Abstract
We tested the hypothesis that swabs from the nasopharynx carry a higher viral load than swabs from the oropharynx in patients with real-time polymerase chain reaction (PCR)-confirmed influenza infection. Using flocked swabs, oropharyngeal and nasopharyngeal samples were harvested from hospital-admitted influenza patients no later than 3 days after the initial detection of influenza virus. Comparison of cycle threshold (CT) values was performed to assess differences in viral load in the specimens. Seventeen patients were diagnosed with influenza B, 14 patients with influenza A(H1N1)pdm09, and one patient with influenza A(H3N2). Nasopharyngeal samples were positive at a lower CT value than the oropharyngeal samples [mean difference in CT 5.75, 95 % confidence interval (CI) 3.8–7.7, p < 0.01], suggesting that, on average, the calculated viral load of the nasopharyngeal samples was 54 times higher (95 % CI 13.7–210.8) than those of the oropharyngeal samples. The corresponding difference in the calculated viral load for influenza A(H1N1)pdm09 virus was 23 times (95 % CI 3.8–136.2, p < 0.01) and for influenza B virus, it was 80 times (95 % CI 9.3–694.6, p < 0.01). In patients with acute influenza, nasopharyngeal swabbing was clearly superior to oropharyngeal swabbing in terms of diagnostic yield by real-time PCR.
Similar content being viewed by others
Introduction
The early diagnosis of influenza is important for timely antiviral treatment and appropriate infection control. Although a number of sampling strategies such as nasal washes, nasal aspirates, nasal swabbing, nasopharyngeal (NP) and oropharyngeal (OP) swabbing, as well as sputum and bronchial lavage are used, no gold standard is established for the acquisition of representative material for this important diagnosis [1]. Furthermore, interpersonal variability in sampling technique may provide materials of highly variable quality [2].
The infectious potential of influenza viruses is directly related to the extent of viral shedding [3]. Prospective studies on naturally acquired influenza have shown a nonlinear decrease in viral shedding with time. Major comorbidities, systemic corticosteroid use, and late antiviral treatment were consistent with slower viral clearance [4–6]. The duration of influenza virus shedding in healthy volunteers is approximately 4.8 days [3]. Samples harvested later in the course of illness are likely to be negative due to a low viral load [5]. In particular with influenza A(H1N1) pandemic 2009 infections [influenza A(H1N1)pdm09], there are conflicting reports on the possible impact of antiviral treatment on the duration of viral shedding [6, 7]. Prolonged shedding (>7 days) was found in approximately 25 % of all hospitalized patients with influenza A(H1N1) [8]. Younger individuals seem to carry a higher viral load and have a longer period of viral shedding compared to older individuals [8, 9]. Although patients suffering from influenza B are less extensively studied, their viral load and shedding pattern appear to be more variable than those observed in patients infected with influenza A [3].
Viral density and persistence are likely to vary with the location of the pathogen in the airways. Thus, samples from the lower airways may still be positive by polymerase chain reaction (PCR) also when virus is no longer detectable in specimens from the nasopharynx. For this reason, some patients may be contagious even after conversion to negative tests by NP swabs [2]. In a study covering several respiratory viruses, we have shown that the viral load in the nasopharynx is higher than in the oropharynx [10]. Although only a few cases of influenza virus infections were included in that study, we assume that, also in patients suffering from an acute influenza virus infection, swabs from the nasopharynx carry a higher viral load than swabs from the oropharynx. The aim of the present study was to test this hypothesis.
Materials and methods
Ethical committee
The study design was approved by the Norwegian Regional Committee of Research Ethics.
Study design and eligibility criteria
During the influenza seasons of November–December 2009 and December 2010–March 2011, all patients above 18 years of age who had a positive PCR test for influenza virus upon hospital admission were invited to take part in this prospective study. To avoid an increased risk of nasal bleeding, patients with an international normalized ratio (INR) of three or more were not eligible for inclusion in the study. A total of 34 patients were eligible for inclusion, and 32 patients agreed to participate. Two patients declined due to previously experienced discomfort during NP sampling.
Swabbing was performed by one of two technicians experienced with NP and OP swabbing using the previously described technique [10]. Two nylon flocked swabs, 502CS01 regular and 503CS01 nasopharyngeal (both Copan Italia, Brescia, Italy), were used in the oropharynx and nasopharynx, respectively. The swabs were inserted into Universal Transport Medium (UTM) tubes (1.5 ml medium, no beads, Copan Italia) and the tubes were shaken for 10 s.
Examination of the patient’s journal was performed to assess for known risk factors for influenza, such as chronic heart, lung, kidney, liver, or neurological diseases, pregnancy, immunodeficiency, autoimmune diseases, and diabetes. For each participant, the time of any vaccination for influenza A(H1N1)pdm09 virus and/or seasonal influenza (H1N1, H3N2, and B) was established. In addition, the date of symptom debut leading to health care contact and of possible initiation of antiviral treatment was registered.
Sample processing and diagnostic procedures
Following swabbing, specimens were kept at room temperature, and there were no differences in the collection, handling, or analysis for the OP and NP specimens. Nucleic acid extraction usually took place within 24 h and always within 48 h. PCR analyses were performed within the next 24 h (or else the nucleic acid eluates were stored at −70 °C until testing). Using duplex PCR methods, the specimens were examined for untyped influenza A and influenza B (Table 1). All influenza A-positive samples were examined for influenza A(H1N1)pdm09. Further laboratory work followed procedures described previously [10]. The amplification efficiency of the PCR assays used in this study was measured to be above 98 % [10]. Samples that were negative for influenza A(H1N1)pdm09 and positive for untyped influenza A were shipped to the Norwegian Institute of Public Health for further subtyping.
Statistics
Student’s t-tests were used when comparing continuous data, and Chi-square tests when comparing categorical data. The statistical analysis for viral load was based on differences in cycle threshold (CT) values of the cDNA derived from the samples [10]. A CT value of 38 was designated as the cut-off value for positive results. The CT values were analyzed as continuous data. A comparative analysis of differences in CT values in influenza subgroups was performed by the use of one-way analysis of variance (ANOVA). Two-tailed p-values less than 0.05 were considered to be significant. Statistical analysis was performed by the use of PASW Statistics 18.0. Calculations of sample size were performed by the use of Sample Power 2.0. From our previous study, a mean difference in viral load between OP and NP samples was found to be 8.16 [95 % confidence interval (CI) 7.0–10.5] [10]. The sample size needed to detect a mean 4× difference in viral load with a significance level 0.05 and power 80 % was 40 study subjects.
Results
Of the 32 patients included in the study, 17 were diagnosed with influenza B infection, 14 with influenza A(H1N1)pdm09, and one with influenza A (H3N2). Samples from the nasopharynx were positive in 25 of these 32 patients, rendering a diagnostic sensitivity of the procedure of 78 %. In comparison, the sensitivity for OP samples was 63 % (20/32) (χ2 < 0.01). Six patients were PCR-positive in their nasopharynx only and one patient only in material obtained from the oropharynx. Subgroup analysis of influenza A(H1N1)pdm09 showed a diagnostic sensitivity of 71 % (10/14 patients) for NP samples and 50 % (7/14) for OP samples (χ2 = 0.02).
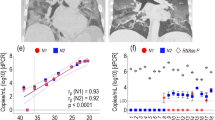
NP samples were positive at a lower CT value than samples from the OP (mean difference in CT 5.75, 95 % CI 3.8–7.7, p < 0.01), corresponding to 54 times (95 % CI 13.7–210.8) higher calculated viral load in the nasopharynx samples compared to those obtained from the oropharynx. Subgroup analysis of influenza A(H1N1)pdm09 showed a mean difference in the calculated viral load of 23 (95 % CI 3.8–136.2, p < 0.01) times between the NP and OP swabs. For influenza B, the mean difference in the calculated viral load was 80 times (95 % CI 9.3–694.6, p < 0.01) higher. No significant difference was found between influenza A(H1N1)pdm09 and influenza B when comparing viral loads.
A total of seven patients with influenza A(H1N1)pdm09 and three patients with influenza B were swabbed more than 24 h after the initiation of antiviral treatment and the mean difference in CT values for these patients was 2.8 (95 % CI 0.7–6.3). This was significantly lower than for the group treated less than 24 h (or not at all), with mean difference in CT values of 7.1 (95 % CI 4.8–9.4, p = 0.035). Four patients with influenza A(H1N1)pdm09 and two with influenza B tested negative in the oropharynx and nasopharynx when swabbed for this study. Only three of these six patients received antiviral treatment. Further patient characteristics are summarized in Table 2.
Fifty-nine percent of the patients had one or more known risk factors for influenza infection. The mean duration of their symptoms before hospitalization was 3.3 (±2.7) days and the mean time from symptom start until swabbing was 4.8 (±2.7) days. The patients were swabbed an average of 1.5 (±0.8) days after hospital admission. Of the six patients who tested negative in both the oropharynx and nasopharynx, the mean [standard deviation (SD)] time from symptom start to swabbing was 6.0 (±1.9) days. Out of the 32 study patients, five were admitted to the intensive care unit, where they spent a median of 3 days (range 1–10 days). Four patients needed respiratory support for a median of 2.5 days (range 1–3 days). No patients received antivirals prior to hospitalization and a total of 13/32 patients received in-hospital antiviral treatment.
Discussion
This study shows that, using the described swabs, material obtained from the nasopharynx contains more influenza virus than that recovered from the oropharynx. The combined calculated diagnostic sensitivity for influenza B and influenza A(H3N2)pdm09 for NP and OP samples were 78 % and 63 %, respectively (χ2 < 0.01).
Our finding that the NP samples contain more virus than the OP samples are in contrast to a prospective study from Kenya, where influenza A(H1N1) were found significantly more often in OP samples and influenza B virus more often in NP samples, whereas with influenza A (H3N2), no significant difference between the two sampling sites were observed[11]. However, the Kenyan study compared a flocked swab used in the nasopharynx with a polyester swab used in the oropharynx. As flocked swabs contain, on average, 4.8 times more virus than rayon swabs, a comparison based on different swab types is difficult to interpret [10]. Lieberman et al. performed NP and OP sampling by rigid cotton swabs as well as by NP washes to detect eight different respiratory virus types [12]. NP washes were found to be superior to NP swabbing, and OP swabbing was the least sensitive of the three sampling methods. As NP washes are difficult to obtain, NP swabbing samples appear to be the most appropriate method, especially in frail individuals [1].
In this selected material of initially PCR-positive individuals, our sampling method reached a sensitivity of 78 % for all observed influenza virus and 71 % for influenza A(H1N1)pdm09 alone. As the initial viral material was collected, on average, 1.5 days before the collection of swabs for the current study, natural viral clearance in some patients would be expected to reduce the sensitivity of the test. On the other hand, the use of absorptive swabs and specially trained technicians probably contributed to the high sensitivity of 78 % for all influenza viruses and 71 % for the H1N1 strain.
In a retrospective cohort study of 25 patients hospitalized for pandemic influenza with community-acquired pneumonia, a comparison between lower respiratory tract specimens and NP swabs revealed a sensitivity of only 63 % for the NP swabs and nearly 98 % for the lower respiratory samples [2]. A lower respiratory tract infection could possibly delay viral clearance in the lungs, leading to a prolonged viral excretion in the lower airways. Therefore, specimens from the bronchial tree could be more suitable than NP samples when investigating patients with pneumonia. However, at least in patients without lower respiratory tract affection, NP swabs represent a low-cost, easily accessible, and quick diagnostic method, and provided a proper swabbing technique.
The viral load is reduced in a nonlinear fashion and different types of influenza virus seem to follow different time lines for viral clearance [3–5, 8, 9, 13]. Samples from influenza A(H1N1)pdm09 patients display a lower viral load than patients suffering from seasonal influenza A (H1N1 and H3N2) [9]. The majority of studies on viral load and clearance have been performed on influenza A subgroups. Although less is known about influenza B in this regard, one study reported continuous viral shedding in 70 % of influenza B patients compared to 33 % among those suffering from influenza A disease after 7 days of illness [4]. We did not find a significant difference in viral load when comparing samples from patients with influenza B and influenza A(H1N1)pdm09. Among our patients, more of the initially influenza A(H1N1)pdm09-positive than influenza B-positive individuals had turned PCR-negative on average 4.8 days after onset of the disease. However, the small numbers involved do not allow any general conclusions be made. The time of sampling, seriousness of the infection, comorbidity, lower respiratory tract infection, sampling method, and vaccination status may all influence the viral load at the sampling site [4, 8, 9, 14]. An assessment of the relative impact on the viral load of such factors would require careful examination of a large group of comparable patients.
One might speculate that the H1N1 dominance of the current study is explained by sampling bias towards detecting influenza A(H1N1)pdm09 rather than influenza A(H3N2).
However, recent subtyping of influenza strains in a comparable study [10] conducted during the influenza season 2008–2009 provided additional H3N2 isolates (Olav Hungnes, Norwegian Institute of Public Health, personal communication). The NP samples for these patients were positive at a lower CT value than the OP samples, with a mean difference in CT value of 10.9 (95 % CI 7.1–14.7, p < 0.001), corresponding to a mean difference in the calculated viral load of 1,911 times (95 % CI 137.2–26,615.9) higher in the nasopharynx than the oropharynx. H1N1 and H3N2 shared the higher affinity for the nasopharynx compared to the oropharynx. This finding makes it unlikely that the H1N1 dominance of the current study is explained by methodological issues.
A strength of our study is the prospective design and the low number of specially trained staff responsible for the swabbing procedure, reducing the possibility of suboptimal techniques. A limitation is that the sensitivity of the diagnostic procedure is evaluated, on average, 1.5 days after admission, and not at the time of hospitalization. The genetic variation in influenza A(H1N1)pdm09 is still limited, with most isolates genetically similar to influenza A/California/07/2009 (H1N1). Therefore, future studies need to be performed when influenza A(H1N1)pdm09 has been thoroughly established as a seasonal pathogen.
To our knowledge, this is the first study comparing the load of influenza viruses in swabbed samples from the oropharynx and nasopharynx of adult patients. We show that, in influenza patients, the etiological diagnosis by PCR is achieved more efficiently by NP than by OP swabbing.
References
Falsey AR, Walsh EE (2006) Viral pneumonia in older adults. Clin Infect Dis 42(4):518–524
Mulrennan S, Tempone SS, Ling IT, Williams SH, Gan GC, Murray RJ, Speers DJ (2010) Pandemic influenza (H1N1) 2009 pneumonia: CURB-65 score for predicting severity and nasopharyngeal sampling for diagnosis are unreliable. PLoS One 5(9):e12849
Carrat F, Vergu E, Ferguson NM, Lemaitre M, Cauchemez S, Leach S, Valleron AJ (2008) Time lines of infection and disease in human influenza: a review of volunteer challenge studies. Am J Epidemiol 167(7):775–785
Lee N, Chan PK, Hui DS, Rainer TH, Wong E, Choi KW, Lui GC, Wong BC, Wong RY, Lam WY, Chu IM, Lai RW, Cockram CS, Sung JJ (2009) Viral loads and duration of viral shedding in adult patients hospitalized with influenza. J Infect Dis 200(4):492–500
Lau LL, Cowling BJ, Fang VJ, Chan KH, Lau EH, Lipsitch M, Cheng CK, Houck PM, Uyeki TM, Peiris JS, Leung GM (2010) Viral shedding and clinical illness in naturally acquired influenza virus infections. J Infect Dis 201(10):1509–1516
Meschi S, Selleri M, Lalle E, Bordi L, Valli MB, Ferraro F, Ippolito G, Petrosillo N, Lauria FN, Capobianchi MR (2011) Duration of viral shedding in hospitalized patients infected with pandemic H1N1. BMC Infect Dis 11:140
Li IW, Hung IF, To KK, Chan KH, Wong SS, Chan JF, Cheng VC, Tsang OT, Lai ST, Lau YL, Yuen KY (2010) The natural viral load profile of patients with pandemic 2009 influenza A(H1N1) and the effect of oseltamivir treatment. Chest 137(4):759–768
Giannella M, Alonso M, Garcia de Viedma D, Lopez Roa P, Catalán P, Padilla B, Muñoz P, Bouza E (2011) Prolonged viral shedding in pandemic influenza A(H1N1): clinical significance and viral load analysis in hospitalized patients. Clin Microbiol Infect 17(8):1160–1165
Lee CK, Lee HK, Loh TP, Lai FY, Tambyah PA, Chiu L, Koay ES, Tang JW (2011) Comparison of pandemic (H1N1) 2009 and seasonal influenza viral loads, Singapore. Emerg Infect Dis 17(2):287–291
Hernes SS, Quarsten H, Hagen E, Lyngroth AL, Pripp AH, Bjorvatn B, Bakke PS (2011) Swabbing for respiratory viral infections in older patients: a comparison of rayon and nylon flocked swabs. Eur J Clin Microbiol Infect Dis 30(2):159–165
Kim C, Ahmed JA, Eidex RB, Nyoka R, Waiboci LW, Erdman D, Tepo A, Mahamud AS, Kabura W, Nguhi M, Muthoka P, Burton W, Breiman RF, Njenga MK, Katz MA (2011) Comparison of nasopharyngeal and oropharyngeal swabs for the diagnosis of eight respiratory viruses by real-time reverse transcription-PCR assays. PLoS One 6(6):e21610
Lieberman D, Lieberman D, Shimoni A, Keren-Naus A, Steinberg R, Shemer-Avni Y (2009) Identification of respiratory viruses in adults: nasopharyngeal versus oropharyngeal sampling. J Clin Microbiol 47(11):3439–3443
Nilsson AC, Brytting M, Serifler F, Björkman P, Persson K, Widell A (2010) Longitudinal clearance of seasonal influenza A viral RNA measured by real-time polymerase chain reaction in patients identified at a hospital emergency department. Scand J Infect Dis 42(9):679–686
Blyth CC, Iredell JR, Dwyer DE (2009) Rapid-test sensitivity for novel swine-origin influenza A (H1N1) virus in humans. N Engl J Med 361(25):2493
Watzinger F, Suda M, Preuner S, Baumgartinger R, Ebner K, Baskova L, Niesters HG, Lawitschka A, Lion T (2004) Real-time quantitative PCR assays for detection and monitoring of pathogenic human viruses in immunosuppressed pediatric patients. J Clin Microbiol 42(11):5189–5198
van Elden LJ, van Essen GA, Boucher CA, van Loon AM, Nijhuis M, Schipper P, Verheij TJ, Hoepelman IM (2001) Clinical diagnosis of influenza virus infection: evaluation of diagnostic tools in general practice. Br J Gen Pract 51(469):630–634
The German National Reference Centre for Influenza (NIC) (2011)
Acknowledgments
The authors are grateful for the skillful help of Rebecca Cox and Olav Hungnes in the preparation of this paper.
This project was financed with the aid of EXTRA funds from the Norwegian Foundation for Health and Rehabilitation in association with the Norwegian Heart and Lung Patient Organisation, from the South-Eastern Norway Regional Health Authority, and from Sorlandet Hospital HF.
Conflict of interest
The nylon flocked swabs and UTM transport tubes were donated free of charge by Copan Italia, Brescia, Italy. The supplier did not participate in the planning, running, evaluation, or reporting of this trial. The authors declare that they have no conflict of interest related to this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hernes, S.S., Quarsten, H., Hamre, R. et al. A comparison of nasopharyngeal and oropharyngeal swabbing for the detection of influenza virus by real-time PCR. Eur J Clin Microbiol Infect Dis 32, 381–385 (2013). https://doi.org/10.1007/s10096-012-1753-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-012-1753-0