Abstract
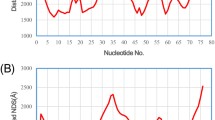
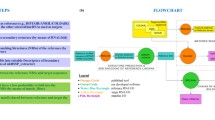
Recently, non-coding RNA which participates with organic activities has been found in non-coding region. Until now, we do not know detailed function of non-coding RNAs very well. To make clear functions of non-coding RNAs, we need to obtain a lot of data about non-coding RNAs and their targets. However, we do not have efficient techniques to analyze relations between non-coding RNAs and their targets. In this paper, we propose a high-speed method that can detect modification domain candidates on the target RNA based on the small nucleolar RNA (snoRNA) sequence. A snoRNA modifies a target RNA by composing complementary base pairs between a part of snoRNA and a part of target RNA. Our method stores the relations between snoRNA sequences and their target in a Fully Indexable Dictionary built by Trie data structure and Level-Order Unary Degree Sequence for high-speed retrieving.









Similar content being viewed by others
References
Yi-Tao Y, Rebecca MT, Michael PT (2005) Mechanisms and functions of RNA-guided RNA modification. Fine-tuning of RNA functions by modification and editing topics in current genetics, 12. pp 223–262
Junichi A (1993) Key search strategies Trie and its applications. Inf Process Soc Japan 34:244–251
Jacobson G (1989), Space-efficient static tree and graphs. Proceedings of the 30th annual symposium on foundations of computer science SFCS, pp 549–554.
Yasuo T (2012) Succinct multibit tree: compact representation of multibit trees by using succinct data structures in chemical fingerprint searches. Workshop on Algorithms in Bioinformatics Lecture Notes in Computer Science, 7534. pp 201–213
Kenmochi N (2008) snOPY (snoRNA Orthological Gene Database). http://snoopy.med.miyazaki-u.ac.jp/
Author information
Authors and Affiliations
Corresponding author
Additional information
This work was presented in part at the 19th International Symposium on Artificial Life and Robotics, Beppu, Oita, January 22–24, 2014.
About this article
Cite this article
Yamamoto, T., Yamamori, K., Kenmochi, N. et al. A detection method for snoRNA modification domain by fully indexable dictionary retrieving. Artif Life Robotics 19, 209–214 (2014). https://doi.org/10.1007/s10015-014-0149-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10015-014-0149-x