Abstract
Since the World Health Organization 2016 classification (2016 WHO), genetic status has been incorporated into the diagnosis of Grade 2/3 gliomas (lower-grade gliomas). Therefore, immunohistochemistry (IHC) of IDH1-R132H, ATRX, and p53 have been used in place of genetic status. We report the associations between histological findings, IHC, and genetic status. We performed IHC of IDH1-R132H, ATRX, and p53 in 76 lower-grade gliomas and discussed its validity based on the 2016 WHO and the upcoming 2021 WHO classification. The sensitivity and specificity of anti-ATRX, p53, and IDH1-R132H IHC were 40.9%/98.1%, 78.6%/85.4%, and 90.5%/84.6%, respectively. Among 21 IDH1-mutant gliomas without 1p/19q codeletion, two gliomas (9.5%) mimicked the so-called classic for oligodendroglioma (CFO) in their morphology. Of the 42 gliomas with 1p/19q codeletion, four cases were difficult to diagnose as oligodendroglioma through morphological examination. Moreover, there were three confusing cases with ATRX mutations but with retained ATRX-IHC positivity. The lessons learned from this study are as follows: (1) ATRX-IHC and p53-IHC should be supplementary to morphological diagnosis, (2) rare IDH mutations other than IDH1 R132H should be considered, and (3) there is no complete alternative test to detect molecular features of glioblastoma under the 2021 WHO classification.
Similar content being viewed by others
Introduction
The 2016 World Health Organization (WHO) classification of gliomas changed the way gliomas are classified [1]. Moreover, cIMPACT NOW and its updates 2–7 have been reported [2,3,4,5,6,7,8], which have added newly discovered genetic alterations found in adulthood and childhood gliomas. These situations seem to have lessened the importance of morphology-based histopathology. However, in routine practice, pathologists are making efforts to evaluate the histology of a given tumor by summarizing limited information and diagnostic tools [9]. Histological evaluations are particularly important in cases (1) requiring determination within an urgent turn-around-time period, (2) with sample insufficiency to perform accurate genetic analysis, and (3) in an institute where a molecular diagnostic technique is not available.
Molecular diagnosis in Grades 2–4 gliomas focuses mostly on the mutation of IDH1/2 and 1p/19q codeletion, although CDKN2A/B homozygous deletion, EGFR amplification, TERT promoter (pTERT) mutation, mitosis, and necrosis need to be considered for subclassification in precision treatments for gliomas [2, 3, 5, 10]. The baseline examination is the mutation of IDH1/2 and 1p/19q codeletion [11,12,13]. However, there are still controversial cases as long as we use markers, IDH1-R132H, ATRX, and p53 immunohistochemistry (IHC). In this article, we investigated how IHC-based diagnosis corresponds to the 2016 WHO and upcoming 2021 WHO classifications [14] and discussed their interpretation.
Materials and methods
Patients and data sets
We used the same tumor samples investigated in a previous study by Yamamichi et al. [15]. A total of 76 (45 male and 31 female) patient tumor tissues were used for analysis in this study, with informed consent from the patients, and ethical approval from Nagoya University. The overall mean age was 43.5 years old (Table 1). All tissues were obtained between 2003 and 2012 by surgery and diagnosed as Grades 2/3 gliomas based on the 2016 WHO classification independently by two pathologists (R.W. and I. I.). When the independent diagnosis was unmatched, a consensus diagnosis was made through discussion. The morphology with microvascular proliferations and necrosis that may be corresponded to glioblastoma (GBM), Grade 4 according to 2021 WHO was excluded in this study [14]. This study included Grades 2 and 3 gliomas together as oligodendroglioma and astrocytoma. The role of molecular diagnostics has become more weighted.
Tumor samples and analysis for molecular genetics
We examined 76 Grade 2 or 3 gliomas at Nagoya University Hospital between 2003 and 2012. The surgically removed tumor tissue was first separated into two samples for histology and molecular genetic analysis, as previously described [15,16,17]. Fresh, unfixed, tumor tissue samples were stored at – 80 °C and subsequently examined by whole-exome sequencing for IDH1/2, ATRX, TP53, pTERT, and by SNP array for 1p/19q codeletion, CDKN2A/B homozygous deletion, EGFR amplification, and chromosome 7 gain and 10 loss as was indicated in Yamamichi’s study [15].
Histology and IHC
Four slides from each case were freshly prepared from formalin-fixed paraffin-embedded tumor tissues. One slide was stained with hematoxylin and eosin (H&E), and the remaining three sections were used for immunostaining. After deparaffinization, and dipping in 0.01 M phosphate buffer saline at pH 7.2 for 10 min, the thin sections were treated by antigen retrieval method using a citrate buffer at pH 6.0 (BOND Epitope Retrieval Solution 1, AR9961, Leica Biosystems) at 100 ℃ for 30 min for the first antibody H09 (DIA-H09, Dianova, Hamburg, dilution × 200) and HMab-2 [18, 19] (dilution × 1000), or using an ethylenediaminetetraacetic acid-based buffer at pH 9.0 (BOND Epitope Solution 2, AR9640, Leica Biosystems) at 100 ℃ for 20 min for the first antibody HPA001906[19] (antibody against ATRX, Sigma-Aldrich, dilution × 200), or at 100 ℃ for 30 min for the first antibody DO-7 [20] (antibody against p53, Cell Signaling, dilution × 500). After rinsing three times for 1–3 min with Bond Wash Solution (AR9590, Leica Biosystems, Melbourne, Australia), each of the three first antibodies indicated above were applied to a thin section of each sample. The first antibodies were diluted with DAKO REAL™ Antibody Diluent (Agilent, Santa Clara, CA, USA). After the first antibodies were reacted for 15 min at room temperature, washing steps were performed and peroxidase-conjugated antibodies were applied using a Leica IHC Refine/NovoLink™ kit (Cat. DS9800), according to the manufacturer’s instructions. All 76 specimens were examined simultaneously using the same procedures by an autostainer machine (BOND-III Automated Stainer, Leica Biosystems) and a detection kit (BOND-Polymer-Refine-Detection kit, Leica Biosystems).
The sections were covered with a thin cover glass and sealing agent and microscopically examined by two pathologists (I. I. and R.W.). First, histological diagnoses were performed using only H&E sections without genetic or immunohistochemical information. Thereafter, immunohistochemical findings on each stained section were estimated in the optimally stained portion of the tumor in each section.
According to “General Rules for Clinical and Pathological Studies on Brain Tumors, Fourth Edition” (2018, in Japanese), samples were judged as IHC-positive/negative for p53, ATRX, H09, or HMab-2 as follows: when 10% or more neoplastic cells showed nuclear positivity for p53 (stronger than wild-type staining on adjacent non-neoplastic cells), it was considered IHC-positive; when neoplastic cells showed no nuclear staining in ATRX IHC in comparison with clear positivity of adjacent non-neoplastic cells, it was considered IHC-negative for ATRX; when neoplastic cells showed nuclear and cytoplasmic positivity for H09 or HMab-2, it was considered IHC-positive.
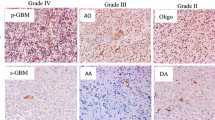
Morphologically, oligodendroglioma was diagnosed when a given case showed a typical perinuclear halo with no cytoplasmic process, namely, classic for oligodendroglioma (CFO) [21]. On the contrary, if a tumor had evident cell processes and a wide cytoplasm, the morphological diagnosis was astrocytoma. Even if a given tumor cell had some amount of cytoplasm, it was diagnosed as oligodendroglioma when small round nuclei were observed in a monotonous manner.
Results
Table 1 indicates each patient’s age, sex, IDH1/2 mutation status and H09-IHC positivity, biallelic T53 genetic status and DO-7-IHC positivity, GBM molecular features such as CDKN2A/B homozygous deletion, pTERT mutation, EGFR amplification, and both chromosome 7 gain and chromosome 10 loss, and diagnose based on only H&E staining, integrated IHC, 2016 WHO classification, or provisional diagnosis based on the upcoming 2021 WHO classification. First, we display morphologically confusing cases and demonstrate the differences between genetic testing and IHC for IDH, ATRX and TP53 status, sensitivity and specificity of ATRX and/or p53 IHC as surrogate markers for 1p/19q codeletion. Finally, we show the association of IHC-based integrated diagnosis with molecular genetic diagnosis based on the 2016 WHO and upcoming 2021 WHO classifications.
Morphology
We attempted to morphologically achieve a preliminary diagnosis using H&E sections. As shown in the column for “H&E” in Table 1, most cases were consistent with” IHC-based” and “WHO2016-based” diagnoses. However, there were some confusing cases (* C No. in Table 1). There were three oligodendrogliomas with 1p/19q codeletion, but with retained ATRX immunopositivity despite having ATRX and pTERT mutation, and the wild-type TP53 as what Yamamichi et al. have already reported [15]: two cases (Cases 64 and 71) had ATRX nonsynonymous mutation and showed many tumor cells with spindle-shaped cytoplasm and bipolar processes (Fig. 1a–d). The remaining case (Case 29) did not have a spindle-shaped cell morphology but had histology conventional to oligodendroglioma (Fig. 1e, f). Notably, case 29 had an IDH2R172K mutation.
Representative cases with discrepancies between morphology and immunohistochemistry. Three gliomas are shown which have discrepancies between the ATRX molecular analysis and IHC results. They have 1p19q codeletion, IDH mutation, and ATRX mutation (cases 64 (a, b), 71 (c, d), and 29 (e, f)). In case 64 (a) and 71 (c), the tumor cells have small oval or short spindle nuclei with narrow, long, pale, or wide eosinophilic cytoplasmic processes and almost no perinuclear halo. ATRX immunohistochemistry (IHC) shows clear nuclear positivity (b and d), despite the non-synonymous ATRX mutation. In case 29 (e), tumor cells are small, round, or oval-shaped nuclei with a perinuclear halo. This case has IDH2R172K mutation and non-synonymous ATRX mutation, whereas ATRX IHC is also positive (f). Scale bar, 20 μm
Cases 3 and 15 were genetically oligodendrogliomas, with 1p/19q codeletion, IDH1R132H mutation, and ATRX wild-type, TP53 wild-type, and pTERT mutation. However, the tumor cells of Case 3 showed small round or oval nuclei, eosinophilic cytoplasm, and bipolar processes without a perinuclear halo (Fig. 2a). The tumor cells of Case 15 had irregularly shaped nuclei, some of which showed short spindles or irregular shapes (Fig. 2b).
Four gliomas suggesting discrepancies between morphology and 1p/19q status. In Case 3 (a), the tumor cells show little or no perinuclear halo, small but irregularly shaped nuclei, and eosinophilic cytoplasmic processes such as astrocytoma. In Case 15 (b), tumor cells have cytoplasmic processes and show nuclear pleomorphism, occasionally binucleated, seemingly astrocytoma. Both tumors have IDH1R132H mutants, ATRX-wild type, and 1p/19q codeletion. In Case 4 (c), a honeycomb pattern is observed, but it has mutations in both IDH1R132H and ATRX and 1p/19q non-codeleted, that is, alterations compatible with astrocytoma. In Case 10 (d), tumor cells have small or medium-sized, round nuclei, with some having a perinuclear halo; however, this case has neither 1p/19q codeletion nor IDH mutation. Scale bar, 20 mm
Likewise, astrocytomas with typical genetic alterations, including mutated IDH1R132H, 1p/19q non-codeleted, mutated ATRX, and mutated TP53 (Cases 4 and 10), showed oligodendroglioma-like morphology; the tumor cells had irregularly shaped round nuclei with a perinuclear halo (Fig. 2c, d).
IDH1-R132H IHC
We determined the sensitivity and specificity of two worldwide anti-IDH1 R132H antibodies: H09 and HMab-2. The sensitivity and specificity of H09 for detecting IDH1 R132H were 98.2% and 84.2%, respectively, while those of HMab-2 were 89.5% and 52.6%, respectively (Table 2). H09 was more sensitive and specific than HMab-2; however, even with H09, among the 57 gliomas with genetic IDH1R132H, one (Case 72) failed to be detected by IDH1-R132H-IHC (H09). In addition, the antibody was positive in 2 out of 13 IDH1/2-wild type tumors (Cases 45 and 20, 15.4%) (Table 1). As for real-world sensitivity and specificity on H09, the number of false negative cases may include other mutations than R132H. From this standpoint, IDH IHC using H09 detected correctly 57 cases out of all kinds of IDH1/2 mutations in our cases and detected falsely 2 cases out of IDH1/2 wild type 13 cases. These data gave sensitivity of 90.5% and specificity of 84.6% on H09 IHC (not included in Table 2).
ATRX mutation
ATRX alteration status included multiple mutations, splicing, frameshift insertion/deletion, nonsynonymous, and stop-gain single nucleotide variants (Table 1). These mutational statuses did not correlate with retained or lost ATRX immunopositivity, resulting in low sensitivity for ATRX IHC (40.9%). However, most ATRX wild-type cases showed retained ATRX positivity (specificity of 98.1%) (Table 3A).
TP53 mutation
TP53 had a variety of alterations such as biallelic nonsynonymous mutation, frameshift deletion, frameshift insertion, multi-mutant/splicing, stop-gain single nucleotide variants, and splicing (Table 1). The sensitivity and specificity of p53 IHC against TP53 mutations were 78.6% and 85.4%, respectively (Table 3C).
Sensitivity and specificity of surrogate markers
Diffuse astrocytoma, IDH-mutant, is most likely to harbor mutations in ATRX and TP53; therefore, this tumor type theoretically displays loss of ARTX immunopositivity and retains p53 positivity. However, when evaluating whether they serve as a surrogate marker for 1p/19q codeletion, the sensitivity and specificity of ATRX IHC were 100% and 29.4%, respectively (Table 3B). Similarly, when we compared p53 IHC negativity with 1p/19q codeletion, the sensitivity and specificity for 1p/19q codeletion status were 90.5% and 73.5%, respectively (Table 3D). Therefore, ATRX and p53 IHC may work complimentarily as surrogate markers for 1p/19q codeletion.
When we combined “ATRX-retained and p53 wild-type” IHC results as a surrogate marker, the specificity for 1p/19q codeletion increased to 76.5%, and the sensitivity remained at 90.5% (Table 3E). Figure 3 shows the flow of IHC-based diagnosis: among 76 cases, 17 cases were H09-negative; they were categorized as diffuse astrocytoma, IDH-wild-type. In fact, there were four oligodendrogliomas, IDH-mutant with 1p/19 codeletion, and two diffuse astrocytomas, IDH-mutant, according to the 2016 WHO classification system. When ATRX was positive and p53 was negative, the tumors were diagnosed as oligodendroglioma, IDH-mutant, and 1p/19q codeletion. In our cohort, 37 patients were of this type. Genetic testing revealed that the three tumors were non-codeleted. Diffuse astrocytoma, IDH-mutant, was diagnosed when ATRX was negative, AND/OR p53 was positive. Among the 22 cases meeting these criteria, four were actually oligodendroglioma, IDH-mutant with 1p/19q codeletion, and two were diffuse astrocytoma, IDH wild-type according to the 2016 WHO scheme. Each case is shown in Table 1.
Comparison among classifications of lower-grade gliomas according to immunohistochemistry for IDH1-R132H, ATRX, and/or p53. IDH1-R132H IHC (H09)-based diagnosis leads to 17 “diffuse astrocytoma (DA), IDH-wild type” diagnoses, among which there are six genetic IDH1/2 mutant cases. ATRX and p53 IHC-based diagnosis leads to 37 “oligodendroglioma (OD), IDH-mutant, 1p/19q codeleted” and 22 “diffuse astrocytoma, IDH-mutant” diagnoses. However, there are 3 and 6 cases that differed from the genetic diagnosis, respectively
Provisional molecular diagnosis based on the upcoming 2021 WHO classification
The 2021 WHO classification is upcoming as of November, 2021. We made a provisional molecular diagnosis according to the 2021 WHO classification, as shown in Table 1. Three cases (Cases 17, 23, and 73) had pTERT mutation, and one case (Case 14) had pTERT mutation and EGFR amplification, and thus these were classified as GBM. Case 59 with wild-type IDH had a CDKN2A/B homozygous deletion. However, it was designated as astrocytoma, IDH wild-type.
Discussion
The diagnosis of lower-grade glioma has changed from a morphology-oriented system to a genetic-event-oriented system since the WHO 2016 classification. Thereby, the prognosis of the disease can be predicted more accurately based on genetic data than on morphology [15]. However, as it would be difficult to fully analyze the genetics of all cases, some immunohistochemical surrogate markers are required. The aim of this study was to investigate the reliability of surrogate markers, that is, ATRX- and p53-IHC, by evaluating the patterns of ATRX alteration and TP53 mutation in next-generation sequencing (NGS) and classical morphology. The lessons learned from this study are as follows: (1) ATRX-IHC and p53-IHC should be supplementary to morphological diagnosis, (2) the possibility of rare IDH mutations other than IDH1 R132H should be considered, and (3) there is no easy alternative test to detect molecular features of GBM under the 2021 WHO classification.
The cIMPACT-NOW Update 2 states that a definite loss of ATRX nuclear expression and/or strong, diffuse p53 immunopositivity is sufficient for the diagnosis of astrocytoma without the need for a 1p/19q test [3]. Sonoda et al. developed a practical procedure in which immunohistochemistry was used as the next step after a simple, but not recommended, diagnosis by pathology alone, and ATRX and p53 IHC could be used as a surrogate for 1p/19q [22]. Our data support these reports by additionally providing the feasibility and reliability of combined ATRX and p53 IHCs in clinical practice.
Few studies have compared potential IHC surrogates and genetic analysis in Grade 2/3 gliomas, while some reports have suggested that the presence of ATRX mutations could be a surrogate marker for 1p/19q codeletion [23,24,25]. These studies limited the utility of immunohistochemistry to provide genetic information on tumors. Hewer et al. stated that ATRX-proficient tumors with IDH1 IHC positivity predicted 1p/19q loss of heterozygosity with a sensitivity of 85% [23]. Rajeswarie et al. classified gliomas according to their immunohistochemical surrogate markers. However, their studies did not provide a genetic status using NGS [26].
There seems to be an incomplete parallel relationship between mutational status and IHC results for ATRX, IDH, and TP53. The IHC results may be susceptible to the examination conditions. It is difficult to set the optimal conditions for IHC staining and to estimate the results. Therefore, the sensitivity and specificity may be affected by inter-laboratory and inter-condition variations. In our study, the cohort was collected from a single institute and analyzed retrospectively. When archived paraffin-embedded tissues are used, one must consider whether the antigenicity is well-preserved [27, 28].
We demonstrated that the specificity and sensitivity for 1p/19q codeletion were higher (76.5%/90.5%) when combined with ATRX-IHC retention and p53-IHC wild pattern (Table 3E). Many sought other surrogates; Rajmohan et al. reported that alpha internexin is a surrogate marker for 1p/19q codeletion, with a specificity of 93.0% and a sensitivity of 87.5% [29]. Kitahama et al. reported that H3K27me3 loss in IHC tended to be a 1p/19q surrogate with high specificity but low sensitivity that was not observed in oligodendrogliomas with IDH mutations other than IDH1-R132H [30]. However, the underlying mode of action remains unknown in these studies and they provide only circumstantial evidence.
The pathological diagnosis began with H&E-based morphological observations, which correctly determined 38 cases of 42 genetic oligodendrogliomas and 26 cases of 34 genetic astrocytomas. However, there are a substantial number of cases that could not be diagnosed correctly using only H&E-based morphological examination, as depicted in Figs. 1 and 2. Integrated with IHC and morphology, the correct diagnosis could be achieved in most cases.
IHC- and morphology-based diagnosis has, however, a limitation. In our cohot, 63 IDH-mutant gliomas harbored six IDH1/2 mutations other than IDH1-R132H. It would be hazardous to determine the IDH mutation status using only IDH1-R132H IHC. Regardless of the tumor grading, IDH-wild type astrocytoma can be classified as a GBM-like entity; thus, patients require intensive radiochemotherapy. This study suggested that there were 15.4% false positives and 9.5% false negatives for IDH1/2 mutations. This error rate would raise a new concern that patients may receive over- or under-treatment when 2021 WHO is applied.
References
Louis DN, Perry A, Reifenberger G et al (2016) The 2016 World health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
Louis DN, Wesseling P, Paulus W et al (2018) cIMPACT-NOW update 1: not otherwise specified (NOS) and not elsewhere classified (NEC). Acta Neuropathol 135:481–484
Louis DN, Giannini C, Capper D et al (2018) cIMPACT-NOW update 2: diagnostic clarifications for diffuse midline glioma, H3 K27M-mutant and diffuse astrocytoma/anaplastic astrocytoma, IDH-mutant. Acta Neuropathol 135:639–642
Brat DJ, Aldape K, Colman H et al (2018) cIMPACT-NOW update 3: recommended diagnostic criteria for “Diffuse astrocytic glioma, IDH-wildtype, with molecular features of glioblastoma, WHO grade IV.” Acta Neuropathol 136:805–810
Ellison DW, Hawkins C, Jones DTW et al (2019) cIMPACT-NOW update 4: diffuse gliomas characterized by MYB, MYBL1, or FGFR1 alterations or BRAF(V600E) mutation. Acta Neuropathol 137:683–687
Brat DJ, Aldape K, Colman H et al (2020) cIMPACT-NOW update 5: recommended grading criteria and terminologies for IDH-mutant astrocytomas. Acta Neuropathol 139:603–608
Louis DN, Wesseling P, Aldape K et al (2020) cIMPACT-NOW update 6: new entity and diagnostic principle recommendations of the cIMPACT-Utrecht meeting on future CNS tumor classification and grading. Brain Pathol 30:844–856
Ellison DW, Aldape KD, Capper D et al (2020) cIMPACT-NOW update 7: advancing the molecular classification of ependymal tumors. Brain Pathol 30:863–866
Paulus W (2017) WHO 2016: Open questions and practical implications. Acta Neurochir (Wien) 159:419–422
Pekmezci M, Rice T, Molinaro AM et al (2017) Adult infiltrating gliomas with WHO 2016 integrated diagnosis: additional prognostic roles of ATRX and TERT. Acta Neuropathol 133:1001–1016
Gorovets D, Kannan K, Shen R et al (2012) IDH mutation and neuroglial developmental features define clinically distinct subclasses of lower grade diffuse astrocytic glioma. Clin Cancer Res 18:2490–2501
Kefenie H, Desta B, Abebe A et al (1989) Prevalence of hepatitis B infection among hospital personnel in Addis Ababa (Ethiopia). Eur J Epidemiol 5:462–467
Cairncross G, Wang M, Shaw E et al (2013) Phase III trial of chemoradiotherapy for anaplastic oligodendroglioma: long-term results of RTOG 9402. J Clin Oncol 31:337–343
Louis DN, Perry A, Wesseling P et al (2021) The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol 23:1231–1251
Yamamichi A, Ohka F, Aoki K et al (2018) Immunohistochemical ATRX expression is not a surrogate for 1p19q codeletion. Brain Tumor Pathol 35:106–113
Suzuki H, Aoki K, Chiba K et al (2015) Mutational landscape and clonal architecture in grade II and III gliomas. Nat Genet 47:458–468
Aoki K, Natsume A (2019) Overview of DNA methylation in adult diffuse gliomas. Brain Tumor Pathol 36:84–91
Kato Y (2015) Specific monoclonal antibodies against IDH1/2 mutations as diagnostic tools for gliomas. Brain Tumor Pathol 32:3–11
Fujii Y, Ogasawara S, Oki H et al (2015) A high-sensitive HMab-2 specifically detects IDH1-R132H, the most common IDH mutation in gliomas. Biochem Biophys Res Commun 466:733–739
Takami H, Yoshida A, Fukushima S et al (2015) Revisiting TP53 mutations and immunohistochemistry–a comparative study in 157 diffuse gliomas. Brain Pathol 25:256–265
Giannini C, Burger PC, Berkey BA et al (2008) Anaplastic oligodendroglial tumors: refining the correlation among histopathology, 1p 19q deletion and clinical outcome in Intergroup Radiation Therapy Oncology Group Trial 9402. Brain Pathol 18:360–369
Sonoda Y, Yokoo H, Tanaka S et al (2019) Practical procedures for the integrated diagnosis of astrocytic and oligodendroglial tumors. Brain Tumor Pathol 36:56–62
Hewer E, Vajtai I, Dettmer MS et al (2016) Combined ATRX/IDH1 immunohistochemistry predicts genotype of oligoastrocytomas. Histopathology 68:272–278
Takano S, Ishikawa E, Sakamoto N et al (2016) Immunohistochemistry on IDH 1/2, ATRX, p53 and Ki-67 substitute molecular genetic testing and predict patient prognosis in grade III adult diffuse gliomas. Brain Tumor Pathol 33:107–116
Leeper HE, Caron AA, Decker PA et al (2015) IDH mutation, 1p19q codeletion and ATRX loss in WHO grade II gliomas. Oncotarget 6:30295–30305
Rajeswarie RT, Rao S, Nandeesh BN et al (2018) A simple algorithmic approach using histology and immunohistochemistry for the current classification of adult diffuse glioma in a resource-limited set-up. J Clin Pathol 71:323–329
Grillo F, Bruzzone M, Pigozzi S et al (2017) Immunohistochemistry on old archival paraffin blocks: is there an expiry date? J Clin Pathol 70:988–993
Combs SE, Han G, Mani N et al (2016) Loss of antigenicity with tissue age in breast cancer. Lab Invest 96:264–269
Rajmohan KS, Sugur HS, Shwetha SD et al (2020) Alpha internexin: a surrogate marker for 1p/19q codeletion and prognostic marker in anaplastic (WHO grade III) Gliomas. Neurol India 68:832–837
Kitahama K, Iijima S, Sumiishi A et al (2021) Reduced H3K27me3 levels in diffuse gliomas: association with 1p/19q codeletion and difference from H3K27me3 loss in malignant peripheral nerve sheath tumors. Brain Tumor Pathol 38:23–29
Acknowledgements
This research was supported by a Grant-in-Aid for Scientific Research on Innovative Areas “Chemistry for Multimolecular Crowding Biosystems” (A.N.) (JSPS KAKENHI Grant No. 17928985). This research was also supported in part by the Japan Agency for Medical Research and Development (AMED) under Grant Number: JP21am0101078 (to Y.K.).
Funding
Japan Society for the Promotion of Science,17928985,Atsushi Natsume,Japan Agency for Medical Research and Development,JP21am0101078,Yukinari Kato
Author information
Authors and Affiliations
Contributions
Conceptualization: A.N., R.W. and I.I.; Methodology: A.N., A.Y., K.M., Y. Kitano, R.W., and I.I.; Conduct of Immunohistochemistry: S.S. and T.O.; Formal analysis and investigation: Y. Kitano, A.Y. S.M., Y.K., A.K., R.S., T. W., R.W., and I.I.; Writing-original draft preparation: T.N., R.W., and I.I.; Writing-review and editing: R.W., I.I., and A.N.; Funding acquisition: A.N. and Y. Kato; Resources: Y.M., F.O., K.A., M.M., J.Y., Y. Kitano and Y. Kato; Supervision: A.N. and I.I.
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nishikawa, T., Watanabe, R., Kitano, Y. et al. Reliability of IDH1-R132H and ATRX and/or p53 immunohistochemistry for molecular subclassification of Grade 2/3 gliomas. Brain Tumor Pathol 39, 14–24 (2022). https://doi.org/10.1007/s10014-021-00418-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10014-021-00418-x