Abstract
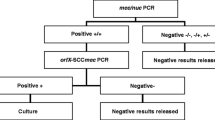
Staphylococcus aureus (SA) nasal carriage screening is usually based on either culture or molecular biology. The aim of the study was to evaluate the performance of the Panther Fusion® MRSA Assay (PF) that proposes a complete automation of the molecular screening for MSSA and MRSA carriage. Four hundred thirty-four nasal samples collected on ESwab™ were screened using PF. Results were compared with standard culture on BBL™ CHROMagar™ Staph aureus and chromID® MRSA agar. Discordant results were analyzed with additional techniques: Xpert SA Nasal Complete on GeneXpert (GX), culture on selective agar after 24 h in broth enrichment, and, if necessary, characterization of mec gene and SCCmec cassette using DNA microarray. The PF presented an overall agreement of 97.5% for SA detection and 97.9% for MRSA detection. Furthermore, 7.1% (31/434) of the samples were SA-negative in primary culture but SA-positive using PF and GX, confirming the greater sensitivity of molecular tests compared with culture. Of note, 4 out of 30 MRSA-positive samples were not detected due to an atypical SCCmec cassette, while 2 samples were falsely detected as MRSA due to co-colonization with a MSSA drop-out strain and a methicillin-resistant coagulase-negative staphylococcal strain. Considering all results, the PF instrument appears as a reliable and rapid (< 3 h) package for MSSA/MRSA nasal screening. This technology using random access capability and direct sampling of the primary container is innovative and corresponds therefore to a new step in complete molecular biology automation in bacteriology.

Similar content being viewed by others
References
Wertheim HFL, Melles DC, Vos MC, van Leeuwen W, van Belkum A, Verbrugh HA et al (2005) The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect Dis 5:751–762. https://doi.org/10.1016/S1473-3099(05)70295-4
Wertheim HFL, Vos MC, Ott A, van Belkum A, Voss A, Kluytmans JAJW et al (2004) Risk and outcome of nosocomial Staphylococcus aureus bacteraemia in nasal carriers versus non-carriers. Lancet 364:703–705. https://doi.org/10.1016/S0140-6736(04)16897-9
Lowy FD (1998) Staphylococcus aureus infections. N Engl J Med 339:520–532. https://doi.org/10.1056/NEJM199808203390806
Chambers HF, Deleo FR (2009) Waves of resistance: Staphylococcus aureus in the antibiotic era. Nat Rev Microbiol 7:629–641. https://doi.org/10.1038/nrmicro2200
Lim D, Strynadka NCJ (2002) Structural basis for the beta lactam resistance of PBP2a from methicillin-resistant Staphylococcus aureus. Nat Struct Biol 9:870–876. https://doi.org/10.1038/nsb858
García-Álvarez L, Holden MTG, Lindsay H, Webb CR, Brown DFJ, Curran MD et al (2011) Meticillin-resistant Staphylococcus aureus with a novel mecA homologue in human and bovine populations in the UK and Denmark: a descriptive study. Lancet Infect Dis 11:595–603. https://doi.org/10.1016/S1473-3099(11)70126-8
Gómez-Sanz E, Schwendener S, Thomann A, Gobeli Brawand S, Perreten V (2015) First staphylococcal cassette chromosome mec containing a mecB-carrying gene complex independent of transposon Tn6045 in a Macrococcus canis isolate from a canine infection. Antimicrob Agents Chemother 59:4577–4583. https://doi.org/10.1128/AAC.05064-14
Comparison of recommendations in national/regional Guidelines for prevention and control of MRSA in thirteen European countries | International Journal of Infection Control n.d.
Gestion préopératoire du risque infectieux - Mise à jour de la conférence de consensus 2004. SF2H n.d. https://sf2h.net/publications/gestion-preoperatoire-risque-infectieux-mise-a-jour-de-conference-de-consensus (accessed November 4, 2018)
Prévention de la transmission croisée : Précautions complémentaires contact. SF2H n.d. https://sf2h.net/publications/prevention-de-transmission-croisee-precautions-complementaires-contact (accessed November 4, 2018)
Morris K, Wilson C, Wilcox MH (2012) Evaluation of chromogenic meticillin-resistant Staphylococcus aureus media: sensitivity versus turnaround time. J Hosp Infect 81:20–24. https://doi.org/10.1016/j.jhin.2012.02.003
Paule SM, Mehta M, Hacek DM, Gonzalzles T-M, Robicsek A, Peterson LR (2009) Chromogenic media vs real-time PCR for nasal surveillance of methicillin-resistant Staphylococcus aureus: impact on detection of MRSA-positive persons. Am J Clin Pathol 131:532–539. https://doi.org/10.1309/AJCP18ONZUTDUGAQ
Trouillet-Assant S, Rasigade J-P, Lustig S, Lhoste Y, Valour F, Guerin C et al (2013) Ward-specific rates of nasal cocolonization with methicillin-susceptible and -resistant Staphylococcus spp. and potential impact on molecular methicillin-resistant Staphylococcus aureus screening tests. J Clin Microbiol 51:2418–2420. https://doi.org/10.1128/JCM.00491-13
Beck ET, Buchan BW, Reymann GC, Ledeboer NA (2016) Comparison of ESwab and wound fiber swab specimen collection devices for use with Xpert SA nasal complete assay. J Clin Microbiol 54:1904–1906. https://doi.org/10.1128/JCM.00449-16
Murakami K, Minamide W, Wada K, Nakamura E, Teraoka H, Watanabe S (1991) Identification of methicillin-resistant strains of staphylococci by polymerase chain reaction. J Clin Microbiol 29:2240–2244
Stegger M, Andersen PS, Kearns A, Pichon B, Holmes MA, Edwards G et al (2012) Rapid detection, differentiation and typing of methicillin-resistant Staphylococcus aureus harbouring either mecA or the new mecA homologue mecA(LGA251). Clin Microbiol Infect 18:395–400. https://doi.org/10.1111/j.1469-0691.2011.03715.x
Monecke S, Slickers P, Ehricht R (2008) Assignment of Staphylococcus aureus isolates to clonal complexes based on microarray analysis and pattern recognition. FEMS Immunol Med Microbiol 53:237–251. https://doi.org/10.1111/j.1574-695X.2008.00426.x
Tenover FC, Tickler IA, Le VM, Dewell S, Mendes RE, Goering RV (2019) Updating molecular diagnostics for detecting methicillin-susceptible and methicillin-resistant Staphylococcus aureus isolates in blood culture bottles. J Clin Microbiol 57. https://doi.org/10.1128/JCM.01195-19
Shore AC, Rossney AS, Brennan OM, Kinnevey PM, Humphreys H, Sullivan DJ et al (2011) Characterization of a novel arginine catabolic mobile element (ACME) and staphylococcal chromosomal cassette mec composite island with significant homology to Staphylococcus epidermidis ACME type II in methicillin-resistant Staphylococcus aureus genotype ST22-MRSA-IV. Antimicrob Agents Chemother 55:1896–1905. https://doi.org/10.1128/AAC.01756-10
Tunsjø HS, Kalyanasundaram S, Worren MM, Leegaard TM, Moen AEF (2017) High frequency of occupied attB regions in Norwegian Staphylococcus aureus isolates supports a two-step MRSA screening algorithm. Eur J Clin Microbiol Infect Dis 36:65–74. https://doi.org/10.1007/s10096-016-2771-0
Donnio P-Y, Oliveira DC, Faria NA, Wilhelm N, Le Coustumier A, de Lencastre H (2005) Partial excision of the chromosomal cassette containing the methicillin resistance determinant results in methicillin-susceptible Staphylococcus aureus. J Clin Microbiol 43:4191–4193. https://doi.org/10.1128/JCM.43.8.4191-4193.2005
Becker K, Pagnier I, Schuhen B, Wenzelburger F, Friedrich AW, Kipp F et al (2006) Does nasal cocolonization by methicillin-resistant coagulase-negative staphylococci and methicillin-susceptible Staphylococcus aureus strains occur frequently enough to represent a risk of false-positive methicillin-resistant S. aureus determinations by molecular methods? J Clin Microbiol 44:229–231. https://doi.org/10.1128/JCM.44.1.229-231.2006
Huletsky A, Giroux R, Rossbach V, Gagnon M, Vaillancourt M, Bernier M et al (2004) New real-time PCR assay for rapid detection of methicillin-resistant Staphylococcus aureus directly from specimens containing a mixture of staphylococci. J Clin Microbiol 42:1875–1884
Keshtgar MRS, Khalili A, Coen PG, Carder C, Macrae B, Jeanes A et al (2008) Impact of rapid molecular screening for meticillin-resistant Staphylococcus aureus in surgical wards. Br J Surg 95:381–386. https://doi.org/10.1002/bjs.6013
Chowers M, Carmeli Y, Shitrit P, Elhayany A, Geffen K (2015) Cost analysis of an intervention to prevent methicillin-resistant Staphylococcus aureus (MRSA) transmission. PLoS One 10:e0138999. https://doi.org/10.1371/journal.pone.0138999
Cunningham R, Jenks P, Northwood J, Wallis M, Ferguson S, Hunt S (2007) Effect on MRSA transmission of rapid PCR testing of patients admitted to critical care. J Hosp Infect 65:24–28. https://doi.org/10.1016/j.jhin.2006.09.019
Hardy K, Price C, Szczepura A, Gossain S, Davies R, Stallard N et al (2010) Reduction in the rate of methicillin-resistant Staphylococcus aureus acquisition in surgical wards by rapid screening for colonization: a prospective, cross-over study. Clin Microbiol Infect 16:333–339. https://doi.org/10.1111/j.1469-0691.2009.02899.x
Tacconelli E, De Angelis G, de Waure C, Cataldo MA, La Torre G, Cauda R (2009) Rapid screening tests for meticillin-resistant Staphylococcus aureus at hospital admission: systematic review and meta-analysis. Lancet Infect Dis 9:546–554. https://doi.org/10.1016/S1473-3099(09)70150-1
Harbarth S, Masuet-Aumatell C, Schrenzel J, Francois P, Akakpo C, Renzi G et al (2006) Evaluation of rapid screening and pre-emptive contact isolation for detecting and controlling methicillin-resistant Staphylococcus aureus in critical care: an interventional cohort study. Crit Care 10:R25. https://doi.org/10.1186/cc3982
Wassenberg MWM, Kluytmans JJW, Bosboom RW, Buiting AGM, van Elzakker EPM, Melchers WJG et al (2011) Rapid diagnostic testing of methicillin-resistant Staphylococcus aureus carriage at different anatomical sites: costs and benefits of less extensive screening regimens. Clin Microbiol Infect 17:1704–1710. https://doi.org/10.1111/j.1469-0691.2011.03502.x
Acknowledgments
We thank Dr. Muriel Rabilloud (Hospices Civils de Lyon) for statistical expertise and Dr. Philip Robinson (Hospices Civils de Lyon) for help in manuscript preparation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Results of this study were presented at the 38th meeting of the RICAI congress (Réunion Interdisciplinaire de Chimiothérapie Anti-Infectieuse) on December 17–18, 2018, in Paris as a paper poster. Results of this study were presented at the 29th ECCMID congress on April 13–16, 2019, in Amsterdam also as a paper poster.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Maurin, E., Ranc, AG., Abad, L. et al. Performance of the Hologic Panther Fusion® MRSA Assay for the nasal screening of methicillin-sensitive and methicillin-resistant Staphylococcus aureus carriage. Eur J Clin Microbiol Infect Dis 39, 2169–2176 (2020). https://doi.org/10.1007/s10096-020-03968-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-020-03968-8