Abstract
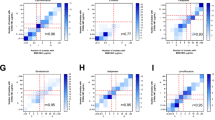
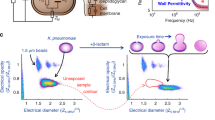
We present the MilliDrop Analyzer (MDA), a droplet-based millifluidic system for digital antimicrobial susceptibility testing (D-AST), which enables us to determine minimum inhibitory concentrations (MICs) precisely and accurately. The MilliDrop technology was validated by using resazurin for fluorescence readout, for comparison with standard methodology, and for conducting reproducibility studies. In this first assessment, the susceptibility of a reference Gram-negative strain Escherichia coli ATCC 25922 to gentamicin, chloramphenicol, and nalidixic acid were tested by the MDA, VITEK®2, and broth microdilution as a reference standard. We measured the susceptibility of clinically relevant Gram-positive strains of Staphylococcus aureus to vancomycin, including vancomycin-intermediate S. aureus (VISA), heterogeneous vancomycin-intermediate S. aureus (hVISA), and vancomycin-susceptible S. aureus (VSSA) strains. The MDA provided results which were much more accurate than those of VITEK®2 and standard broth microdilution. The enhanced accuracy enabled us to reliably discriminate between VSSA and hVISA strains.


Similar content being viewed by others
References
Zillich AJ, Sutherland JM, Wilson SJ, Diekema DJ, Ernst EJ, Vaughn TE, Doebbeling BN (2006) Antimicrobial use control measures to prevent and control antimicrobial resistance in US hospitals. Infect Control Hosp Epidemiol 27:1088–1095
World Health Organization (WHO) (2014) Antimicrobial resistance: global report on surveillance 2014. Available online at: http://www.who.int/drugresistance/documents/surveillancereport/en
Leekha S, Terrell CL, Edson RS (2011) General principles of antimicrobial therapy. Mayo Clin Proc 86:156–167
Wheat PF (2001) History and development of antimicrobial susceptibility testing methodology. J Antimicrob Chemother 48:1–4
Pulido MR, García-Quintanilla M, Martín-Peña R, Cisneros JM, McConnell MJ (2013) Progress on the development of rapid methods for antimicrobial susceptibility testing. J Antimicrob Chemother 68:2710–2717
van Belkum A, Durand G, Peyret M, Chatellier S, Zambardi G, Schrenzel J, Shortridge D, Engelhardt A, Dunne WM Jr (2013) Rapid clinical bacteriology and its future impact. Ann Lab Med 33:14–27
van Belkum A, Dunne WM Jr (2013) Next-generation antimicrobial susceptibility testing. J Clin Microbiol 51:2018–2024
Mittman SA, Huard RC, Della-Latta P, Whittier S (2009) Comparison of BD phoenix to Vitek 2, MicroScan MICroSTREP, and Etest for antimicrobial susceptibility testing of Streptococcus pneumoniae. J Clin Microbiol 47:3557–3561
Clinical and Laboratory Standards Institute (CLSI) (2013) Performance standards for antimicrobial susceptibility testing; Twenty-third informational supplement. CLSI document M100-S23. CLSI, Pennsylvania
Conville PS, Brown-Elliott BA, Wallace RJ Jr, Witebsky FG, Koziol D, Hall GS, Killian SB, Knapp CC, Warshauer D, Van T, Wengenack NL, Deml S, Woods GL (2012) Multisite reproducibility of the broth microdilution method for susceptibility testing of nocardia species. J Clin Microbiol 50:1270–1280
Howden BP, Davies JK, Johnson PDR, Stinear TP, Grayson ML (2010) Reduced vancomycin susceptibility in Staphylococcus aureus, including vancomycin-intermediate and heterogeneous vancomycin-intermediate strains: resistance mechanisms, laboratory detection, and clinical implications. Clin Microbiol Rev 23:99–139
Lowy FD (2003) Antimicrobial resistance: the example of Staphylococcus aureus. J Clin Invest 111:1265–1273
Brouzes E, Medkova M, Savenelli N, Marran D, Twardowski M, Hutchison JB, Rothberg JM, Link DR, Perrimon N, Samuels ML (2009) Droplet microfluidic technology for single-cell high-throughput screening. Proc Natl Acad Sci U S A 106:14195–14200
Fallah-Araghi A, Baret JC, Ryckelynck M, Griffiths AD (2012) A completely in vitro ultrahigh-throughput droplet-based microfluidic screening system for protein engineering and directed evolution. Lab Chip 12:882–891
Guo MT, Rotem A, Heyman JA, Weitz DA (2012) Droplet microfluidics for high-throughput biological assays. Lab Chip 12:2146–2155
Kelly BT, Baret JC, Taly V, Griffiths AD (2007) Miniaturizing chemistry and biology in microdroplets. Chem Commun (Camb) 14:1773–1788
Theberge AB, Courtois F, Schaerli Y, Fischlechner M, Abell C, Hollfelder F, Huck WTS (2010) Microdroplets in microfluidics: an evolving platform for discoveries in chemistry and biology. Angew Chem Int Ed Engl 49:5846–5868
Abate AR, Hung T, Sperling RA, Mary P, Rotem A, Agresti JJ, Weiner MA, Weitz DA (2013) DNA sequence analysis with droplet-based microfluidics. Lab Chip 13:4864–4869
Boitard L, Cottinet D, Kleinschmitt C, Bremond N, Baudry J, Yvert G, Bibette J (2012) Monitoring single-cell bioenergetics via the coarsening of emulsion droplets. Proc Natl Acad Sci U S A 109:7181–7186
Clausell-Tormos J, Lieber D, Baret JC, El-Harrak A, Miller OJ, Frenz L, Blouwolff J, Humphry KJ, Köster S, Duan H, Holtze C, Weitz DA, Griffiths AD, Merten CA (2008) Droplet-based microfluidic platforms for the encapsulation and screening of mammalian cells and multicellular organisms. Chem Biol 15:427–437
Kintses B, Hein C, Mohamed MF, Fischlechner M, Courtois F, Lainé C, Hollfelder F (2012) Picoliter cell lysate assays in microfluidic droplet compartments for directed enzyme evolution. Chem Biol 19:1001–1009
Pekin D, Skhiri Y, Baret JC, Le Corre D, Mazutis L, Ben Salem C, Millot F, El Harrak A, Hutchison JB, Larson JW, Link DR, Laurent-Puig P, Griffiths AD, Taly V (2011) Quantitative and sensitive detection of rare mutations using droplet-based microfluidics. Lab Chip 11:2156–2166
Lagus TP, Edd JF (2013) A review of the theory, methods and recent applications of high-throughput single-cell droplet microfluidics. J Phys D Appl Phys 46:114005
Baraban L, Bertholle F, Salverda MLM, Bremond N, Panizza P, Baudry J, de Visser JAGM, Bibette J (2011) Millifluidic droplet analyser for microbiology. Lab Chip 11:4057–4062
Churski K, Kaminski TS, Jakiela S, Kamysz W, Baranska-Rybak W, Weibel DB, Garstecki P (2012) Rapid screening of antibiotic toxicity in an automated microdroplet system. Lab Chip 12:1629–1637
Martin K, Henkel T, Baier V, Grodrian A, Schön T, Roth M, Köhler JM, Metze J (2003) Generation of larger numbers of separated microbial populations by cultivation in segmented-flow microdevices. Lab Chip 3:202–207
Grodrian A, Metze J, Henkel T, Martin K, Roth M, Köhler JM (2004) Segmented flow generation by chip reactors for highly parallelized cell cultivation. Biosens Bioelectron 19(11):1421–1428
Funfak A, Hartung R, Cao J, Martin K, Wiesmüller KH, Wolfbeis OS, Köhler JM (2009) Highly resolved dose–response functions for drug-modulated bacteria cultivation obtained by fluorometric and photometric flow-through sensing in microsegmented flow. Sensors Actuators B Chem 142(1):66–72
Boedicker JQ, Li L, Kline TR, Ismagilov RF (2008) Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 8:1265–1272
Zwietering MH, Jongenburger I, Rombouts FM, van ’t Riet K (1990) Modeling of the bacterial growth curve. Appl Environ Microbiol 56:1875–1881
Kahm M, Hasenbrink G, Lichtenberg-Fraté H, Ludwig J, Kschischo M (2010) grofit: fitting biological growth curves with R. J Stat Softw 33:1–21
Jacobs MR (2001) Optimisation of antimicrobial therapy using pharmacokinetic and pharmacodynamic parameters. Clin Microbiol Infect 7:589–596
European Committee on Antimicrobial Susceptibility Testing (EUCAST) (2014) Breakpoint tables for interpretation of MICs and zone diameters. Version 4.0, 2014. Available online at: http://www.eucast.org/clinical_breakpoints/
Eng RH, Padberg FT, Smith SM, Tan EN, Cherubin CE (1991) Bactericidal effects of antibiotics on slowly growing and nongrowing bacteria. Antimicrob Agents Chemother 35:1824–1828
MacKenzie FM, Gould IM (1993) The post-antibiotic effect. J Antimicrob Chemother 32:519–537
Acknowledgments
This work is supported by a public grant overseen by the French National Research Agency (ANR) as part of the “Investissements d’Avenir” program (reference: ANR-10-NANB-0002-06). bioMérieux provided materials for the study and a post-doctoral grant. The authors are thankful to Alex van Belkum for the stimulating discussions and proofreading.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Patrick Broyer, Anne-Coline Chareire, Pierrot Bourne-Branchu, Pierre Mahé, Maud Tournoud, Christine Franceschi, and Gilles Zambardi are scientists employed by bioMérieux. Laurent Boitard, Jean Baudry, and Jérôme Bibette are founders of MilliDrop Instruments SAS. Lianmei Jiang declares no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1925 kb)
Rights and permissions
About this article
Cite this article
Jiang, L., Boitard, L., Broyer, P. et al. Digital antimicrobial susceptibility testing using the MilliDrop technology. Eur J Clin Microbiol Infect Dis 35, 415–422 (2016). https://doi.org/10.1007/s10096-015-2554-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-015-2554-z