Abstract
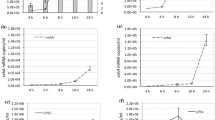
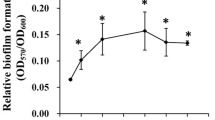
Effects of elevated intracellular 3′,5′-cyclic diguanylic acid (c-di-GMP) levels on biofilm formation and transcription profiles were evaluated to assess the functions of c-di-GMP in Vibrio vulnificus. Elevated c-di-GMP levels promoted biofilm formation and rugose colony development. Microarray analysis revealed that c-di-GMP influenced expression of genes belonging to different functional categories and more than 5% of the V. vulnificus genome. Among these, 10 genes potentially involved in biofilm formation were experimentally verified as subject to c-di-GMP regulation. c-di-GMP contributes to biofilm formation based on modulation of diverse cellular processes in V. vulnificus.
Similar content being viewed by others
References
Flemming HC, Wingender J. The biofilm matrix. Nat. Rev. Microbiol. 8: 623–633 (2012)
Costerton JW, Cheng KJ, Geesey GG, Ladd TI, Nickel JC, Dasgupta M, Marrie TJ. Bacterial biofilms in nature and disease. Annu. Rev. Microbiol. 41: 435–464 (1987)
Johnson LR. Microcolony and biolm formation as a survival strategy for bacteria. J. Theor. Biol. 251: 24–34 (2008)
Marco-Noales E, Milán M, Fouz B, Sanjuán E, Amaro C. Transmission to eels, portals of entry, and putative reservoirs of Vibrio vulnificus serovar E (biotype 2). Appl. Environ. Microb. 67: 4717–4725 (2001)
Guo Y, Rowe-Magnus DA. Identification of a c-di-GMP-regulated polysaccharide locus governing stress resistance and biofilm and rugose colony formation in Vibrio vulnificus. Infect. Immun. 78: 1390–1402 (2010)
Guo Y, Rowe-Magnus DA. Overlapping and unique contributions of two conserved polysaccharide loci in governing distinct survival phenotypes in Vibrio vulnificus. Environ. Microbiol. 13: 2888–2990 (2011)
Grau BL, Henk MC, Pettis GS. High-frequency phase variation of Vibrio vulnificus 1003: Isolation and characterization of a rugose phenotypic variant. J. Bacteriol. 187: 2519–2525 (2005)
Paranjpye RN, Johnson AB, Baxter AE, Strom MS. Role of type IV pilins in persistence of Vibrio vulnificus in Crassostrea virginica oysters. Appl. Environ. Microb. 73: 5041–5044 (2007)
Kim SM, Park JH, Lee HS, Kim WB, Ryu JM, Han HJ, Choi SH. LuxR homologue SmcR is essential for Vibrio vulnificus pathogenesis and biofilm detachment, and its expression is induced by host cells. Infect. Immun. 81: 3721–3730 (2013)
Boyd CD, O’Toole GA. Second messenger regulation of biofilm formation: Breakthroughs in understanding c-di-GMP effector systems. Annu. Rev. Cell Dev. Bi. 28: 439–462 (2012)
Hengge R. Principles of c-di-GMP signaling in bacteria. Nat. Rev. Microbiol. 7: 263–273 (2009)
Nakhamchik A, Wilde C, Rowe-Magnus DA. Cyclic-di-GMP regulates extracellular polysaccharide production, biofilm formation, and rugose colony development by Vibrio vulnificus. Appl. Environ. Microb. 74: 4199–4209 (2008)
Goo SY, Lee HJ, Kim WH, Han KL, Park DK, Lee HJ, Kim SM, Kim KS, Lee KH, Park SJ. Identication of OmpU of Vibrio vulnicus as a bronectin-binding protein and its role in bacterial pathogenesis. Infect. Immun. 74: 5586–5594 (2006)
Irie Y, Borlee BR, O’Connor JR, Hill PJ, Harwood CS, Wozniak DJ, Parsek MR. Self-produced exopolysaccharide is a signal that stimulates biolm formation in Pseudomonas aeruginosa. P. Natl. Acad. Sci. USA 109: 20632–20636 (2012)
Russo DM, Williams A, Edwards A, Posadas DM, Finnie C, Dankert M, Downie JA, Zorreguieta A. Proteins exported via the PrsD-PrsE type I secretion system and the acidic exopolysaccharide are involved in biofilm formation by Rhizobium leguminosarum. J. Bacteriol. 188: 4474–4486 (2006)
Kim HS, Park SJ, Lee KH. Role of NtrC-regulated exopolysaccharides in the biolm formation and pathogenic interaction of Vibrio vulnificus. Mol. Microbiol. 74: 436–453 (2009)
Greenberg EP, Hastings JW, Ulitzur S. Induction of luciferase synthesis in Beneckea harveyi by other marine bacteria. Arch. Microbiol. 120: 87–91 (1979)
Edgar R, Domrachev M, Lash AE. Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30: 207–210 (2002)
Barrios AF, Zuo R, Ren D, Wood TK. Hha, YbaJ, and OmpA regulate Escherichia coli K12 biolm formation and conjugation plasmids abolish motility. Biotechnol. Bioeng. 93: 188–200 (2006)
Smith SG, Mahon V, Lambert MA, Fagan RP. A molecular Swiss army knife: OmpA structure, function, and expression. FEMS Microbiol. Lett. 273: 1–11 (2007)
Enos-Berlage JL, Guvener ZT, Keenan CE, McCarter LL. Genetic determinants of biofilm development of opaque and translucent Vibrio parahaemolyticus. Mol. Microbiol. 55: 1160–1182 (2005)
Liang Y, Gao H, Chen J, Dong Y, Wu L, He Z, Liu X, Qiu G, Zhou J. Pellicle formation in Shewanella oneidensis. BMC Microbiol. 10: 291 (2010)
Ferreira RB, Chodur DM, Antunes LC, Trimble MJ, McCarter LL. Output targets and transcriptional regulation by a cyclic dimeric GMP-responsive circuit in the Vibrio parahaemolyticus Scr network. J. Bacteriol. 194: 914–924 (2012)
Satchell KJ. Structure and function of MARTX toxins and other large repetitive RTX proteins. Annu. Rev. Microbiol. 65: 71–90 (2011)
Casper-Lindley C, Yildiz FH. VpsT is a transcriptional regulator required for expression of vps biosynthesis genes and the development of rugose colonial morphology in Vibrio cholerae O1 El Tor. J. Bacteriol. 186: 1574–1578 (2004)
Simon R, Priefer U, Pühler A. A broad host range mobilization system for in vivo genetic engineering transposon mutagenesis in gram negative bacteria. Nat. Biotechnol. 1: 784–791 (1983)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Park, J.H., Lim, J.G. & Choi, S.H. Effects of elevated intracellular cyclic di-GMP levels on biofilm formation and transcription profiles of Vibrio vulnificus . Food Sci Biotechnol 24, 771–776 (2015). https://doi.org/10.1007/s10068-015-0100-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10068-015-0100-5