Fang, H.; Zhou, R. Destructive extraction of phospholipids from Escherichia coli membranes by graphene nanosheets. Nat. Nanotechnol. 2013, 8, 594–601.
Marrink, S. J.; de Vries, A. H.; Tieleman, D. P. Lipids on the move: Simulations of membrane pores, domains, stalks and curves. Biochim. Biophys. Acta 2009, 1788, 149–168.
Marrink, S. J.; de Vries, A. H.; Mark, A. E. Coarse grained model for semiquantitative lipid simulations. J. Phys. Chem. B 2004, 108, 750–760.
Reynwar, B. J.; Illya, G.; Harmandaris, V. A.; Müller, M. M.; Kremer, K.; Deserno, M. Aggregation and vesiculation of membrane proteins by curvature-mediated interactions. Nature 2007, 447, 461–464.
Pluhackova, K.; Böckmann, R. A. Biomembranes in atomistic and coarse-grained simulations. J. Phys.: Condens. Matter 2015, 27, 323103.
Klauda, J. B.; Venable, R. M.; Freites, J. A.; O’Connor, J. W.; Tobias, D. J.; Mondragon-Ramirez, C.; Vorobyov, I.; MacKerell, A. D.,Jr.; Pastor, R. W. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 2010, 114, 7830–7843.
Provasi, D.; Bortolato, A.; Filizola, M. Exploring molecular mechanisms of ligand recognition by opioid receptors with metadynamics. Biochemistry 2009, 48, 10020–10029.
Berka, K.; Hendrychová, T.; Anzenbacher, P.; Otyepka, M. Membrane position of Ibuprofen agrees with suggested access path entrance to cytochrome P450 2C9 Active Site. J. Phys. Chem. A 2011, 115, 11248–11255.
Schmid, N.; Eichenberger, A. P.; Choutko, A.; Riniker, S.; Winger, M.; Mark, A. E.; van Gunsteren, W. F. Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur. Biophys. J. 2011, 40, 843–856.
Venturoli, M.; Sperotto, M.; Kranenburg, M.; Smit, B. J. P. R. Mesoscopic models of biological membranes. Phys. Rep. 2006, 437, 1–54.
Marrink, S. J.; Risselada, H. J.; Yefimov, S.; Tieleman, D. P.; de Vries, A. H. The MARTINI force field: coarse grained model for biomolecular simulations. J. Phys. Chem. B 2007, 111, 7812–7824.
Español, P.; Warren, P. Statistical mechanics of dissipative particle dynamics. Europhys. Lett. 1995, 30, 191.
Groot, R. D.; Warren, P. B. Dissipative particle dynamics: Bridging the gap between atomistic and mesoscopic simulation. J. Chem. Phys. 1997, 107, 4423–4435.
Hoogerbrugge, P. J.; Koelman, J. M. V. A. Simulating microscopic hydrodynamic phenomena with dissipative particle dynamics. Europhys. Lett. 1992, 19, 155–160.
Cooke, I. R.; Deserno, M. Solvent-free model for self-assembling fluid bilayer membranes: stabilization of the fluid phase based on broad attractive tail potentials. J. Chem. Phys. 2005, 123, 224710.
Cooke, I. R.; Kremer, K.; Deserno, M. Tunable generic model for fluid bilayer membranes. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2005, 72, 011506.
Yuan, H.; Huang, C.; Li, J.; Lykotrafitis, G.; Zhang, S. One-particle-thick, solvent-free, coarse-grained model for biological and biomimetic fluid membranes. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2010, 82, 011905.
Hollingsworth, S. A.; Dror, R. O. Molecular dynamics simulation for all. Neuron 2018, 99, 1129–1143.
Plimpton, S. Fast parallel algorithms for short-range molecular dynamics. J. Comput. Phys. 1995, 117, 1–19.
Chávez Thielemann, H.; Cardellini, A.; Fasano, M.; Bergamasco, L.; Alberghini, M.; Ciorra, G.; Chiavazzo, E.; Asinari, P. From GROMACS to LAMMPS: GRO2LAM. J. Mol. Model. 2019, 25, 147.
Lyubartsev, A. P. Multiscale modeling of lipids and lipid bilayers. Eur. Biophys. J. 2005, 35, 53–61.
Chen, P.; Xu, Z.; Zhu, G.; Dai, X.; Yan, L. T. Cellular uptake of active particles. Phys. Rev. Lett. 2020, 124, 198102.
Lelimousin, M.; Limongelli, V.; Sansom, M. S. P. Conformational changes in the epidermal growth factor receptor: role of the transmembrane domain investigated by coarse-grained metadynamics free energy calculations. J. Am. Chem. Soc. 2016, 138, 10611–10622.
Deng, Z. X.; Li, J. L.; Yuan, B.; Yang, K. Residue-specialized membrane poration kinetics of melittin and its variants: insight from mechanistic landscapes. Commun. Theor. Phys. 2019, 71, 887–902.
Duncan, A. L.; Corey, R. A.; Sansom, M. S. P. Defining how multiple lipid species interact with inward rectifier potassium (Kir2) channels. Proc. Natl. Acad. Sci. U.S.A. 0200, 117, 7803–7813.
Ma, W.; Jiang, X.; Dou, Y.; Zhang, Z.; Li, J.; Yuan, B.; Yang, K. Biophysical impact of lipid a modification caused by mobile colistin resistance gene on bacterial outer membranes. J. Phys. Chem. Lett. 2021, 12, 11629–11635.
Yuan, B.; Liu, J.; Deng, Z.; Wei, L.; Li, W.; Dou, Y.; Chen, Z.; Zhang, C.; Xia, Y.; Wang, J.; Zhang, M.; Yang, K.; Ma, Y.; Kang, Z. A molecular architectural design that promises potent antimicrobial activity against multidrug-resistant pathogens. NPG Asia Mater. 2021, 13, 18.
Deng, Z.; Yuan, B.; Yang, K. Cardiolipin selectively binds to the interface of VsSemiSWEET and regulates its dimerization. J Phys. Chem. Lett. 2021, 12, 1940–1946.
Liu, G.; Xu, Z.; Dai, X.; Zeng, Y.; Wei, Y.; He, X.; Yan, L. T.; Tao, L. De Novo design of entropy-driven polymers resistant to bacterial attachment via multicomponent reactions. J. Am. Chem. Soc. 2021, 143, 17250–17260.
Li, J.; Lu, X. M.; Ma, W. D.; Chen, Z. L.; Sun, S. Q.; Wang, Q. H.; Yuan, B.; Yang, K. Cholesterols work as a molecular regulator of the antimicrobial peptide-membrane interactions. Front. Mol. Biosci. 2021, 8, 638988.
Zhang, S. L.; Li, J.; Lykotrafitis, G.; Bao, G.; Suresh, S. Size-dependent endocytosis of nanoparticles. Adv. Mater. 2009, 21, 419–424.
He, K.; Wei, Y.; Zhang, Z.; Chen, H.; Yuan, B.; Pang, H. B.; Yang, K. Membrane-curvature-mediated co-endocytosis of bystander and functional nanoparticles. Nanoscale 2021, 13, 9626–9633.
Wei, Y.; Chen, H.; Li, Y. X.; He, K.; Yang, K.; Pang, H. B. Synergistic entry of individual nanoparticles into mammalian cells driven by free energy decline and regulated by their sizes. ACS Nano 2022, 16, 5885–5897.
Yang, K.; Ma, Y. Q. Computer simulation of the translocation of nanoparticles with different shapes across a lipid bilayer. Nat. Nanotechnol. 2010, 5, 579–583.
Ding, H. M.; Tian, W. D.; Ma, Y. Q. Designing nanoparticle translocation through membranes by computer simulations. ACS Nano 2012, 6, 1230–1238.
Wang, Y.-F.; Zhang, Q.; Tian, F.; Wang, H.; Wang, Y.; Ma, X.; Huang, Q.; Cai, M.; Ji, Y.; Wu, X.; Gan, Y.; Yan, Y.; Dawson, K. A.; Guo, S.; Zhang, J.; Shi, X.; Shan, Y.; Liang, X. J. Spatiotemporal tracing of the cellular internalization process of rod-shaped nanostructures. ACS Nano 2022, 16, 4059–4071.
Ding, H.; Li, J.; Chen, N.; Hu, X.; Yang, X.; Guo, L.; Li, Q.; Zuo, X.; Wang, L.; Ma, Y.; Fan, C. DNA nanostructure-programmed like-charge attraction at the cell-membrane interface. ACS Cent. Sci. 2018, 4, 1344–1351.
Li, Y.; Yue, T.; Yang, K.; Zhang, X. Molecular modeling of the relationship between nanoparticle shape anisotropy and endocytosis kinetics. Biomaterials 2012, 33, 4965–4973.
Shi, X.; von dem Bussche, A.; Hurt, R. H.; Kane, A. B.; Gao, H. Cell entry of one-dimensional nanomaterials occurs by tip recognition and rotation. Nat. Nanotechnol. 2011, 6, 714–719.
Guo, R.; Mao, J.; Yan, L. T. Computer simulation of cell entry of graphene nanosheet. Biomaterials 2013, 34, 4296–4301.
Mao, J.; Chen, P.; Liang, J.; Guo, R.; Yan, L. T. Receptor-mediated endocytosis of two-dimensional nanomaterials undergoes flat vesiculation and occurs by revolution and self-rotation. ACS Nano 2016, 10, 1493–1502.
Lu, X. M.; Liu, J. J.; Gou, L.; Li, J. L.; Yuan, B.; Yang, K.; Ma, Y. Q. Designing melittin-graphene hybrid complexes for enhanced antibacterial activity. Adv. Healthc. Mater. 2019, 8, 1801521.
Qian, R.; Maiti, D.; Zhong, J.; Xiong, S. S.; Zhou, H. L.; Zhu, R.; Wan, J. M.; Yang, K. Multifunctional nano-graphene based nanocomposites for multimodal imaging guided combined radioisotope therapy and chemotherapy. Carbon 2019, 149, 55–62.
Li, Y.; Feng, D.; Zhang, X.; Cao, D. Design strategy of cell-penetrating copolymers for high efficient drug delivery. Biomaterials 2015, 52, 171–179.
Ou, L.; Chen, H.; Yuan, B.; Yang, K. Membrane-specific binding of 4 nm lipid nanoparticles mediated by an entropy-driven interaction mechanism. ACS Nano 2022, 16, 18090–18100.
Tang, Y.; Bera, S.; Yao, Y.; Zeng, J.; Lao, Z.; Dong, X.; Gazit, E.; Wei, G. Prediction and characterization of liquid-liquid phase separation of minimalistic peptides. Cell Rep. Phys. Sci. 2021, 2, 100579.
Dong, X.; Bera, S.; Qiao, Q.; Tang, Y.; Lao, Z.; Luo, Y.; Gazit, E.; Wei, G. Liquid-liquid phase separation of tau protein is encoded at the monomeric level. J. Phys. Chem. Lett. 2021, 12, 2576–2586.
Ren, C.-L.; Shan, Y.; Zhang, P.; Ding, H. M.; Ma, Y. Q. Uncovering the molecular mechanism for dual effect of ATP on phase separation in FUS solution. Sci. Adv. 2022, 8, eabo7885.
Sun, J. S.; Zhang, L.; Wang, J. L.; Feng, Q.; Liu, D. B.; Yin, Q. F.; Xu, D. Y.; Wei, Y. J.; Ding, B. Q.; Shi, X. H.; Jiang, X. Y. Tunable rigidity of (polymeric core)-(lipid shell) nanoparticles for regulated cellular uptake. Adv. Mater. 2015, 27, 1402–1407.
Yi, X.; Shi, X.; Gao, H. Cellular uptake of elastic nanoparticles. Phys. Rev. Lett. 2011, 107, 098101.
Hu, C.-M. J.; Fang, R. H.; Wang, K.-C.; Luk, B. T.; Thamphiwatana, S.; Dehaini, D.; Nguyen, P.; Angsantikul, P.; Wen, C. H.; Kroll, A. V.; Carpenter, C.; Ramesh, M.; Qu, V.; Patel, S. H.; Zhu, J.; Shi, W.; Hofman, F. M.; Chen, T. C.; Gao, W.; Zhang, K.; Chien, S.; Zhang, L. Nanoparticle biointerfacing by platelet membrane cloaking. Nature 2015, 526, 118–121.
Xia, Q. S.; Zhu, T.; Jiang, Z. Y.; Ding, H. M.; Ma, Y. Q. Enhancing the targeting ability of nanoparticles via protected copolymers. Nanoscale 2020, 12, 7804–7813.
Xu, C.; Ma, W.; Wang, K.; He, K.; Chen, Z.; Liu, J.; Yang, K.; Yuan, B. Correlation between single-molecule dynamics and biological functions of antimicrobial peptide melittin. J. Phys. Chem. Lett. 2020, 11, 4834–4841.
Chen, P.; Huang, Z.; Liang, J.; Cui, T.; Zhang, X.; Miao, B.; Yan, L. T. Diffusion and directionality of charged nanoparticles on lipid bilayer membrane. ACS Nano 2016, 10, 11541–11547.
Dai, X.; Zhu, Z.; Li, Y.; Yang, B.; Xu, J. F.; Dong, Y.; Zhou, X.; Yan, L. T.; Liu, D. “Shutter” effects enhance protein diffusion in dynamic and rigid molecular networks. J. Am. Chem. Soc. 2022, 144, 19017–19025.
Cho, W. K.; Spille, J. H.; Hecht, M.; Lee, C.; Li, C.; Grube, V.; Cisse, I. I. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 2018, 361, 412–415.
Franzmann, T. M.; Jahnel, M.; Pozniakovsky, A.; Mahamid, J.; Holehouse, A. S.; Nuske, E.; Richter, D.; Baumeister, W.; Grill, S. W.; Pappu, R. V.; Hyman, A. A.; Alberti, S. Phase separation of a yeast prion protein promotes cellular fitness. Science 2018, 359, eaao5654.
Zhang, H.; Ji, X.; Li, P.; Liu, C.; Lou, J.; Wang, Z.; Wen, W.; Xiao, Y.; Zhang, M.; Zhu, X. Liquid-liquid phase separation in biology: mechanisms, physiological functions and human diseases. Sci. China: Life Sci. 2020, 63, 953–985.
Li, P.; Banjade, S.; Cheng, H.-C.; Kim, S.; Chen, B.; Guo, L.; Llaguno, M.; Hollingsworth, J. V.; King, D. S.; Banani, S. F.; Russo, P. S.; Jiang, Q.-X.; Nixon, B. T.; Rosen, M. K. Phase transitions in the assembly of multivalent signalling proteins. Nature 2012, 483, 336–340.
Patel, A.; Lee, Hyun O.; Jawerth, L.; Maharana, S.; Jahnel, M.; Hein, Marco Y.; Stoynov, S.; Mahamid, J.; Saha, S.; Franzmann, Titus M.; Pozniakovski, A.; Poser, I.; Maghelli, N.; Royer, Loic A.; Weigert, M.; Myers, Eugene W.; Grill, S.; Drechsel, D.; Hyman, Anthony A.; Alberti, S. A Liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 2015, 162, 1066–1077.
Murray, D. T.; Kato, M.; Lin, Y.; Thurber, K. R.; Hung, I.; McKnight, S. L.; Tycko, R. Structure of FUS protein fibrils and its relevance to self-assembly and phase separation of low-complexity domains. Cell 2017, 171, 615–627.
Wei, X.; Zhou, J.; Wang, Y.; Meng, F. Modeling elastically mediated liquid-liquid phase separation. Phys. Rev. Lett. 2020, 125, 268001.
Zhang, L.; Granick, S. Slaved diffusion in phospholipid bilayers. Proc. Natl. Acad. Sci. USA 2005, 102, 9118–9121.
Wang, B.; Anthony, S. M.; Bae, S. C.; Granick, S. Anomalous yet Brownian. Proc. Natl. Acad. Sci. USA 2009, 106, 15160–15164.
Barkai, E.; Garini, Y.; Metzler, R. Strange kinetics of single molecules in living cells. Phys. Today 2012, 65, 29–35.
Wang, B.; Kuo, J.; Bae, S. C.; Granick, S. When Brownian diffusion is not Gaussian. Nat. Mater. 2012, 11, 481–485.
Saffman, P. G.; Delbruck, M. Brownian-motion in biological-membranes. Proc. Natl. Acad. Sci. U.S.A. 1975, 72, 3111–3113.
Javanainen, M.; Martinez-Seara, H.; Metzler, R.; Vattulainen, I. Diffusion of integral membrane proteins in protein-rich membranes. J. Phys. Chem. Lett. 2017, 8, 4308–4313.
Jeon, J. H.; Monne, H. M. S.; Javanainen, M.; Metzler, R. Anomalous diffusion of phospholipids and cholesterols in a lipid bilayer and its origins. Phys. Rev. Lett. 2012, 109, 188103.
Jeon, J. H.; Javanainen, M.; Martinez-Seara, H.; Metzler, R.; Vattulainen, I. Protein crowding in lipid bilayers gives rise to non-Gaussian anomalous lateral diffusion of phospholipids and proteins. Phys. Rev. X 2016, 6, 021006.
Ji, Q. J.; Yuan, B.; Lu, X. M.; Yang, K.; Ma, Y. Q. Controlling the nanoscale rotational behaviors of nanoparticles on the cell membranes: a computational model. Small 2016, 12, 1140–1146.
Osaki, F.; Kanamori, T.; Sando, S.; Sera, T.; Aoyama, Y. A Quantum dot conjugated sugar ball and its cellular uptake. on the size effects of endocytosis in the subviral region. J. Am. Chem. Soc. 2004, 126, 6520–6521.
Chithrani, B. D.; Ghazani, A. A.; Chan, W. C. W. Determining the size and shape dependence of gold nanoparticle uptake into mammalian cells. Nano Lett. 2006, 6, 662–668.
Gao, H. J.; Shi, W. D.; Freund, L. B. Mechanics of receptor-mediated endocytosis. Proc. Natl. Acad. Sci. U.S.A. 2005, 102, 9469–9474.
Gu, Y.; Sun, W.; Wang, G. F.; Zimmermann, M. T.; Jernigan, R. L.; Fang, N. Revealing rotational modes of functionalized gold nanorods on live cell membranes. Small 2013, 9, 785–792.
Chu, Z.; Zhang, S.; Zhang, B.; Zhang, C.; Fang, C. Y.; Rehor, I.; Cigler, P.; Chang, H.-C.; Lin, G.; Liu, R.; Li, Q. Unambiguous observation of shape effects on cellular fate of nanoparticles. Sci. Rep. 2014, 4, 4495.
Cho, E. C.; Xie, J.; Wurm, P. A.; Xia, Y. Understanding the role of surface charges in cellular adsorption versus internalization by selectively removing gold nanoparticles on the cell surface with a I-2/KI Etchant. Nano Lett. 2009, 9, 1080–1084.
Ding, H. M.; Ma, Y. Q. Interactions between Janus particles and membranes. Nanoscale 2012, 4, 1116–1122.
Gao, Y.; Yu, Y. How half-coated janus particles enter cells. J. Am. Chem. Soc. 2013, 135, 19091–19094.
Ding, H. M.; Ma, Y. Q. Role of physicochemical properties of coating ligands in receptor-mediated endocytosis of nanoparticles. Biomaterials 2012, 33, 5798–5802.
Chelladurai, R.; Debnath, K.; Jana, N. R.; Basu, J. K. Nanoscale heterogeneities drive enhanced binding and anomalous diffusion of nanoparticles in model biomembranes. Langmuir 2018, 34, 1691–1699.
Salvati, A.; Pitek, A. S.; Monopoli, M. P.; Prapainop, K.; Bombelli, F. B.; Hristov, D. R.; Kelly, P. M.; Åberg, C.; Mahon, E.; Dawson, K. A. Transferrin-functionalized nanoparticles lose their targeting capabilities when a biomolecule corona adsorbs on the surface. Nat. Nanotechnol. 2013, 8, 137–143.
Ding, H. M.; Ma, Y. Q. Computer simulation of the role of protein corona in cellular delivery of nanoparticles. Biomaterials 2014, 35, 8703–8710.
Hartmann, R.; Weidenbach, M.; Neubauer, M.; Fery, A.; Parak, W. J. Stiffness-dependent in vitro uptake and lysosomal acidification of colloidal particles. Angew. Chem. Int. Ed. 2015, 54, 1365–1368.
Dai, X.; Zhang, X.; Gao, L.; Yan, L. T. Superentropy effect and macromolecular entropy control strategy. Acta Polymerica Sinica (in Chinese) 2021, 52, 1076–1099.
Beningo, K. A.; Wang, Y. L. Fc-receptor-mediated phagocytosis is regulated by mechanical properties of the target. J. Cell Sci. 2002, 115, 849–856.
Banquy, X.; Suarez, F.; Argaw, A.; Rabanel, J. M.; Grutter, P.; Bouchard, J. F.; Hildgen, P.; Giasson, S. Effect of mechanical properties of hydrogel nanoparticles on macrophage cell uptake. Soft Matter 2009, 5, 3984–3991.
Dai, X.; Zhang, X.; Gao, L.; Xu, Z.; Yan, L. T. Topology mediates transport of nanoparticles in macromolecular networks. Nat. Commun. 2022, 13, 4094.
Guo, R.; Mao, J.; Yan, L. T. Unique dynamical approach of fully wrapping dendrimer-like soft nanoparticles by lipid bilayer membrane. ACS Nano 2013, 7, 10646–10653.
Teng, Z. G.; Wang, C. Y.; Tang, Y. X.; Li, W.; Bao, L.; Zhang, X. H.; Su, X. D.; Zhang, F.; Zhang, J. J.; Wang, S. J.; Zhao, D. Y.; Lu, G. M. Deformable hollow periodic mesoporous organosilica nanocapsules for significantly improved cellular uptake. J. Am. Chem. Soc. 2018, 140, 1385–1393.
Acknowledgments
This work was financially supported by the National Natural Science Foundation of China (Nos. 22025302, 21873053 and 32230063). K.Y. thanks the financial support from the open research fund of Songshan Lake Materials Laboratory (No. 2021SLABFK10).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare no interest conflict.
Additional information
Biographies
Kai Yang received his Ph.D. degree in soft matter physics from Nanjing University in 2009. He was appointed as an Associate Professor at Soochow University in 2010 and was promoted as a full Professor in 2015. During 2014–2015, he was a visiting professor at the Cornell University with Prof. Gerald W. Feigenson. His research interests include biophysics, Nano-Bio interactions, and simulations and theory of soft matter systems.
Li-Tang Yan received his Ph.D. in polymer physics and chemistry at Tsinghua University in 2007. Then he went to Bayreuth University in Germany as a Humboldt Research Fellowship. In 2010, he joined Prof. Anna Balazs’ group at University of Pittsburgh in USA as a Postdoctoral Research Fellowship. He returned to Tsinghua University as a faculty from May 2011, and now is a full professor with tenure. He leads a polymer theory and physics group working on polymer theory and simulation, soft condense matter physics, biophysics and nonequilibrium physics.
Rights and permissions
About this article
Cite this article
Wan, HX., Xu, D., Dong, XW. et al. Insight into Biophysicochemical Principles of Biopolymers through Simulation and Theory. Chin J Polym Sci 41, 1342–1354 (2023). https://doi.org/10.1007/s10118-023-2954-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10118-023-2954-y