Abstract
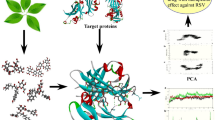
The discovery of antiviral approaches to prevent or cure respiratory syncytial virus (RSV) infections is critical, particularly because RSV is one of the most common causes of infant respiratory problems. There is currently no approved vaccination available to treat RSV infections. FDA has approved the drug ribavirin, but it is not sufficient to treat RSV. This work aimed to find and study in silico anti-RSV drugs that target matrix protein and nucleoprotein. In this study, we have identified five drug candidates that had better binding energies than ribavirin. Garenoxacin appeared as top lead compounds between them. AutoDock Vina was used to execute molecular docking of a library of chosen chemicals. The high-score compound was then confirmed using the Maestro 12.3 module’s molecular dynamics simulation and the binding energies derived using Prime/Molecular Mechanics Generalized Born Surface Area (Prime/MM-GBSA). Comparative molecular dynamics simulations revealed that garenoxacin has better stability and high residue contacts with high binding affinity than ribavirin. This study showed garenoxacin could prevent RSV infection better than ribavirin. In pursuing a more effective RSV control drug, additional research into these chemicals in vitro and in vivo is essential.











Similar content being viewed by others
References
Heylen E, Neyts J, Jochmans D (2017) Drug candidates and model systems in respiratory syncytial virus antiviral drug discovery. Biochem Pharmacol 127:1–12
Cancellieri M, Bassetto M, Widjaja I et al (2015) In silico structure-based design and synthesis of novel anti-RSV compounds. Antiviral Res 122:46–50
Bloom-Feshbach K, Alonso WJ, Charu V et al (2013) Latitudinal variations in seasonal activity of influenza and respiratory syncytial virus (RSV): a global comparative review. PloS one 8:e54445
Hall CB, Weinberg GA, Iwane MK et al (2009) The burden of respiratory syncytial virus infection in young children. N Engl J Med 360:588–598
Meerhoff TJ, Paget JW, Kimpen JL et al (2009) Variation of respiratory syncytial virus and the relation with meteorological factors in different winter seasons. Pediatr Infect Dis J 28:860–866
Tan L, Coenjaerts FE, Houspie L et al (2013) The comparative genomics of human respiratory syncytial virus subgroups A and B: genetic variability and molecular evolutionary dynamics. J Virol 87:8213–8226
Eckardt-Michel J, Lorek M, Baxmann D et al (2008) The fusion protein of respiratory syncytial virus triggers p53-dependent apoptosis. J Virol 82:3236–3249
Saravolatz LD, Empey KM, Peebles Jr RS et al (2010) Pharmacologic advances in the treatment and prevention of respiratory syncytial virus. Clin Infect Dis 50:1258–1267
Piras S, Sanna G, Carta A et al (2019) Dichloro-phenyl-benzotriazoles: a new selective class of human respiratory syncytial virus entry inhibitors. Front Chem 7:247
Cox RM, Toots M, Yoon JJ et al (2018) Development of an allosteric inhibitor class blocking RNA elongation by the respiratory syncytial virus polymerase complex. J Biol Chemm 293:16761–16777
Cichero E, Calautti A, Francesconi V et al (2021) Probing in silico the benzimidazole privileged scaffold for the development of drug-like anti-RSV agents. Pharmaceuticals 14:1307
Kim S, Thiessen PA, Bolton EE et al (2016) PubChem substance and compound databases. Nucleic acids Res 44:D1202–D1213
Dong J, Cao DS, Miao HY et al (2015) ChemDes: an integrated web-based platform for molecular descriptor and fingerprint computation. J Cheminform 7:1–10
Kenny PW (2019) The nature of ligand efficiency. J Cheminform 11:1–18
Burley SK, Berman HM, Kleywegt GJ et al (2017) Protein Data Bank (PDB): the single global macromolecular structure archive. Prot Crystallo:627–641
Costello HM, Ray WC, Chaiwatpongsakorn S et al (2012) Targeting RSV with vaccines and small molecule drugs. Infect Disord Drug Target 12:110–128
Bharaj P, Wang YE, Dawes BE et al (2016) The matrix protein of Nipah virus targets the E3-ubiquitin ligase TRIM6 to inhibit the IKKε kinase-mediated type-I IFN antiviral response. PLoS pathogens 12:e1005880
Zhang W, Pei J, Lai L (2017) Computational multitarget drug design. J Chem Inf Model 57:403–412
Kumar A, Tiwari A, Sharma A (2018) Changing paradigm from one target one ligand towards multi-target directed ligand design for key drug targets of Alzheimer disease: an important role of in silico methods in multi-target directed ligands design. Curr Neuropharmacol 16:726–739
Tian W, Chen C, Lei X et al (2018) CASTp 3.0: computed atlas of surface topography of proteins. Nucleic acids Res 46:W363–W367
Forli W, Halliday S, Belew R et al (2012) AutoDock Version 4.2. J Med Chem 55:623–638
Mitra D, Dey A, Biswas I et al (2021) Bioactive compounds as a potential inhibitor of colorectal cancer; an insilico study of gallic acid and pyrogallol. Anna Colorec Res 9:32–39
Mitra D, Mohapatra PK (2022) Effect of natural compounds to inhibit human respiratory syncytial virus. Smart Environmental Science, Technology and Management 97–101
Mitra D, Paul M, Thatoi H et al (2021) Study of potentiality of dexamethasone and its derivatives against Covid-19. J Biomol Struc Dyna:1–11
Khanna V, Ranganathan S (2009) Physicochemical property space distribution among human metabolites, drugs and toxins. In BMC bioinformatics 10:1–18
Mitra D, Mohapatra PKD (2021) Inhibition of SARS-CoV-2 protein by bioactive compounds of edible mushroom; a bioinformatics insight. Int J Adv Res Sci Eng Technol 9:84–88
DeLano WL (2002) PyMOL: an open-source molecular graphics tool. CCP4 Newsletter on protein crystallography 40:82-92.
Ononamadu CJ, Abdalla M, Ihegboro GO et al (2021) In silico identification and study of potential anti-mosquito juvenile hormone binding protein (MJHBP) compounds as candidates for dengue virus-vector insecticides. Biochem Biophys Rep 28:101178
Baby K, Maity S, Mehta CH et al (2020) Targeting SARS-CoV-2 RNA-dependent RNA polymerase: an in silico drug repurposing for COVID-19. F1000Research 9:1166.
Jin Z, Wang Y, Yu XF et al (2020) Structure-based virtual screening of influenza virus RNA polymerase inhibitors from natural compounds: molecular dynamics simulation and MM-GBSA calculation. Comput Biol Chem 85:107241
Di L, Kerns EH, Carter GT (2009) Drug-like property concepts in pharmaceutical design. Curr Pharm Des 15:2184–2194
Lipinski CA (2004) Lead-and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341
Benet LZ, Hosey CM, Ursu O et al (2016) BDDCS, the rule of 5 and drugability. Adv Drug Deliv Rev 101:89–98
Porter CJ, Trevaskis NL, Charman WN (2007) Lipids and lipid-based formulations: optimizing the oral delivery of lipophilic drugs. Nat Rev Drug Discov 6:231–248
Lobo S (2020) Is there enough focus on lipophilicity in drug discovery? Expert Opin Drug Discov 15:261–263
Clark DE (2011) What has polar surface area ever done for drug discovery? Future Med Chem 3:469–484
Hitchcock SA, Pennington LD (2006) Structure− brain exposure relationships. J Med Chem 49:7559–7783
Lucas MC, Bhagirath N, Chiao E et al (2014) Using ovality to predict nonmutagenic, orally efficacious pyridazine amides as cell specific spleen tyrosine kinase inhibitors. J Med Chem 57:2683–2691
Stank A, Kokh DB, Fuller JC et al (2016) Protein binding pocket dynamics. Acco Chem Res 49:809–815
Guedes IA, de Magalhães CS, Dardenne LE (2014) Receptor–ligand molecular docking. Biophy Rev 6:75–87
Shah P, Westwell AD (2007) The role of fluorine in medicinal chemistry. J Enzym Inhib Med Chem 22:527–540
Cheng T, Li Q, Zhou Z, Wang Y et al (2012) Structure-based virtual screening for drug discovery: a problem-centric review. The AAPS J 14:133–141
Sarkar C, Abdalla M, Mondal M et al (2021) Ebselen suitably interacts with the potential SARS-CoV-2 targets: an in-silico approach. J Biomol Struc Dyna 23:1–16
Beura S, Chetti P (2021) In-silico strategies for probing chloroquine based inhibitors against SARS-CoV-2. J Biomol Struc Dyna 39:3747–3759
Ahmed S, Mahtarin R, Ahmed SS et al (2021) Investigating the binding affinity, interaction, and structure-activity-relationship of 76 prescription antiviral drugs targeting RdRp and Mpro of SARS-CoV-2. J Biomol Struc Dyna 39:6290–6305
Availability of data and material
NA
Author information
Authors and Affiliations
Contributions
D.M. conceived and designed the project. P.K.D.M. conducted initial manual verifications. Target protein and ligand compounds were identified by D.M. Molecular docking was performed by D.M. Molecular dynamic simulations were performed by S.A and M.A. Analysis of those results was done by M.A. Draft of the manuscript was prepared by D.M and S.A. Final version of the manuscript was edited by M.A. The whole work was done under the supervision of P.K.D.M and M.A.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mitra, D., Afreen, S., Das Mohapatra, P.K. et al. Threat of respiratory syncytial virus infection knocking the door: a proposed potential drug candidate through molecular dynamics simulations, a future alternative. J Mol Model 29, 91 (2023). https://doi.org/10.1007/s00894-023-05489-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-023-05489-5