Abstract
Background
Parkinson’s disease (PD) is a neurodegenerative disorder characterized by resting tremor, bradykinesia, muscle rigidity, and abnormal gait. The low-density lipoprotein receptor–related protein 10 (LRP10) was recently shown to be a causal gene for PD, and different ethnic cohorts have distinct frequencies and spectrum of LRP10 variants.
Methods
We sequenced the full coding regions and exon–intron boundaries of LRP10 in 129 patients with sporadic Chinese PD to further investigate the connection of LRP10 with PD in a sample of Chinese patients.
Results
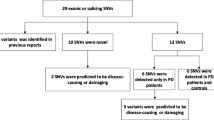
In this study, we identified four potentially pathogenic mutations, including one novel mutation of p.Gly328Asp and three known mutations of p.Cys165Tyr, p.Arg230Trp, and p.Arg661His in four of the 129 Chinese patients with PD.
Conclusion
According to our study, the LRP10 gene may attribute to PD pathogenesis.

Similar content being viewed by others
References
Li, C, Chen, Y, Ou, R, Gu, X, Wei, Q, Cao, B, Zhang, L, Hou, Y, Liu, K, Chen, X, Song, W, Zhao, B, Wu, Y, Shang, H (2021) Mutation analysis of LRP10 in a large Chinese familial Parkinson disease cohort.Neurobiol Aging 99(99 e1–99 e6. https://doi.org/10.1016/j.neurobiolaging.2020.08.015
Obeso JA, Stamelou M, Goetz CG, Poewe W, Lang AE, Weintraub D, Burn D, Halliday GM, Bezard E, Przedborski S, Lehericy S, Brooks DJ, Rothwell JC, Hallett M, Delong MR, Marras C, Tanner CM, Ross GW, Langston JW, Klein C, Bonifati V, Jankovic J, Lozano AM, Deuschl G, Bergman H, Tolosa E, Rodriguez-Violante M, Fahn S, Postuma RB, Berg D, Marek K, Standaert DG, Surmeier DJ, Olanow CW, Kordower JH, Calabresi P, Schapira AHV, Stoessl AJ (2017) Past, present, and future of Parkinson’s disease: a special essay on the 200th Anniversary of the Shaking Palsy. Mov Disord 32(9):1264–1310. https://doi.org/10.1002/mds.27115
Morgante F, Valente EM (2019) LRP10: a novel disease gene bridging Parkinson’s disease and dementia with Lewy body. Mov Disord 34(1):47. https://doi.org/10.1002/mds.27593
Iwaki, H, Blauwendraat, C, Leonard, HL, Kim, JJ, Liu, G, Maple-Grodem, J, Corvol, JC, Pihlstrom, L, Van Nimwegen, M, Hutten, SJ, Nguyen, KH, Rick, J, Eberly, S, Faghri, F, Auinger, P, Scott, KM, Wijeyekoon, R, Van Deerlin, VM, Hernandez, DG, Gibbs, JR, International Parkinson’s Disease Genomics, C, Chitrala, KN, Day-Williams, AG, Brice, A, Alves, G, Noyce, AJ, Tysnes, OB, Evans, JR, Breen, DP, Estrada, K, Wegel, CE, Danjou, F, Simon, DK, Andreassen, O, Ravina, B, Toft, M, Heutink, P, Bloem, BR, Weintraub, D, Barker, RA, Williams-Gray, CH, Van De Warrenburg, BP, Van Hilten, JJ, Scherzer, CR, Singleton, AB, Nalls, MA (2019) Genomewide association study of Parkinson’s disease clinical biomarkers in 12 longitudinal patients’ cohorts. Mov Disord 34(12):1839–1850. https://doi.org/10.1002/mds.27845
Blauwendraat C, Nalls MA, Singleton AB (2020) The genetic architecture of Parkinson’s disease. Lancet Neurol 19(2):170–178. https://doi.org/10.1016/S1474-4422(19)30287-X
Lunati A, Lesage S, Brice A (2018) The genetic landscape of Parkinson’s disease. Rev Neurol (Paris) 174(9):628–643. https://doi.org/10.1016/j.neurol.2018.08.004
Quadri, M, Mandemakers, W, Grochowska, MM, Masius, R, Geut, H, Fabrizio, E, Breedveld, GJ, Kuipers, D, Minneboo, M, Vergouw, LJM, Carreras Mascaro, A, Yonova-Doing, E, Simons, E, Zhao, T, Di Fonzo, AB, Chang, HC, Parchi, P, Melis, M, Correia Guedes, L, Criscuolo, C, Thomas, A, Brouwer, RWW, Heijsman, D, Ingrassia, AMT, Calandra Buonaura, G, Rood, JP, Capellari, S, Rozemuller, AJ, Sarchioto, M, Fen Chien, H, Vanacore, N, Olgiati, S, Wu-Chou, YH, Yeh, TH, Boon, AJW, Hoogers, SE, Ghazvini, M, As, IJ, Van, IWFJ, Onofrj, M, Barone, P, Nicholl, DJ, Puschmann, A, De Mari, M, Kievit, AJ, Barbosa, E, De Michele, G, Majoor-Krakauer, D, Van Swieten, JC, De Jong, FJ, Ferreira, JJ, Cossu, G, Lu, CS, Meco, G, Cortelli, P, Van De Berg, WDJ, Bonifati, V, International Parkinsonism Genetics, N (2018) LRP10 genetic variants in familial Parkinson’s disease and dementia with Lewy bodies: a genome-wide linkage and sequencing study. Lancet Neurol 17(7):597–608. https://doi.org/10.1016/S1474-4422(18)30179-0
Liao TW, Wang CC, Chung WH, Su SC, Chin SH, Fung HC, Wu YR (2021) Role of LRP10 in Parkinson’s disease in a Taiwanese cohort Parkinsonism. Relat Disord 89:79–83. https://doi.org/10.1016/j.parkreldis.2021.06.026
Zhao Q, Ning P, Yang X, Shi C, Xu Y, Shen Q, Huang H, Xie D, Chen Y, Xu Y (2021) LRP10 mutations may correlate with sporadic Parkinson’s disease in China. Mol Neurobiol 58(3):1212–1216. https://doi.org/10.1007/s12035-020-02186-9
Manini, A, Straniero, L, Monfrini, E, Percetti, M, Vizziello, M, Franco, G, Rimoldi, V, Zecchinelli, A, Pezzoli, G, Corti, S, Comi, GP, Duga, S, Di Fonzo, A (2021) Screening of LRP10 mutations in Parkinson’s disease patients from Italy.Parkinsonism Relat Disord 89(17-21. https://doi.org/10.1016/j.parkreldis.2021.06.014
Perinan MT, Macias-Garcia D, Buiza-Rueda D, Guijarro-Albaladejo B, Jesus S, Adarmes-Gomez AD, Escuela R, Vigo-Ortega R, Gomez-Garre P, Mir P (2020) Analysis of p Tyr307Asn variant in the LRP10 gene in Parkinson’s disease in southern Spain. Neurobiol Aging 93142(e1–142):e3. https://doi.org/10.1016/j.neurobiolaging.2020.04.007
Vergouw LJM, Geut H, Breedveld G, Kuipers DJS, Quadri M, Netherlands Brain B, Rozemuller AJM, Van Swieten JC, De Jong FJ, Van De Berg WDJ, Bonifati V (2020) Clinical and pathological phenotypes of LRP10 variant carriers with dementia. J Alzheimers Dis 76(3):1161–1170. https://doi.org/10.3233/JAD-200318
Chen Y, Cen Z, Zheng X, Pan Q, Chen X, Zhu L, Chen S, Wu H, Xie F, Wang H, Yang D, Wang L, Zhang B, Luo W (2019) LRP10 in autosomal-dominant Parkinson’s disease. Mov Disord 34(6):912–916. https://doi.org/10.1002/mds.27693
Vergouw LJM, Ruitenberg A, Wong TH, Melhem S, Breedveld GJ, Criscuolo C, De Michele G, De Jong FJ, Bonifati V, Van Swieten JC, Quadri M (2019) LRP10 variants in Parkinson’s disease and dementia with Lewy bodies in the South-West of the Netherlands. Parkinsonism. Relat Disord 65:243–247. https://doi.org/10.1016/j.parkreldis.2019.05.037
Shi, CH, Luo, HY, Fan, Y, Li, YS, Xu, YM (2018) LRP10 in alpha-synucleinopathies.Lancet Neurol 17(12):1034–1035. https://doi.org/10.1016/S1474-4422(18)30402-2
Gagliardi M, Procopio R, Nicoletti G, Morelli M, D’amelio M, Quattrone A, Annesi G (2021) Analysis of the LRP10 gene in patients with Parkinson’s disease and dementia with Lewy bodies from Southern Italy. Neurol Sci 42(1):305–308. https://doi.org/10.1007/s10072-020-04747-1
Daida, K, Nishioka, K, Li, Y, Yoshino, H, Kikuchi, A, Hasegawa, T, Funayama, M, Hattori, N (2019) Mutation analysis of LRP10 in Japanese patients with familial Parkinson’s disease, progressive supranuclear palsy, and frontotemporal dementia. Neurobiol Aging 84 235 e11–235 e16. https://doi.org/10.1016/j.neurobiolaging.2019.08.030
Kia, DA, Sabir, MS, Ahmed, S, Trinh, J, Bandres-Ciga, S, International Parkinson’s disease genomics, C (2018) LRP10 in alpha-synucleinopathies. Lancet Neurol 17(12):1032. https://doi.org/10.1016/S1474-4422(18)30401-0
Guerreiro, R, Orme, T, Neto, JL, Bras, J, International, DLBGC (2018) LRP10 in alpha-synucleinopathies. Lancet Neurol 17(12):1032–1033. https://doi.org/10.1016/S1474-4422(18)30399-5
Pihlstrom, L, Schottlaender, L, Chelban, V, Houlden, H, Consortium, MSAE (2018) LRP10 in alpha-synucleinopathies. Lancet Neurol 17(12):1033–1034. https://doi.org/10.1016/S1474-4422(18)30407-1
Tesson C, Brefel-Courbon C, Corvol JC, Lesage S, Brice A, French Parkinson’s disease genetics study, G, (2018) LRP10 in alpha-synucleinopathies. Lancet Neurol 17(12):1034. https://doi.org/10.1016/S1474-4422(18)30400-9
Zhao Y, Qin L, Pan H, Liu Z, Jiang L, He Y, Zeng Q, Zhou X, Zhou X, Zhou Y, Fang Z, Wang Z, Xiang Y, Yang H, Wang Y, Zhang K, Zhang R, He R, Zhou X, Zhou Z, Yang N, Liang D, Chen J, Zhang X, Zhou Y, Liu H, Deng P, Xu K, Xu K, Zhou C, Zhong J, Xu Q, Sun Q, Li B, Zhao G, Wang T, Chen L, Shang H, Liu W, Chan P, Xue Z, Wang Q, Guo L, Wang X, Xu C, Zhang Z, Chen T, Lei L, Zhang H, Wang C, Tan J, Yan X, Shen L, Jiang H, Zhang Z, Hu Z, Xia K, Yue Z, Li J, Guo J, Tang B (2020) The role of genetics in Parkinson’s disease: a large cohort study in Chinese mainland population. Brain 143(7):2220–2234. https://doi.org/10.1093/brain/awaa167
Postuma RB, Berg D, Stern M, Poewe W, Olanow CW, Oertel W, Obeso J, Marek K, Litvan I, Lang AE, Halliday G, Goetz CG, Gasser T, Dubois B, Chan P, Bloem BR, Adler CH, Deuschl G (2015) MDS clinical diagnostic criteria for Parkinson’s disease. Mov Disord 30(12):1591–601. https://doi.org/10.1002/mds.26424
Postuma RB, Poewe W, Litvan I, Lewis S, Lang AE, Halliday G, Goetz CG, Chan P, Slow E, Seppi K, Schaffer E, Rios-Romenets S, Mi T, Maetzler C, Li Y, Heim B, Bledsoe IO, Berg D (2018) Validation of the MDS clinical diagnostic criteria for Parkinson’s disease. Mov Disord 33(10):1601–1608. https://doi.org/10.1002/mds.27362
Song N, Wang W, Chen C, Niu J, Guojin Y, Guo C, Han F (2018) Analysis of SMPD1 gene mutations in Chinese patients with Parkinson’s disease. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 35(3):319–323. https://doi.org/10.3760/cma.j.issn.1003-9406.2018.03.003
Del Rey NL, Quiroga-Varela A, Garbayo E, Carballo-Carbajal I, Fernandez-Santiago R, Monje MHG, Trigo-Damas I, Blanco-Prieto MJ, Blesa J (2018) Advances in Parkinson’s disease: 200 years later. Front Neuroanat 12:113. https://doi.org/10.3389/fnana.2018.00113
Vilarino-Guell C, Wider C, Ross OA, Dachsel JC, Kachergus JM, Lincoln SJ, Soto-Ortolaza AI, Cobb SA, Wilhoite GJ, Bacon JA, Behrouz B, Melrose HL, Hentati E, Puschmann A, Evans DM, Conibear E, Wasserman WW, Aasly JO, Burkhard PR, Djaldetti R, Ghika J, Hentati F, Krygowska-Wajs A, Lynch T, Melamed E, Rajput A, Rajput AH, Solida A, Wu RM, Uitti RJ, Wszolek ZK, Vingerhoets F, Farrer MJ (2011) VPS35 mutations in Parkinson disease. Am J Hum Genet 89(1):162–167. https://doi.org/10.1016/j.ajhg.2011.06.001
Wang J, Li Y, Gao L, Yan F, Gao G, Li L (2018) GSK-3beta inhibitor alsterpaullone attenuates MPP(+)-induced cell damage in a c-Myc-dependent manner in SH-SY5Y cells. Front Cell Neurosci 12:283. https://doi.org/10.3389/fncel.2018.00283
Arias-Fuenzalida J, Jarazo J, Qing X, Walter J, Gomez-Giro G, Nickels SL, Zaehres H, Scholer HR, Schwamborn JC (2017) FACS-assisted CRISPR-Cas9 genome editing facilitates Parkinson’s disease modeling. Stem Cell Reports 9(5):1423–1431. https://doi.org/10.1016/j.stemcr.2017.08.026
Acknowledgements
We thank all the participants.
Funding
This study was supported by grants from the Shandong Provincial Natural Science Foundation (Nos. ZR2019ZD39 and ZR2020KH026), National Natural Science Foundation of China (Nos. 81800980 and 81972989).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics approval
This study was approved by the Ethical Review Committees of Liaocheng People’s Hospital.
Consent to participate
The informed consent was signed by each participant.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Song, N., Wang, Y., Zhou, L. et al. Genetic analysis of the LRP10 gene in Chinese patients with Parkinson’s disease. Neurol Sci 44, 905–912 (2023). https://doi.org/10.1007/s10072-022-06496-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10072-022-06496-9