Abstract
Coronavirus disease 2019 (COVID-19), caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is a new infectious disease that first emerged in Hubei province, China, in December 2019, which was found to be associated with a large seafood and animal market in Wuhan. Airway epithelial cells from infected patients were used to isolate a novel coronavirus, named the SARS-CoV-2, on January 12, 2020, which is the seventh member of the coronavirus family to infect humans. Phylogenetic analysis of full-length genome sequences obtained from infected patients showed that SARS-CoV-2 is similar to severe acute respiratory syndrome coronavirus (SARS-CoV) and uses the same cell entry receptor, angiotensin-converting enzyme 2 (ACE2), as SARS-CoV. The possible person-to-person disease rapidly spread to many provinces in China as well as other countries. Without a therapeutic vaccine or specific antiviral drugs, early detection and isolation become essential against novel Coronavirus. In this review, we introduced current diagnostic methods and criteria for the SARS-CoV-2 in China and discuss the advantages and limitations of the current diagnostic methods, including chest imaging and laboratory detection.
Similar content being viewed by others
Introduction
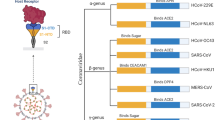
Coronaviruses are unsegmented single-stranded RNA viruses ranging from 26 to 32 kilobases in length, belonging to the subfamily Coronavirinae of the family Coronaviridae of the order Nidovirales [1]. According to the serotype and genomic characteristics, the Coronavirinae subfamily is divided into four major genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus [2]. The former two genera primarily infect mammals, whereas the latter two predominantly infect birds [3]. Coronaviruses mainly cause respiratory and gastrointestinal tract infections; six kinds of human CoVs have been previously identified, including the HCoV-NL63 and the HCoV-229E, which belong to the Alphacoronavirus genus, and the HCoV-OC43, the HCoVHKU1, the severe acute respiratory syndrome coronavirus (SARS-CoV), and the Middle East respiratory syndrome coronavirus (MERS-CoV), which belong to the Betacoronavirus genus [4]. Given the high prevalence and wide distribution of coronaviruses in animals, the large genetic diversity and frequent recombination of their genomes, and increasing human-animal interface activities and frequent cross-species infections, novel coronaviruses are likely to emerge periodically in humans [5].
In December 2019, a group of pneumonia cases was reported at a wholesale seafood market in Wuhan, Hubei province, which was found to be caused by previously unknown Coronaviruses [6]. On December 29, 2019, the local hospitals using a surveillance mechanism for “pneumonia of an unknown etiology,” which was established in the wake of the 2003 severe acute respiratory syndrome (SARS) outbreak, identified the first 4 cases which were all associated with the Huanan (Southern China) Seafood Wholesale Market. On December 31, 2019, the Chinese Center for Disease Control and Prevention (China CDC) dispatched a rapid response team to accompany Hubei provincial and Wuhan city health authorities and to conduct an epidemiologic and etiologic investigation. Similar cases were subsequently reported in Wuhan, and many of these patients did not have contacts with the Huanan Seafood Wholesale Markets or animals. Epidemiological investigation showed that about only 1% of the patients had direct contact with the live-animal market trade, while more than three quarters were local residents of Wuhan or had made contact with people from Wuhan, suggesting a person-to-person transmission of this novel coronavirus was possible [7]. Airway epithelial cells from infected patients were used to isolate a novel coronavirus, temporarily named 2019-nCoV [8], but later, the Coronavirus Research Group (CSG) of the International Committee for the classification of viruses found that the new coronavirus is related to the SARS virus (SARS-CoV) that swept China in 2003. Both belong to a “species” category called severe acute respiratory syndrome-related coronavirus. Therefore, on February 11, 2020, the International Committee for the classification of viruses designated the name of this coronavirus as the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [9]. In addition, the World Health Organization has named the disease caused by the SARS-CoV-2 as coronavirus disease 2019 (COVID-19). The possible person-to-person transmission rapidly spreads to many provinces in China as well as other countries. By February 27, 2020, 78,824 cases were laboratory-confirmed, and 2788 died in China [10]. The current public health emergency is partially similar to the SARS outbreak in southern China in 2002. The two cases share similarities. Both occurred during the winter with initial cases related to an exposure to live animals sold at animal markets, and the amino acid sequence identity between the SARS-CoV-2 and the SARS-CoV S-proteins is 76.47% [11]. The current knowledge of the physical and chemical properties of Coronaviruses is mainly derived from the study of the SARS-CoV and the MERS-CoV. The Coronaviruses are sensitive to exposure to heat (56 °C for 30 min), as well as solvents including ether, 75% ethanol, chlorine-containing disinfectant, peroxyacetic acid, and chloroform. Other lipid solvents can also effectively inactivate the virus except for chlorhexidine [12]. According to Zhong’s latest pilot experiment, 4 out of the 62 stool specimens tested positive to the SARS-CoV-2, suggesting oral-fecal route might have played a role in the rapid transmission of SARS-CoV-2 [7]. However, no cases of transmission via the fecal-oral route have yet been reported for SARS-CoV-2. Contamination of fomite is more likely to be caused by airway/hands. At present, respiratory transmission and direct contact transmission are the main routes for SARS-CoV-2.
Genetic sequence and origin of the SARS-CoV-2
The genome of Coronaviruses, ranging from 26 to 32 kilobases in length, includes a variable number of open reading frames (ORFs) [13]. The SARS-CoV-2 genome was reported to possess 14 ORFs encoding 27 proteins [14]. The spike surface glycoprotein plays an essential role in binding to receptors on the host cell and is crucial for determining host tropism and transmission capacity, mediating receptor binding and membrane fusion [15]. Generally, the spike protein of Coronaviruses is functionally divided into the S1 domain, responsible for receptor binding, and the S2 domain, responsible for cell membrane fusion [16]. The eight accessory proteins (3a, 3b, p6, 7a, 7b, 8b, 9b, and orf14) and four major structural proteins, including the spike surface glycoprotein (S), small envelope protein (E), matrix protein (M), and nucleocapsid protein (N), are located in the 3′-terminus of the SARS-CoV-2 genome [14]. When researchers compare the SARS-CoV-2 with the SARS-CoV at the amino acid level, they found the SARS-CoV-2 was quite similar to the SARS-CoV, but there were some notable differences in the 8a, 8b, and 3b protein [14]. When researchers compared the SARS-CoV-2 with the MERS-CoV, they found that the SARS-CoV-2 was distant from and less related to the MERS-CoVs. From the phylogenetic tree based on whole genomes, the SARS-CoV-2 is parallel to the SARS-like bat CoVs, while the SARS-CoV has descended from the SARS-like bat CoV lineage, indicating that SARS-CoV-2 is closer to the SARS-like bat CoVs than the SARS-CoVs based on of the whole-genome sequence [14]. Analysis of the genome from nine patients’ samples also confirmed that the SARS-CoV-2 was more similar to two SARS-like bat CoVs from Zhoushan in eastern China, bat-SL-CoVZC45 and bat-SL-CoVZXC21, than to the SARS-CoV and the MERS-CoV [17]. At the whole-genome level, the SARS-CoV-2 shares an 87.99% sequence identity with the bat-SL-CoVZC45 and 87.23% sequence identity with the bat-SL-CoVZXC2, less genetically similar to the SARS-CoV (about 79%) and MERS-CoV (about 50%) [17]. At the protein level, the lengths of most of the proteins encoded by the SARS-CoV-2, the bat-SL-CoVZC45, and the bat-SL-CoVZXC21 were similar, with only a few minor insertions or deletions [17]. Although the SARS-CoV-2 was closer to the bat-SL-CoVZC45 and the bat-SL-CoVZXC21 at the whole-genome level, the receptor-binding domain of the SARS-CoV-2 located in lineage B was closer to that of the SARS-CoV [17]. Given the close relationship between the SARS-CoV-2 and the SARS-CoVs or the SARS-like bat CoVs, further studies of the amino acid substitutions in different proteins could explain how the SARS-CoV-2 differs structurally and functionally from the SARS-CoVs and how these differences affect the functionality and pathogenesis of the SARS-CoV-2.
It was reported that 27 of the first 41 infected patients had been exposed to the Huanan Seafood Market [18]. Thus, it was believed that the new coronavirus originated from the Huanan Seafood Market in Wuhan and spread from animal hosts to humans in the process of wildlife trade, transportation, slaughter, and trade. Bats have the most variety of coronaviruses in their bodies and are the hosts of many kinds of coronaviruses, such as the SARS-CoV and the MERS-CoV [19]. The SARS-CoV and the MERS-CoV are considered highly pathogenic, and it is very likely that the SARS-CoV was transmitted from bats to palm civets and the MERS-CoV was transmitted from bats to dromedary camels and finally to humans [20, 21]. Given the high sequence similarity between the SARS-CoV-2 and the SARS-like bat CoVs from Hipposideros bats in China, the natural host of the SARS-CoV-2 may be the Hipposideros bat. The discovery that pangolin coronavirus genomes have 85.5% to 92.4% sequence similarity to SARS-CoV-2 suggests pangolins should be considered as possible hosts in the emergence of SARS-CoV-2 [22].
Diagnosis
According to the seventh edition of Pneumonia Diagnosis and Treatment program for novel coronavirus infection reported by the National Health Commission of the People’s Republic of China, suspected cases were defined as patients having fever or respiratory symptoms, a typical ground-glass opacity chest imaging as well as a history of exposure to wildlife in the Wuhan seafood market, and a travel history or contact with people from Wuhan within 2 weeks of diagnosis [12]. Confirmed cases with the SARS-CoV-2 were identified as a positive result of a high-throughput sequencing or an RT-PCR assay for respiratory specimens including nasal and pharyngeal swab specimens, bronchoalveolar lavage fluid, sputum, or bronchial aspirates or a positive result of anti-SARS-CoV-2 IgM/IgG or the titer of anti-SARS-CoV-2 IgG antibody in the recovery period was four times or more higher than in the acute period [12]. At present, the diagnosis of the COVID-19 is mainly based on clinical characteristics, epidemiological history, chest imaging, and laboratory detection.
Clinical characteristics and epidemiological history
The most common symptoms of confirmed patients were fever, cough, and myalgia or fatigue, whereas sputum production, headache, diarrhea, and vomiting were rare [23,23,24,26]. Mild cases only have a low fever and mild fatigue, without pneumonia. Severe and moderate cases had clinical manifestations of dyspnea, lymphopenia, and hypoalbuminemia, which mainly occurred in elderly patients [23]. It is worth noting that patients with severe or critical illness may have a moderate or low fever, or even no significant fever [12]. The elderly and those with chronic diseases, including diabetes, hypertension, and cardiovascular disease, have poor prognoses [12]. Most severe patient died of severe pneumonia, severe respiratory failure, and multiple organ failure [26]. Epidemiological investigations indicate that most patients were local residents of Wuhan or had direct exposure to the Huanan Seafood Market, a travel history to Wuhan, or contact with confirmed cases [7]. In addition, outbreaks within family clusters have been reported from several provinces in China [27]. An increasing number of cluster cases including family cluster cases are occurring [24, 25].
Chest imaging
The most common patterns seen on chest CT were bilateral, peripheral, and ground-glass opacity [28, 29]. Less common CT findings were nodules, cystic changes, bronchiolectasis, pleural effusion, and lymphadenopathy [28, 29]. Chest CT images of an early-stage COVID-19 patients showed multiple small plaques and interstitial changes. The findings of a progressive stage chest CT images included a bilateral multiple ground-glass opacity and an infiltrating opacity with consolidation, interstitial thickening or fibrous stripes [29,29,31]. The diffuse lesions in bilateral lungs could be seen in the most seriously affected patients, whose CT showed as “white lungs” [31].
Laboratory detection
Specific laboratory detection
Isolation of the causal agent and determination of its partial genome sequence provided the basis for next-generation sequencing or real-time reverse transcriptase-polymerase chain reaction (RT-PCR) methods for the SARS-CoV-2 [14, 17]. After the SARS-CoV-2 was isolated from a lower respiratory tract specimen, a diagnostic RT-PCR test was developed. RT-PCR tests were based on the RNA-dependent RNA polymerase (RdRp) gene of the ORF1ab sequence, E gene, N gene, and S gene of the SARS-CoV-2 genome [32,32,33,35]. Among these assays, RT-PCR assays targeting the RdRp assay had the highest analytical sensitivity [32]. The SARS-CoV-2 nucleic acid can be detected in nasal and pharyngeal swab specimens, bronchoalveolar lavage fluid, sputum, bronchial aspirates, blood, anal swab, and other samples by an RT-PCR [36, 37]. In a case with severe peptic ulcers after the onset of symptoms, the SARS-CoV-2 was directly detected in the esophageal erosion and at the bleeding site [7]. Some patients infected with the SARS-CoV-2 also displayed gastrointestinal symptoms such as diarrhea [23, 38] because some viruses may enter the digestive tract through the throat, infecting the intestinal epithelial cells and activating the intestinal immune response. Thus, the SARS-CoV-2 nucleic acid can also be detected in the stool samples of some patients [7, 36, 37]. High-throughput sequencing or an RT-PCR assay has become a standard and formative assessment for the diagnosis of the COVID-19 [12]. However, nucleic acid amplification kits sometimes produced false-negative results among patients whose clinical features, chest imaging, and laboratory detection accorded with the COVID-19 [30, 39]. There are several possible reasons for the false-negative results from the nucleic acid kit. Firstly, although older age was correlated with higher viral load [40], it is not clear whether the viral load in body fluids has a positive linear correlation with the severity of symptoms after infection. If the virus in the suspected patients remains to be rapidly replicated and released in the lungs, the nasal and pharyngeal swabs sampling may not collect enough virus for diagnosis. Secondly, the current common sampling method is to collect nasal and pharyngeal swabs, sputum, or the alveolar lavage fluid [36, 40, 41]. Few patients with the SARS-CoV-2 infection had prominent signs and symptoms of the respiratory tract, indicating that the target cells may be located in the lower airway [18]. The viral nucleic acid is most easily detected in the alveolar lavage fluid, followed by sputum, nasal, and pharyngeal swabs [41,41,43]. A study of 4880 cases showed that the alveolar lavage fluid exhibited the most highest positive rate of 100% for SARS-CoV-2 ORF1ab gene; the sputum exhibited a 49.12% positive rate, and the nasal and pharyngeal swabs samples showed a poor positive rate of 38.25% [41]. Alveolar lavage fluid collection is generally suitable for patients with a severe or critical illness, not mild cases. Sputum specimens are also more difficult to obtain because few patients with the SARS-CoV-2 infection had sputum production [7, 18]. Due to the limitations associated with operations and patient acceptance, the most common sampling method in clinical practice is nasal and pharyngeal swab collection. However, respiratory samples collected from 80 individuals at different stages of infection showed a median of 7·99 × 104 in nasal and pharyngeal swab samples and 7·52 × 105 in sputum samples [36]. Sputum samples generally showed higher viral loads than throat swab samples [36, 43]. The low viral load in nasal and pharyngeal swab makes the diagnosis of the SARS-CoV-2 more difficult. On the other hand, RT-PCR test results of pharyngeal swab specimens were variable and potentially unstable [44]. It was reported that patients with initial non-positive results were eventually confirmed with COVID-19 by 3~5 repeated swab PCR tests [44]. The phenomenon of SARS-CoV-2 positive in the stool samples but negative nucleic acid in throat swab specimens indicated that selecting fecal samples for a nucleic acid test may be an alternative strategy [45]. Considering that the SARS-CoV-2 nucleic acid can be detected in nasal and pharyngeal swab specimens, bronchoalveolar lavage fluid, sputum, bronchial aspirates, blood, and anal swab [36, 37], it is suggested to collect samples from multiple site of the same patient at different stages and combine them for detection to improve the positive rate. Thirdly, the SARS-CoV-2 is an RNA virus with low stability, which is easily degraded by RNA enzymes released after exogenous or cellular destruction, affecting the final detection efficiency. Improper sampling location, insufficient sampling strength, and irregular sample delivery process account for the false-negative results of the nucleic acid kit test [39]. Besides, in order to improve the sensitivity of detection, most manufacturers choose two or more regions of viral nucleic acid sequence for detection, including the ORF1ab sequence, E gene, N gene, and S gene of the SARS-CoV-2 genome [32,32,33,35]. In actual tests, there is a certain proportion of positive results of a single target gene locus indicating that the sensitivity of the reagent to different gene regions is indeed different [41], which may also be caused by the competition between the loci of two or three target genes. Furthermore, reagent reaction conditions, reaction system, and nucleic acid addition amount may affect the sensitivity of detection and analysis [46]. It is an effective measure for the clinical laboratory to carry out quality control for each batch of reagents by using the confirmed negative and positive samples before routine work.
Based on the above reasons, detection of the viral RNA using RT-PCR can only achieve a sensitivity of 30~60% [41, 47, 48], depending on the course and condition of the patient, the type and number of clinical specimens collected, and the protocol used. The older had higher positive rate than the young [41] which may be explained by the finding that the older was correlated with higher viral load [40]. Supplement serum IgM/IgG antibody detection against the SARS-CoV-2 internal nucleoprotein (NP) and surface spike protein receptor-binding domain (RBD) can make up for the shortcomings of RT-PCR in some cases [40, 49]. The antibody is the product of a humoral immune response after infection with the virus. Generally, IgM antibodies rise within a few days after a viral infection and can be detected as soon as a week of incubation, and IgG antibodies appear in the middle and late stages of the infection. There is a process of a continuous increase in the antibody titer, and it remains in the blood circulation for a long time. At the moment, the most widely used methods for serodiagnosis of the SARS-CoV-2 infection in clinical microbiology laboratories are antibody detection in acute- and convalescent-phase sera by colloidal gold immunochromatography and enzyme-linked immunosorbent assay (ELISA) [40]. In short, a test for IgM/IgG antibodies can also determine whether a patient has been infected with the SARS-CoV-2 recently or previously and act as a supplementary detection to identify patients with high clinical suspicion of the SARS-CoV-2 infection but negative RT-PCR findings [40, 49]. The new serological diagnostic kits for IgM and IgG antibodies for SARS-CoV-2 have the advantages of high sensitivity and early diagnosis. In addition, the operational requirements of antibody detection in clinical microbiology laboratories are relatively low, fast, capable of large quantities, and can be completed in basic laboratories compared with the nucleic acid test. Anti-SARS-CoV-2 IgM antibody was positive at 3~5 days after onset, and the titer of anti-SARS-CoV-2 IgG antibody in the recovery period was four times or more higher than in the acute period [12]. Although the supplementary antibody test can make up for the missed diagnosis of RT-PCR, it still cannot diagnose all infected patients. The detection of IgM and IgG antibodies can only achieve a sensitivity of 70% at 4~6 days after admission for COVID-19 patients (unpublished data from our group). The detection of IgM and IgG antibodies may be futile for the elderly, because of hypoimmunity and a weak antibody production capacity.
Nonspecific laboratory detection
The laboratory examination of patients at an early stage showed leucopenia, lymphopenia, high level of aspartate aminotransferase, C-reactive protein (CRP), and erythrocyte sedimentation rate [18]. Most patients had normal serum levels of procalcitonin. Compared with moderate cases, severe cases more frequently had lymphopenia, with higher levels of alanine aminotransferase, lactate dehydrogenase, C-reactive protein, ferritin, and D-dimer as well as markedly higher levels of IL-2R, IL-6, IL-10, and TNF-α [23]. Typical abnormal laboratory findings in pediatric patients were elevated creatine kinase MB, decreased lymphocytes, leucopenia, and elevated procalcitonin [24]. Recent studies have also shown another potential diagnostic biomarker for the SARS-CoV-2 diagnosis. Renin cleaves liver-derived angiotensinogen (AGT) into angiotensin I, which is then further processed by the angiotensin-converting enzyme (ACE) into the octapeptide angiotensin II. The abnormal increase of angiotensin II has been reported to be associated with hypertension, heart failure, and lung and kidney dysfunction as well as several pathophysiological features, including inflammation, metabolic dysfunction, and aging [50, 51]. Xu et al. performed structural modeling of the S-protein of the SARS-CoV-2 to evaluate its ability to interact with human angiotensin-converting enzyme 2 (ACE2) molecules. Because of the loss of hydrogen bond interactions due to replacing Arg426 with Asn426 in the SARS-CoV-2 S-protein, the binding free energy for the SARS-CoV-2 S-protein increased by 28 kcal mol–1 when compared with the SARS-CoV S-protein binding. The results revealed that the SARS-CoV-2 S-protein has a strong binding affinity to human ACE2 [11]. A study discovered the markedly increased level of angiotensin II in the plasma samples from SARS-CoV-2-infected patients was linearly correlated with viral load and lung injury [52]. It is suggested that the imbalance of the renin-angiotensin-aldosterone system is caused by the SARS-CoV-2, and angiotensin receptor blocker (ARB) drugs may be used as a potential repurposing treatment of the SARS-CoV-2 infection. Similar studies have demonstrated that the SARS-CoV could bind to its receptor ACE2, downregulating its expressions, resulting in increased angiotensin II levels in mouse blood samples, signaling through angiotensin II receptor 1, leading to an acute lung injury [53]. Besides, markedly, elevation of angiotensin II level in the H7N9-infected patients was associated with the disease severity and outcomes [54].
Discussion
Chest CT imaging showed that 76.4% of infected patients manifested as pneumonia on admission, which was mainly ground-glass opacity (50%) and bilateral patchy shadowing (46.4%). The majority of severe patients could be diagnosed by chest X-ray and chest CT imaging. Despite these predominant manifestations, it was reported that 221 out of the 926 (23.87%) in severe cases compared with 9 out of the 173 non-severe cases (5.20%) who had no abnormal radiological findings were diagnosed by symptoms plus RT-PCR positive findings, suggesting that not all patients had abnormal chest radiological findings of pneumonia. Chest CT images of the early-stage COVID-19 patients showed unilateral or bilateral ground-glass opacity, which was similar to some non-COVID-19 images of patients with the respiratory syncytial viral (RSV), mycoplasma, and parainfluenza virus, suggesting that chest CT scans cannot the identify COVID-19 patients and the non-COVID-19 patients in some cases. Co-infection with other viruses such as influenza A/B, rhino/enterovirus, respiratory syncytial virus, other atypical pathogens, fungi, and bacteria has been reported in the COVID-19 patients [49, 55]. Mixed infection among COVID-19 patients makes the diagnosis of chest CT images more difficult. Besides, positive respiratory pathogen results cannot serve as evidence for the exclusion of SARS-COV-2 infection. Methods of pathogen-specific detection are mainly divided into four types, including virus culture, nucleic acid detection, antigen detection, and antibody detection. In terms of virus culture, the cultivation of the SARS-CoV-2 requires biosafety level 3 laboratory facilities, which are not available in most clinical microbiology laboratories. Thus, the cultivation of the SARS-CoV-2 is mainly used for scientific research. Commercial antigen detection kits require the preparation of monoclonal antibodies and polyclonal antibodies, whereas it costs a long time from production to extraction during antibody preparation, and the preparation process is complicated. Detection of the viral nucleic acid using an RT-PCR assay has become a standard and formative assessment for the diagnosis of COVID-19. However, detection of viral RNA using RT-PCR can only achieve a sensitivity of 30~60%, depending on the course and condition of the patient, the type and number of clinical specimens collected, and the protocol used. In order to improve the positive rate of detection, it is suggested to collect multiple site samples of the same patient at different stages repeatedly and combine them for detection. The phenomenon of SARS-CoV-2 positive in the stool samples but negative nucleic acid in throat swab specimens should be taken seriously. Patients with early or mild illness may have a low viral load in nasal and pharyngeal swabs, resulting in false-negative nucleic acid tests. Thus, selecting fecal samples for a nucleic acid test may be an alternative strategy, regardless of the presence or absence of gastrointestinal symptoms such as diarrhea. In addition, a fecal-oral transmission might exist in the transmission of 2019-nCoV; thus, the transmission via gastrointestinal secretions should be fully considered to control the rapid spread worldwide. Whole genome sequencing (WGS) method can overcome the mutation problems which cause false-negative results in RT-PCR [55, 56], whereas it is not applicable to clinical practice considering the economic status of patients. For individuals with high clinical suspicion of the SARS-CoV-2 infection but negative RT-PCR findings, the detection of IgM/IgG antibodies should be considered. We recommend IgM antibody testing 1 week after infection and IgG antibody testing 4 weeks after infection. Although the supplementary antibody test can make up for the missed diagnosis of RT-PCR, it cannot diagnose all the infected patients. Collectively, for chest CT scans, RT-PCR assays, and the detection of IgM/IgG antibodies, multiple and repetitive tests should be considered during different stages of the COVID-19. Further research of SARS-CoV-2 and the development of more sensitive detection methods will facilitate the diagnosis of COVID-19. In addition, the development of broad-spectrum antiviral drugs and vaccines will enhance the ability to manage future outbreaks caused by this cluster of viruses.
References
Weiss SR, Leibowitz JL (2011) Coronavirus pathogenesis. Adv Virus Res 81:85–164
Li F (2016) Structure, function, and evolution of coronavirus spike proteins. Annu Rev Virol 3:237–261. https://doi.org/10.1146/annurev-virology-110615-042301
Tang Q, Song Y, Shi M, Cheng Y, Zhang W, Xia XQ (2015) Inferring the hosts of coronavirus using dual statistical models based on nucleotide composition. Sci Rep 5:17155. https://doi.org/10.1038/srep17155
Su S, Wong G, Shi W et al (2016) Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol 24:490–502. https://doi.org/10.1016/j.tim.2016.03.003
Zhu N, Zhang D, Wang W et al (2020) A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med 382:727–733. https://doi.org/10.1056/NEJMoa2001017
Wuhan Municipal Health Commission. Report of clustering pneumonia of unknown etiology in Wuhan City. http://wjw.wuhan.gov.cn/front/web/showDetail/2019123108989. Accessed 31 Dec 2019
Guan WJ, Ni ZY, Hu Y et al (2020) Clinical characteristics of 2019 novel coronavirus infection in China. Medrxiv [Preprint]. [cited 2020 Feb 29]. https://www.medrxiv.org/content/10.1101/2020.02.06.20020974v1
WHO. Novel coronavirus (2019-nCoV) situation reports. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports/. Accessed 20 Jan 2020
Coronavirus Study Group (2020) Severe acute respiratory syndrome-related coronavirus: the species and its viruses – a statement of the Coronavirus Study Group. Biorxiv [Preprint]. [cited 2020 Fab 29]. https://www.biorxiv.org/content/10.1101/2020.02.07.937862v1.full
National Health Commission Update on February 27, 2020. http://www.nhc.gov.cn/xcs/yqtb/202002/d5e15557ee534fcbb5aaa9301ea5235f.shtml. Accessed 27 Feb 2020
Xu X, Chen P, Wang J et al (2020) Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Sci China Life Sci 63:457–460. https://doi.org/10.1007/s11427-020-1637-5
National Health Commission of the People’s Republic of China. New coronavirus pneumonia prevention and control program (trial version 7th ed). http://www.nhc.gov.cn/yzygj/s7653p/202003/46c9294a7dfe4cef80dc7f5912eb1989.shtml Accessed 4 Apr 2020
Song Z, Xu Y, Bao L et al (2019) From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses 11. https://doi.org/10.3390/v11010059
Wu A, Peng Y, Huang B et al (2020) Genome composition and divergence of the novel coronavirus (2019-nCoV) originating in China. Cell Host Microbe. https://doi.org/10.1016/j.chom.2020.02.001
Zhu Z, Zhang Z, Chen W et al (2018) Predicting the receptor-binding domain usage of the coronavirus based on kmer frequency on spike protein. Infect Genet Evol 61:183–184. https://doi.org/10.1016/j.meegid.2018.03.028
He Y, Zhou Y, Liu S et al (2004) Receptor-binding domain of SARS-CoV spike protein induces highly potent neutralizing antibodies: implication for developing subunit vaccine. Biochem Biophys Res Commun 324:773–781. https://doi.org/10.1016/j.bbrc.2004.09.106
Lu R, Zhao X, Li J et al (2020) Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 395:565–574. https://doi.org/10.1016/S0140-6736(20)30251-8
Huang C, Wang Y, Li X et al (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395:497–506. https://doi.org/10.1016/S0140-6736(20)30183-5
Cui J, Li F, Shi ZL (2019) Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol 17:181–192. https://doi.org/10.1038/s41579-018-0118-9
Guan Y, Zheng BJ, He YQ et al (2003) Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science. 302:276–278. https://doi.org/10.1126/science.1087139
Drosten C, Kellam P, Memish ZA (2014) Evidence for camel-to-human transmission of MERS coronavirus. N Engl J Med 371:1359–1360. https://doi.org/10.1126/science.1087139
Lam TT, Shum MH, Zhu HC et al (2020) Identifying SARS-CoV-2 related coronaviruses in Malayan pangolins. Nature. https://doi.org/10.1038/s41586-020-2169-0
Chen G, Wu D, Guo W et al (2020) Clinical and immunologic features in severe and moderate coronavirus disease 2019. J Clin Invest. https://doi.org/10.1172/JCI137244
Qiu H, Wu J, Hong L et al (2020) Clinical and epidemiological features of 36 children with coronavirus disease 2019 (COVID-19) in Zhejiang, China: an observational cohort study. Lancet Infect Dis. https://doi.org/10.1016/S1473-3099(20)30198-5
Tian S, Hu N, Lou J et al (2020) Characteristics of COVID-19 infection in Beijing. J Inf Secur 80:401–406. https://doi.org/10.1016/j.jinf.2020.02.018
Chen N, Zhou M, Dong X et al (2020) Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 395:507–513. https://doi.org/10.1016/S0140-6736(20)30211-7
Chan JF, Yuan S, Kok KH et al (2020) A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 395:514–523. https://doi.org/10.1016/S0140-6736(20)30154-9
Wong HYF, Lam HYS, Fong AH et al (2019) Frequency and distribution of chest radiographic findings in COVID-19 positive patients. Radiology 201160. https://doi.org/10.1148/radiol.2020201160
Shi H, Han X, Jiang N et al (2020) Radiological findings from 81 patients with COVID-19 pneumonia in Wuhan, China: a descriptive study. Lancet Infect Dis 20:425–434. https://doi.org/10.1016/S1473-3099(20)30086-4
Cheng Z, Lu Y, Cao Q et al Clinical Features and Chest CT Manifestations of Coronavirus Disease 2019 (COVID-19) in a single-center study in Shanghai, China. AJR Am J Roentgenol 2020:1–6. https://doi.org/10.2214/AJR.20.22959
Xiong Y, Sun D, Liu Y et al (2020) Clinical and high-resolution CT Features of the COVID-19 infection: comparison of the initial and follow-up changes. Investig Radiol. https://doi.org/10.1097/RLI.0000000000000674
Corman VM, Landt O, Kaiser M et al (2020) Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill 25. https://doi.org/10.2807/1560-7917.ES.2020.25.3.2000045
Konrad R, Eberle U, Dangel A et al (2020) Rapid establishment of laboratory diagnostics for the novel coronavirus SARS-CoV-2 in Bavaria, Germany, February 2020. Euro Surveill 25. https://doi.org/10.2807/1560-7917
Chan JF, Yip CC, To KK et al (2020) Improved molecular diagnosis of COVID-19 by the novel, highly sensitive and specific COVID-19-RdRp/Hel real-time reverse transcription-polymerase chain reaction assay validated in vitro and with clinical specimens. J Clin Microbiol. https://doi.org/10.1128/JCM.00310-20
Reusken CBEM, Broberg EK, Haagmans B et al (2020) Laboratory readiness and response for novel coronavirus (2019-nCoV) in expert laboratories in 30 EU/EEA countries, January 2020. Euro Surveill 25. https://doi.org/10.2807/1560-7917.ES.2020.25.6.2000082
Pan Y, Zhang D, Yang P et al (2020) Viral load of SARS-CoV-2 in clinical samples. Lancet Infect Dis 20:411–412. https://doi.org/10.1016/S1473-3099(20)30113-4
Chen W, Lan Y, Yuan X et al (2020) Detectable 2019-nCoV viral RNA in blood is a strong indicator for the further clinical severity. Emerg Microbes Infect 9:469–473. https://doi.org/10.1080/22221751.2020.1732837
Yu N, Li W, Kang Q et al (2020) Clinical features and obstetric and neonatal outcomes of pregnant patients with COVID-19 in Wuhan, China: a retrospective, single-centre, descriptive study. Lancet Infect Dis. https://doi.org/10.1016/S1473-3099(20)30176-6
Xie X, Zhong Z, Zhao W, Zheng C, Wang F, Liu J (2020) Chest CT for typical 2019-nCoV pneumonia: relationship to negative RT-PCR testing. Radiology. https://doi.org/10.1148/radiol.2020200343
To KK, Tsang OT, Leung WS et al (2020) Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis. https://doi.org/10.1016/S1473-3099(20)30196-1
Liu R, Han H, Liu F et al (2020) Positive rate of RT-PCR detection of SARS-CoV-2 infection in 4880 cases from one hospital in Wuhan, China, from Jan to Feb 2020. Clin Chim Acta 505:172–175. https://doi.org/10.1016/j.cca.2020.03.009
Yang Y, Yang MH, Shen CG et al Evaluating the accuracy of different respiratory specimens in the laboratory diagnosis and monitoring the viral shedding of 2019-nCoVinfections. https://www.medrxiv.org/content/10.1101/2020.02.11.20021493v2. Accessed 17 Feb 2020
Yu F, Yan L, Wang N et al (2020) Quantitative detection and viral load analysis of SARS-CoV-2 in infected patients. Clin Infect Dis. https://doi.org/10.1093/cid/ciaa345
Li Y, Yao L, Li J et al (2020) Stability issues of RT-PCR testing of SARS-CoV-2 for hospitalized patients clinically diagnosed with COVID-19. J Med Virol. https://doi.org/10.1002/jmv.25786
Zhang T, Cui X, Zhao X et al (2020) Detectable SARS-CoV-2 viral RNA in feces of three children during recovery period of COVID-19 pneumonia. J Med Virol. https://doi.org/10.1002/jmv.25795
Lippi G, Simundic AM, Plebani M (2020) Potential preanalytical and analytical vulnerabilities in the laboratory diagnosis of coronavirus disease 2019 (COVID-19). Clin Chem Lab Med. https://doi.org/10.1515/cclm-2020-0285
Ai T, Yang Z, Hou H et al (2020) Correlation of chest CT and RT-PCR testing in coronavirus disease 2019 (COVID-19) in China: a report of 1014 cases. Radiology. 200642. https://doi.org/10.1148/radiol.2020200642
Xie C, Jiang L, Huang G et al (2020) Comparison of different samples for 2019 novel coronavirus detection by nucleic acid amplification tests. Int J Infect Dis 93:264–267. https://doi.org/10.1016/j.ijid.2020.02.050
Dong X, Cao YY, Lu XX et al (2020) Eleven faces of coronavirus disease 2019. Allergy. https://doi.org/10.1111/all.14289
Damman K, Gori M, Claggett B et al (2018) Renal effects and associated outcomes during angiotensin-Neprilysin inhibition in heart failure. JACC Heart Fail 6:489–498. https://doi.org/10.1016/j.jchf.2018.02.004
Fröhlich H, Nelges C, Täger T et al (2016) Long-term changes of renal function in relation to ace inhibitor/angiotensin receptor blocker dosing in patients with heart failure and chronic kidney disease. Am Heart J 178:28–36. https://doi.org/10.1016/j.ahj.2016.03.024
Liu Y, Yang Y, Zhang C et al (2020) Clinical and biochemical biomarkerses from SARS-CoV-2 infected patients linked to viral loads and lung injury. Sci China Life Sci. https://doi.org/10.1007/s11427-020-1643-8
Imai Y, Kuba K, Penninger JM (2008) Lessons from SARS: a new potential therapy for acute respiratory distress syndrome (ARDS) with angiotensin converting enzyme 2 (ACE2). Masui. 57:302–310
Huang F, Guo J, Zou Z et al (2014) Angiotensin II plasma levels are linked to disease severity and predict fatal outcomes in H7N9-infected patients. Nat Commun 5:3595
Jing-Wen Ai, Hao-Cheng Zhang, Teng Xu et al (2020) Optimizing diagnostic strategy for novel coronavirus pneumonia, a multi-center study in Eastern China. Medrxiv [Preprint]. [cited 2020 Mar 29]. https://www.medrxiv.org/content/10.1101/2020.02.13.20022673v1
Gong YN, Yang SL, Chen GW et al (2017) A metagenomics study for the identification of respiratory viruses in mixed clinical specimens: an application of the iterative mapping approach. Arch Virol 162:2003–2012. https://doi.org/10.1007/s00705-017-3367-4
Funding
This study was supported by the Natural Science Foundation of China (grant nos. 81571572, 81201488, and 30801088).
Author information
Authors and Affiliations
Contributions
HW and XL were major contributors to writing the manuscript. HW, XL, SZ, LW, JL, XW, TL, YX, and WW checked and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, H., Li, X., Li, T. et al. The genetic sequence, origin, and diagnosis of SARS-CoV-2. Eur J Clin Microbiol Infect Dis 39, 1629–1635 (2020). https://doi.org/10.1007/s10096-020-03899-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-020-03899-4