Abstract
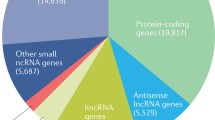
Pervasive transcription of the genome produces a diverse array of functional non-coding RNAs (ncRNAs). One particular class of ncRNAs, long intervening non-coding RNAs (lincRNAs) are thought to play a role in regulating gene expression and may be a major contributor to organism and tissue complexity. The human brain with its heterogeneous cellular make-up is a rich source of lincRNAs; however, the functions of the majority of lincRNAs are unknown. Recently, by completing RNA sequencing (RNA-Seq) of the human frontal cortex, we identified linc00320 as being highly expressed in the white matter compared to grey matter in multiple system atrophy (MSA) brain. Here, we further investigate the expression patterns of linc00320 and conclude that it is involved in specific brain regions rather than having involvement in the MSA disease process. We also show that the full-length linc00320 is only expressed in human brain tissue and not in other primates, suggesting that it may be involved in improved functional connectivity for higher human brain cognition.





Similar content being viewed by others
References
Vinogradov AE, Anatskaya OV (2007) Organismal complexity, cell differentiation and gene expression: human over mouse. Nucleic Acids Res 35(19):6350–6356. doi:10.1093/nar/gkm723
Nolte J, Sundsten J (2002) The human brain: an introduction to its functional anatomy, 5th edn. Mosby, China
Saladin K (2010) Anatomy and physiology: the unity of form and function, 5th edn. McGraw-Hill, New York
Martin J, Howard J, Leonard M (2004) Neuroanatomy: text and atlas, vol., 3rd edn. McGraw-Hill, USA
Underwood J, Cross S (2009) General and systematic pathology, 5th edn. Elsevier Limited, China
Schoenemann PT, Sheehan MJ, Glotzer LD (2005) Prefrontal white matter volume is disproportionately larger in humans than in other primates. Nat Neurosci 8(2):242–252. doi:10.1038/nn1394
Smaers JB, Schleicher A, Zilles K, Vinicius L (2010) Frontal white matter volume is associated with brain enlargement and higher structural connectivity in anthropoid primates. PLoS One 5(2):e9123. doi:10.1371/journal.pone.0009123
Zhang K, Sejnowski TJ (2000) A universal scaling law between gray matter and white matter of cerebral cortex. Proc Natl Acad Sci U S A 97(10):5621–5626. doi:10.1073/pnas.090504197
Mills JD, Kavanagh T, Kim WS, Chen BJ, Kawahara Y, Halliday GM, Janitz M (2013) Unique transcriptome patterns of the white and grey matter corroborate structural and functional heterogeneity in the human frontal lobe. PLoS One 8(10):e78480. doi:10.1371/journal.pone.0078480
Mattick JS (2011) The central role of RNA in human development and cognition. FEBS Lett 585(11):1600–1616. doi:10.1016/j.febslet.2011.05.001
Mattick JS (2001) Non-coding RNAs: the architects of eukaryotic complexity. EMBO Rep 2(11):986–991. doi:10.1093/embo-reports/kve230
Qureshi IA, Mehler MF (2012) Emerging roles of non-coding RNAs in brain evolution, development, plasticity and disease. Nat Rev Neurosci 13(8):528–541. doi:10.1038/nrn3234
Qureshi IA, Mattick JS, Mehler MF (2010) Long non-coding RNAs in nervous system function and disease. Brain Res 1338:20–35. doi:10.1016/j.brainres.2010.03.110
Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL (2011) Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25(18):1915–1927. doi:10.1101/gad.17446611
Duret L, Chureau C, Samain S, Weissenbach J, Avner P (2006) The Xist RNA gene evolved in eutherians by pseudogenization of a protein-coding gene. Science 312(5780):1653–1655. doi:10.1126/science.1126316
Ulitsky I, Shkumatava A, Jan CH, Sive H, Bartel DP (2011) Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147(7):1537–1550. doi:10.1016/j.cell.2011.11.055
Sone M, Hayashi T, Tarui H, Agata K, Takeichi M, Nakagawa S (2007) The mRNA-like noncoding RNA Gomafu constitutes a novel nuclear domain in a subset of neurons. J Cell Sci 120(Pt 15):2498–2506. doi:10.1242/jcs.009357
Rapicavoli NA, Poth EM, Blackshaw S (2010) The long noncoding RNA RNCR2 directs mouse retinal cell specification. BMC Dev Biol 10:49. doi:10.1186/1471-213X-10-49
Mills JD, Kim WS, Halliday GM, Janitz M (2014) Transcriptome analysis of grey and white matter cortical tissue in multiple system atrophy. Neurogenetics. doi:10.1007/s10048-014-0430-0
Mills JD, Kavanagh T, Kim WS, Chen B, Waters PD, Halliday GM, Janitz M (2015) High expression of long intervening non-coding RNA OLMALINC in the human cortical white matter is associated with regulation of oligodendrocyte maturation. Mol Brain 8(1):2. doi:10.1186/s13041-014-0091-9
Gilman S, Wenning G, Low P, Brooks D, Mathias C, Trojanowski J, Wood NW, Colosimo C, Dürr A, Fowler C, Kaufmann H, Klockgether T, Lees A, Poewe W, Quinn N, Revesz T, Robertson D, Sandroni P, Seppi K, Vidailhet M (2008) Second consensus statement on the diagnosis of multiple system atrophy. Neurology 71(9):670–676
Lu CF, Soong BW, Wu HM, Teng S, Wang PS, Wu YT (2013) Disrupted cerebellar connectivity reduces whole‐brain network efficiency in multiple system atrophy. Mov Disord 28(3):362–369
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7(3):562–578. doi:10.1038/nprot.2012.016
Iyer A, Zurolo E, Prabowo A, Fluiter K, Spliet WG, van Rijen PC, Gorter JA, Aronica E (2012) MicroRNA-146a: a key regulator of astrocyte-mediated inflammatory response. PLoS One 7(9):e44789. doi:10.1371/journal.pone.0044789
Wenning GK, Tison F, Seppi K, Sampaio C, Diem A, Yekhlef F, Ghorayeb I, Ory F, Galitzky M, Scaravilli T, Bozi M, Colosimo C, Gilman S, Shults CW, Quinn NP, Rascol O, Poewe W, Multiple System Atrophy Study G (2004) Development and validation of the Unified Multiple System Atrophy Rating Scale (UMSARS). Mov Disord 19(12):1391–1402. doi:10.1002/mds.20255
Aronica E, Boer K, Becker A, Redeker S, Spliet WG, van Rijen PC, Wittink F, Breit T, Wadman WJ, Lopes da Silva FH, Troost D, Gorter JA (2008) Gene expression profile analysis of epilepsy-associated gangliogliomas. Neuroscience 151(1):272–292. doi:10.1016/j.neuroscience.2007.10.036
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 25(4):402–408. doi:10.1006/meth.2001.1262
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, Genome Project Data Processing S (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. doi:10.1093/bioinformatics/btp352
Brawand D, Soumillon M, Necsulea A, Julien P, Csardi G, Harrigan P, Weier M, Liechti A, Aximu-Petri A, Kircher M, Albert FW, Zeller U, Khaitovich P, Grutzner F, Bergmann S, Nielsen R, Paabo S, Kaessmann H (2011) The evolution of gene expression levels in mammalian organs. Nature 478(7369):343–348. doi:10.1038/nature10532
Gruber AR, Lorenz R, Bernhart SH, Neubock R, Hofacker IL (2008) The Vienna RNA websuite. Nucleic Acids Res 36(Web Server issue):W70–W74. doi:10.1093/nar/gkn188
Lorenz R, Bernhart SH, Honer Zu Siederdissen C, Tafer H, Flamm C, Stadler PF, Hofacker IL (2011) ViennaRNA package 2.0. Algorithms Mol Biol 6:26. doi:10.1186/1748-7188-6-26
Hamada M, Kiryu H, Sato K, Mituyama T, Asai K (2009) Prediction of RNA secondary structure using generalized centroid estimators. Bioinformatics 25(4):465–473. doi:10.1093/bioinformatics/btn601
Agostini F, Zanzoni A, Klus P, Marchese D, Cirillo D, Tartaglia GG (2013) catRAPID omics: a web server for large-scale prediction of protein-RNA interactions. Bioinformatics 29(22):2928–2930. doi:10.1093/bioinformatics/btt495
Bellucci M, Agostini F, Masin M, Tartaglia GG (2011) Predicting protein associations with long noncoding RNAs. Nat Methods 8(6):444–445. doi:10.1038/nmeth.1611
Kellis M, Wold B, Snyder MP, Bernstein BE, Kundaje A, Marinov GK, Ward LD, Birney E, Crawford GE, Dekker J (2014) Defining functional DNA elements in the human genome. Proc Natl Acad Sci U S A 111(17):6131–6138
Hangauer MJ, Vaughn IW, McManus MT (2013) Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS Genet 9(6):e1003569. doi:10.1371/journal.pgen.1003569
Mercer TR, Mattick JS (2013) Structure and function of long noncoding RNAs in epigenetic regulation. Nat Struct Mol Biol 20(3):300–307. doi:10.1038/nsmb.2480
Schaeffer C, Bardoni B, Mandel JL, Ehresmann B, Ehresmann C, Moine H (2001) The fragile X mental retardation protein binds specifically to its mRNA via a purine quartet motif. EMBO J 20(17):4803–4813. doi:10.1093/emboj/20.17.4803
Royce-Tolland ME, Andersen AA, Koyfman HR, Talbot DJ, Wutz A, Tonks ID, Kay GF, Panning B (2010) The A-repeat links ASF/SF2-dependent Xist RNA processing with random choice during X inactivation. Nat Struct Mol Biol 17(8):948–954. doi:10.1038/nsmb.1877
Gautier-Courteille C, Le Clainche C, Barreau C, Audic Y, Graindorge A, Maniey D, Osborne HB, Paillard L (2004) EDEN-BP-dependent post-transcriptional regulation of gene expression in Xenopus somitic segmentation. Development 131(24):6107–6117. doi:10.1242/dev.01528
Blech-Hermoni Y, Stillwagon SJ, Ladd AN (2013) Diversity and conservation of CELF1 and CELF2 RNA and protein expression patterns during embryonic development. Dev Dyn 242(6):767–777. doi:10.1002/dvdy.23959
Kress C, Gautier-Courteille C, Osborne HB, Babinet C, Paillard L (2007) Inactivation of CUG-BP1/CELF1 causes growth, viability, and spermatogenesis defects in mice. Mol Cell Biol 27(3):1146–1157. doi:10.1128/MCB. 01009-06
Campos AR, Grossman D, White K (1985) Mutant alleles at the locus elav in Drosophila melanogaster lead to nervous system defects. A developmental-genetic analysis. J Neurogenet 2(3):197–218
Chagnovich D, Fayos BE, Cohn SL (1996) Differential activity of ELAV-like RNA-binding proteins in human neuroblastoma. J Biol Chem 271(52):33587–33591
Heck MV, Azizov M, Stehning T, Walter M, Kedersha N, Auburger G (2014) Dysregulated expression of lipid storage and membrane dynamics factors in Tia1 knockout mouse nervous tissue. Neurogenetics 15(2):135–144. doi:10.1007/s10048-014-0397-x
Jin K, Li W, Nagayama T, He X, Sinor AD, Chang J, Mao X, Graham SH, Simon RP, Greenberg DA (2000) Expression of the RNA-binding protein TIAR is increased in neurons after ischemic cerebral injury. J Neurosci Res 59(6):767–774
Beck AR, Miller IJ, Anderson P, Streuli M (1998) RNA-binding protein TIAR is essential for primordial germ cell development. Proc Natl Acad Sci U S A 95(5):2331–2336
Tapia-Paez I, Tammimies K, Massinen S, Roy AL, Kere J (2008) The complex of TFII-I, PARP1, and SFPQ proteins regulates the DYX1C1 gene implicated in neuronal migration and dyslexia. FASEB J 22(8):3001–3009. doi:10.1096/fj.07-104455
Anczukow O, Rosenberg AZ, Akerman M, Das S, Zhan L, Karni R, Muthuswamy SK, Krainer AR (2012) The splicing factor SRSF1 regulates apoptosis and proliferation to promote mammary epithelial cell transformation. Nat Struct Mol Biol 19(2):220–228. doi:10.1038/nsmb.2207
Vance C, Rogelj B, Hortobagyi T, De Vos KJ, Nishimura AL, Sreedharan J, Hu X, Smith B, Ruddy D, Wright P, Ganesalingam J, Williams KL, Tripathi V, Al-Saraj S, Al-Chalabi A, Leigh PN, Blair IP, Nicholson G, de Belleroche J, Gallo JM, Miller CC, Shaw CE (2009) Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323(5918):1208–1211. doi:10.1126/science.1165942
Timchenko NA, Cai ZJ, Welm AL, Reddy S, Ashizawa T, Timchenko LT (2001) RNA CUG repeats sequester CUGBP1 and alter protein levels and activity of CUGBP1. J Biol Chem 276(11):7820–7826. doi:10.1074/jbc.M005960200
Guttman M, Garber M, Levin JZ, Donaghey J, Robinson J, Adiconis X, Fan L, Koziol MJ, Gnirke A, Nusbaum C (2010) Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat Biotechnol 28(5):503–510
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28(5):511–515
Mercer TR, Dinger ME, Sunkin SM, Mehler MF, Mattick JS (2008) Specific expression of long noncoding RNAs in the mouse brain. Proc Natl Acad Sci U S A 105(2):716–721. doi:10.1073/pnas.0706729105
Pan C, Kumar C, Bohl S, Klingmueller U, Mann M (2009) Comparative proteomic phenotyping of cell lines and primary cells to assess preservation of cell type-specific functions. Mol Cell Proteomics 8(3):443–450. doi:10.1074/mcp. M800258-MCP200
Burdall SE, Hanby AM, Lansdown MR, Speirs V (2003) Breast cancer cell lines: friend or foe? Breast Cancer Res 5(2):89–95
Peters A (2004) A fourth type of neuroglial cell in the adult central nervous system. J Neurocytol 33(3):345–357. doi:10.1023/B:NEUR.0000044195.64009.27
Ishizawa K, Komori T, Sasaki S, Arai N, Mizutani T, Hirose T (2004) Microglial activation parallels system degeneration in multiple system atrophy. J Neuropathol Exp Neurol 63(1):43–52
Stefanova N, Reindl M, Neumann M, Kahle PJ, Poewe W, Wenning GK (2007) Microglial activation mediates neurodegeneration related to oligodendroglial alpha-synucleinopathy: implications for multiple system atrophy. Mov Disord 22(15):2196–2203. doi:10.1002/mds.21671
Acknowledgments
Tissues were received from the Sydney Brain Bank at Neuroscience Research Australia and the New South Wales Tissue Resource Centre at the University of Sydney which are supported by the National Health and Medical Research Council of Australia (NHMRC), University of New South Wales, Neuroscience Research Australia, Schizophrenia Research Institute, and National Institute of Alcohol Abuse and Alcoholism (NIH (NIAAA) R24AA012725). This research was supported by the National Health & Medical Research Council of Australia (Project grant #1022325 to WSK and Fellowship #630434 to GMH) and Brain Foundation Australia (to MJ). EA is supported by the Framework Programme FP7/2007-2013 under the project EPISTOP (grant agreement n°: 602391) and (NeuroGeM grant 733051052).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Figure 1
Conservation and expression of linc00320 in mammals. For each species a brain RNA-seq read coverage track across the locus is shaded blue. Read depth scales are different between species. On the human track the linc00320 exons were numbered and highlighted in green. Expression of the last exon was only observed in human. Light blue shaded tracks (for species other than human) are BLASTN hits of each human linc00320 (ENST00000416768) exon to the respective genomes. Exons are colored differently, and the percentage of homology shared with the human linc00320 is listed after the exon names. (PDF 1585 kb)
Supplementary Figure 2
RNA secondary structure as predicted by RNAfold and region that interacts with the selected RNA-binding proteins. The color of the bases indicates how likely those bases are to pair, with red indicated maximum likely hood and blue indicating unlikely. The common 3’ exon of each isoforms results in a common secondary structure for all of the linc00320 isoforms. The area of RNA-binding protein interaction is highlighted in purple. Interestingly the RNA-binding protein interaction area comprises a similar structural motif in each isoform. The starting point of each exon is indicated with a black bar and indication of the exon number, as well as the 5’ and 3’ end of the folded transcript. A. linc00320-002 B. linc00320-006 C. linc00320-007. (GIF 5 kb)
(GIF 5 kb)
(GIF 5 kb)
High Resolution Image
(TIFF 8719 kb)
High Resolution Image
(TIFF 9322 kb)
High Resolution Image
(TIFF 8092 kb)
Supplementary Figure 3
Interaction profile of each linc00320 transcript and selected RNA-binding proteins. A. Linc00320-006 B. Linc00320-007. The x-axis indicates the nucleotide position along the transcript. The interaction score on the y-axis indicates how likely it is for the RNA-binding protein to interact with this region of the transcript. The peak of 3.5 between approximately 150–300 indicates that this is a likely area of interaction. Two experimentally validated predictions, the interaction between the Fmr1 protein and its mRNA and the interaction between Srsf1 and the long non-coding RNA Xist give interaction scores of close to 4 [30, 31]. The prediction is based on structural properties, further analysis also showed that sequence motifs are present for each of the RNA-binding proteins. (GIF 478 kb)
(GIF 472 kb)
High Resolution Image
(TIFF 6665 kb)
High Resolution Image
(TIFF 6586 kb)
Rights and permissions
About this article
Cite this article
Mills, J.D., Chen, J., Kim, W.S. et al. Long intervening non-coding RNA 00320 is human brain-specific and highly expressed in the cortical white matter. Neurogenetics 16, 201–213 (2015). https://doi.org/10.1007/s10048-015-0445-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-015-0445-1