Abstract
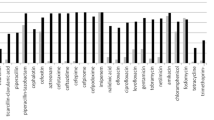
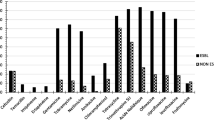
Thanks to the recent advent of matrix-assisted laser desorption/ionisation time-of-flight (MALDI-TOF) technology, Helcococcus kunzii is now easily identifiable and considered as an opportunistic pathogen. However, data about antimicrobial susceptibilities remain very limited. The aim of the study was, then, to assess its in vitro susceptibility to 18 antimicrobial agents and to investigate the genetic basis of macrolide and tetracycline resistance. Thirty-nine human clinical isolates of H. kunzii collected from 2008 to 2013 were studied, as well as the type strain ATCC 51366T. Minimum inhibitory concentrations (MICs) of penicillin G, amoxicillin, cefotaxime, imipenem, gentamicin, erythromycin, clindamycin, quinupristin–dalfopristin, ciprofloxacin, levofloxacin, tetracycline, tigecycline, vancomycin, teicoplanin, linezolid, daptomycin, cotrimoxazole and rifampin were determined by the microdilution method. Screening for macrolide [erm(A) including erm(TR), erm(B), erm(C), erm(F), erm(T), erm(X), msr(A) and mef(A)] and tetracycline [tet(L), tet(M) and tet(O)] resistance genes was performed, as well as the detection of mutations in 23S rRNA. Except for one strain resistant to cefotaxime, all strains were categorised as susceptible to β-lactams, glycopeptides, linezolid, daptomycin and tigecycline. Whereas ciprofloxacin and gentamicin exhibited limited activity, 95 % of strains were categorised as susceptible to levofloxacin. Concerning erythromycin, a bimodal distribution was observed, with 29 ‘wild-type’ strains (MICs from 0.25 to 2 mg/L) and 11 ‘resistant’ strains (MICs ≥256 mg/L), including ten harbouring erm(TR). Two isolates exhibited acquired tetracycline resistance (MICs of 16 mg/L) by the production of tet(M). This large study on the in vitro antimicrobial susceptibility of H. kunzii suggests that β-lactams (especially penicillins) should be preferred for the treatment.

Similar content being viewed by others
References
Caliendo AM, Jordan CD, Ruoff KL (1995) Helcococcus, a new genus of catalase-negative, gram-positive cocci isolated from clinical specimens. J Clin Microbiol 33:1638
Collins MD, Facklam RR, Rodrigues UM et al (1993) Phylogenetic analysis of some Aerococcus-like organisms from clinical sources: description of Helcococcus kunzii gen. nov., sp. nov. Int J Syst Bacteriol 43:425–429
Collins MD, Falsen E, Foster G et al (1999) Helcococcus ovis sp. nov., a gram-positive organism from sheep. Int J Syst Bacteriol 49(Pt 4):1429–1432
Collins MD, Falsen E, Brownlee K et al (2004) Helcococcus sueciensis sp. nov., isolated from a human wound. Int J Syst Evol Microbiol 54:1557–1560
Chow SK, Clarridge JE 3rd (2014) Identification and clinical significance of Helcococcus species, with description of Helcococcus seattlensis sp. nov. from a patient with urosepsis. J Clin Microbiol 52:854–858
Woo PC, Tse H, Wong SS et al (2005) Life-threatening invasive Helcococcus kunzii infections in intravenous-drug users and ermA-mediated erythromycin resistance. J Clin Microbiol 43:6205–6208
McNicholas S, McAdam B, Flynn M et al (2011) The challenges of implantable cardiac device infection due to Helcococcus kunzii. J Hosp Infect 78:337–338
Pérez-Jorge C, Cordero J, Marin M et al (2012) Prosthetic joint infection caused by Helcococcus kunzii. J Clin Microbiol 50:528–530
Sridhar S, Chan JF, Yuen KY (2014) First report of brain abscess caused by a satelliting phenotypic variant of Helcococcus kunzii. J Clin Microbiol 52:370–373
Park JH, Woo BM, Hong SK et al (2014) First Korean case of Helcococcus kunzii bacteremia in a patient with diabetes. Ann Lab Med 34:484–486
Vergne A, Guérin F, Lienhard R et al (2015) Identification and clinical significance of Helcococcus kunzii in human samples. J Clin Microbiol (in press)
Leclercq R (2002) Mechanisms of resistance to macrolides and lincosamides: nature of the resistance elements and their clinical implications. Clin Infect Dis 34:482–492
Roberts MC (2005) Update on acquired tetracycline resistance genes. FEMS Microbiol Lett 245:195–203
Dortet L, Legrand P, Soussy CJ et al (2006) Bacterial identification, clinical significance, and antimicrobial susceptibilities of Acinetobacter ursingii and Acinetobacter schindleri, two frequently misidentified opportunistic pathogens. J Clin Microbiol 44:4471–4478
Clinical and Laboratory Standards Institute (CLSI) (2013) Performance standards for antimicrobial susceptibility testing; Twenty-third informational supplement. CLSI document M100-S23. CLSI, Wayne, PA
Hays C, Lienhard R, Auzou M et al (2014) Erm(X)-mediated resistance to macrolides, lincosamides and streptogramins in Actinobaculum schaalii. J Antimicrob Chemother 69:2056–2060
Aminov RI, Garrigues-Jeanjean N, Mackie RI (2001) Molecular ecology of tetracycline resistance: development and validation of primers for detection of tetracycline resistance genes encoding ribosomal protection proteins. Appl Environ Microbiol 67:22–32
Malhotra-Kumar S, Lammens C, Piessens J et al (2005) Multiplex PCR for simultaneous detection of macrolide and tetracycline resistance determinants in streptococci. Antimicrob Agents Chemother 49:4798–4800
Bilk S, Nordhoff M, Schulze C et al (2011) Antimicrobial susceptibilities and occurrence of resistance genes in bovine Helcococcus ovis isolates. Vet Microbiol 149:488–491
Burdett V (1990) Nucleotide sequence of the tet(M) gene of Tn916. Nucleic Acids Res 18:6137
Acknowledgements
The technical assistance of Michel Auzou is gratefully acknowledged.
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
A. Vergne and F. Guérin contributed equally to this work.
Rights and permissions
About this article
Cite this article
Vergne, A., Guérin, F., Lienhard, R. et al. In vitro antimicrobial susceptibility of Helcococcus kunzii and molecular analysis of macrolide and tetracycline resistance. Eur J Clin Microbiol Infect Dis 34, 2057–2061 (2015). https://doi.org/10.1007/s10096-015-2451-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-015-2451-5