Abstract
Objective
Rheumatoid arthritis (RA) is a common disease of rheumatic diseases. The aim of this study was to identify gene signatures in RA and uncover their potential mechanisms.
Method
Gene expression profiles of GSE1919, GSE55235, GSE55457, and GSE77928 were downloaded from GEO database. The above four series contained 76 samples, including 44 RA patients and 32 normal controls. The gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed, and protein–protein interaction (PPI) network of the differentially expressed genes (DEGs) was constructed by Cytoscape software.
Results
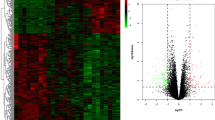
Up-regulated DEGs were significantly enriched in biological processes, including immune response, positive regulation of immune system process and regulation of immune system process, while down-regulated DEGs were significantly enriched in biological processes, including response to oxygen-containing compound, cellular lipid metabolic process, and lipid metabolic process. KEGG pathway analysis showed the up-regulated DEGs were enriched in cytokine–cytokine receptor interaction, chemokine signaling pathway, and primary immunodeficiency. The 104 hub genes, which were significantly differently expressed between patients and normal controls in at least two datasets, were identified from the PPI network, and subnetworks revealed that these genes were involved in significant pathways, including cytokine–cytokine receptor interaction, chemokine signaling pathway, and primary immunodeficiency.
Conclusion
The present study indicated that the identified DEGs and hub genes promote our understanding of molecular mechanisms underlying the development of RA, such as C-C motif chemokine 5 (CCL5), might have a negative impact in the development of RA. CCL5 and its related genes might be the potential diagnostic biomarkers for the therapeutic strategies of RA.



Similar content being viewed by others
References
Siebert S, Lyall DM, Mackay DF, Porter D, McInnes IB, Sattar N, Pell JP (2016) Characteristics of rheumatoid arthritis and its association with major comorbid conditions: cross-sectional study of 502 649 UK Biobank participants. RMD Open 2(1):e267
McInnes IB, Schett G (2017) Pathogenetic insights from the treatment of rheumatoid arthritis. Lancet 389(10086):2328–2337
McInnes IB, Schett G (2011) The pathogenesis of rheumatoid arthritis. N Engl J Med 365(23):2205–2219
Brennan P, Ollier B, Worthington J, Hajeer A, Silman A (1996) Are both genetic and reproductive associations with rheumatoid arthritis linked to prolactin? Lancet 348(9020):106–109
Y Okada DW, Trynka G, Raj T, Terao C, Ikari K, Kochi Y, Ohmura K, Suzuki A, Yoshida S, Graham RR, Manoharan A, Ortmann W, Bhangale T, Denny JC, Carroll RJ, Eyler AE, Greenberg JD, Kremer JM, Pappas DA, Jiang L, Yin J, L Ye DFS, Yang J, Xie G, Keystone E, Westra HJ, Esko T, Metspalu A, Zhou X, Gupta N, Mirel D, Stahl EA, Diogo D, Cui J, Liao K, Guo MH, Myouzen K, Kawaguchi T, Coenen MJ, van Riel PL, van de Laar MA, Guchelaar HJ, Huizinga TW, Dieude P, Mariette X, Bridges SJ, Zhernakova A, Toes RE, Tak PP, Miceli-Richard C, Bang SY, Lee HS, Martin J, Gonzalez-Gay MA, Rodriguez-Rodriguez L, Rantapaa-Dahlqvist S, Arlestig L, Choi HK, Kamatani Y, Galan P, Lathrop M, Eyre S, Bowes J, Barton A, de Vries N, Moreland LW, Criswell LA, Karlson EW, Taniguchi A, Yamada R, Kubo M, Liu JS, Bae SC, Worthington J, Padyukov L, Klareskog L, Gregersen PK, Raychaudhuri S, Stranger BE, De Jager PL, Franke L, Visscher PM, Brown MA, Yamanaka H, Mimori T, A Takahashi HX, Behrens TW, Siminovitch KA, Momohara S, Matsuda F, Yamamoto K, Plenge RM (2014) Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506(7488):376–381
Gavrila BI, Ciofu C, Stoica V (2016) Biomarkers in rheumatoid arthritis, what is new? J Med Life 9(2):144–148
Harley JB, Chen X, Pujato M, Miller D, Maddox A, Forney C, Magnusen AF, Lynch A, Chetal K, Yukawa M, Barski A, Salomonis N, Kaufman KM, Kottyan LC, Weirauch MT (2018) Transcription factors operate across disease loci, with EBNA2 implicated in autoimmunity. Nat Genet 50(5):699–707
Ishigaki K, Kochi Y, Suzuki A, Tsuchida Y, Tsuchiya H, Sumitomo S, Yamaguchi K, Nagafuchi Y, Nakachi S, Kato R, Sakurai K, Shoda H, Ikari K, Taniguchi A, Yamanaka H, Miya F, Tsunoda T, Okada Y, Momozawa Y, Kamatani Y, Yamada R, Kubo M, Fujio K, Yamamoto K (2017) Polygenic burdens on cell-specific pathways underlie the risk of rheumatoid arthritis. Nat Genet 49(7):1120–1125
Park PJ (2009) ChIP–seq: advantages and challenges of a maturing technology. Nat Rev Genet 10(10):669–680
Zipporah EB, Govarthanan K, Shyamsunder P, Verma RS (2018) Expression profiling of differentially regulated genes in Fanconi anemia. Methods Mol Biol 1783:243–258
Sedlazeck FJ, Lee H, Darby CA, Schatz MC (2018) Piercing the dark matter: bioinformatics of long-range sequencing and mapping. Nat Rev Genet 19(6):329–346
Woetzel D, Huber R, Kupfer P, Pohlers D, Pfaff M, Driesch D, Haupl T, Koczan D, Stiehl P, Guthke R, Kinne RW (2014) Identification of rheumatoid arthritis and osteoarthritis patients by transcriptome-based rule set generation. Arthritis Res. Ther. 16(2):R84
Ungethuem U, Haeupl T, Witt H, Koczan D, Krenn V, Huber H, von Helversen TM, Drungowski M, Seyfert C, Zacher J, Pruss A, Neidel J, Lehrach H, Thiesen HJ, Ruiz P, Blass S (2010) Molecular signatures and new candidates to target the pathogenesis of rheumatoid arthritis. Physiol Genomics 42A(4):267–282
Broeren MG, de Vries M, Bennink MB, Arntz OJ, Blom AB, Koenders MI, van Lent PL, van der Kraan PM, van den Berg WB, van de Loo FA (2016) Disease-regulated gene therapy with anti-inflammatory interleukin-10 under the control of the CXCL10 promoter for the treatment of rheumatoid arthritis. Hum Gene Ther 27(3):244–254
Jin S, Li M, Fang Y, Li Q, Liu J, Duan X, Liu Y, R W, Shi X, Wang Y, Jiang Z, Wang Y, C Y, Wang Q, Tian X, Zhao Y, Zeng X (2017) Chinese Registry of rheumatoid arthritis (CREDIT): II. Prevalence and risk factors of major comorbidities in Chinese patients with rheumatoid arthritis. Arthritis Res. Ther. 19(1):251
Shafia S, Dilafroze FAS, Rasool R, Javeed S, Shah ZA (2014) Rheumatoid arthritis and genetic variations in cytokine genes: a population-based study in Kashmir Valley. Immunol Investig 43(4):349–359
Glant TT, Mikecz K, Rauch TA (2014) Epigenetics in the pathogenesis of rheumatoid arthritis. BMC Med 12:35
Klein K, Gay S (2015) Epigenetics in rheumatoid arthritis. Curr Opin Rheumatol 27(1):76–82
Chen YJ, Chang WA, Hsu YL, Chen CH, Kuo PL (2017) Deduction of novel Genes potentially involved in osteoblasts of rheumatoid arthritis using next-generation sequencing and bioinformatic approaches. Int J Mol Sci 18(11):pii:E2396
Rombouts Y, Willemze A, van Beers JJ, Shi J, Kerkman PF, van Toorn L, Janssen GM, Zaldumbide A, Hoeben RC, Pruijn GJ, Deelder AM, Wolbink G, Rispens T, van Veelen PA, Huizinga TW, Wuhrer M, Trouw LA, Scherer HU, Toes RE (2016) Extensive glycosylation of ACPA-IgG variable domains modulates binding to citrullinated antigens in rheumatoid arthritis. Ann Rheum Dis 75(3):578–585
Luterek-Puszynska K, Malinowski D, Paradowska-Gorycka A, Safranow K, Pawlik A (2017) CD28, CTLA-4 and CCL5 gene polymorphisms in patients with rheumatoid arthritis. Clin Rheumatol 36(5):1129–1135
Withrow J, Murphy C, Liu Y, Hunter M, Fulzele S, Hamrick MW (2016) Extracellular vesicles in the pathogenesis of rheumatoid arthritis and osteoarthritis. Arthritis Res. Ther. 18(1):286
Cosenza S, Ruiz M, Maumus M, Jorgensen C, Noel D (2017) Pathogenic or therapeutic extracellular vesicles in rheumatic diseases: role of mesenchymal stem cell-derived vesicles.30. B-cell phenotype and IgD-CD27- memory B cells are affected by TNF-inhibitors and tocilizumab treatment in rheumatoid arthritis. Int J Mol Sci 18(4)
Song J, Kim D, Han J, Kim Y, Lee M, Jin EJ (2015) PBMC and exosome-derived Hotair is a critical regulator and potent marker for rheumatoid arthritis. Clin Exp Med 15(1):121–126
Yoo J, Lee SK, Lim M, Sheen D, Choi EH, Kim SA (2017) Exosomal amyloid A and lymphatic vessel endothelial hyaluronic acid receptor-1 proteins are associated with disease activity in rheumatoid arthritis. Arthritis Res Ther 19(1):119
Wallace DJ, Gavin IM, Karpenko O, Barkhordar F, Gillis BS (2015) Cytokine and chemokine profiles in fibromyalgia, rheumatoid arthritis and systemic lupus erythematosus: a potentially useful tool in differential diagnosis. Rheumatol Int 35(6):991–996
Mo YQ, Dai L, Zheng DH, Zhu LJ, Wei XN, Pessler F, Shen J, Zhang BY (2011) Synovial infiltration with CD79a-positive B cells, but not other B cell lineage markers, correlates with joint destruction in rheumatoid arthritis. J Rheumatol 38(11):2301–2308
Farag AK, Elkamhawy A, Londhe AM, Lee KT, Pae AN, Roh EJ (2017) Novel LCK/FMS inhibitors based on phenoxypyrimidine scaffold as potential treatment for inflammatory disorders. Eur J Med Chem 141:657–675
Han BK, Olsen NJ, Bottaro A (2016) The CD27–CD70 pathway and pathogenesis of autoimmune disease. Semin Arthritis Rheum 45(4):496–501
Moura RA, Quaresma C, Vieira AR, Goncalves MJ, Polido-Pereira J, Romao VC, Martins N, Canhao H, Fonseca JE (2017) PLoS One 12(9):e182927
Burtea C, Laurent S, Sanli T, Fanfone D, Devalckeneer A, Sauvage S, Beckers MC, Rorive S, Salmon I, Vander EL, Lauwerys BR, Muller RN (2016) Screening for peptides targeted to IL-7Rα for molecular imaging of rheumatoid arthritis synovium. Arthritis Res. Ther. 18(1):230
Szekanecz Z, Koch AE (2016) Successes and failures of chemokine-pathway targeting in rheumatoid arthritis. Nat Rev Rheumatol 12(1):5–13
Yoshida S, Arakawa F, Higuchi F, Ishibashi Y, Goto M, Sugita Y, Nomura Y, Niino D, Shimizu K, Aoki R, Hashikawa K, Kimura Y, Yasuda K, Tashiro K, Kuhara S, Nagata K, Ohshima K (2012) Gene expression analysis of rheumatoid arthritis synovial lining regions by cDNA microarray combined with laser microdissection: up-regulation of inflammation-associated STAT1, IRF1, CXCL9, CXCL10, and CCL5. Scand J Rheumatol 41(3):170–179
Yarilina A, Park-Min KH, T Antoniv XH, Ivashkiv LB (2008) TNF activates an IRF1-dependent autocrine loop leading to sustained expression of chemokines and STAT1-dependent type I interferon–response genes. Nat Immunol 9(4):378–387
Pandya JM, Lundell AC, Andersson K, Nordstrom I, Theander E, Rudin A (2017) Blood chemokine profile in untreated early rheumatoid arthritis: CXCL10 as a disease activity marker. Arthritis Res Ther 19(1):20
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosures
None.
Human participants and animal rights
The article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Huang, Y., Zheng, S., Wang, R. et al. CCL5 and related genes might be the potential diagnostic biomarkers for the therapeutic strategies of rheumatoid arthritis. Clin Rheumatol 38, 2629–2635 (2019). https://doi.org/10.1007/s10067-019-04533-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10067-019-04533-1