Abstract
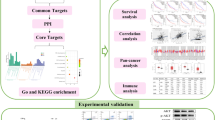
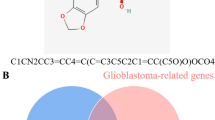
Based on molecular docking and molecular dynamics simulation, to find a new target and mechanism of MEK inhibitor Selumetinib in the treatment of low-grade glioma (LGG), and to provide theoretical guidance for its clinical medication. All possible targets of Selumetinib were fished through the compound target prediction database. New targets of Selumetinib in the treatment of LGG were found and its mechanism was evaluated employing molecular docking, gene difference analysis, molecular dynamics simulation, and protein subcellular localization prediction. A total of 100 Selumetinib targets and 85 LGG-related targets were screened in this study. There were 7 active targets at the intersection of the two. Through protein interaction (PPI), gene enrichment analysis, and gene difference analysis, one effective target of Selumetinib was finally screened, CDK2 mainly existing in the cytoplasm, endoplasmic reticulum, and plasma membrane; the target plays a role in the treatment of LGG by inhibiting the signal pathways of PI3K Akt and participating in biological processes such as peptide amino acid modification, regulation of intracellular signal transduction, and positive regulation of cell metabolism. CDK2 may be a new direction of Selumetinib in the clinical treatment of LGG.








Similar content being viewed by others
Data Availability
The datasets generated and/or analyzed during the current study are not publicly available due the limited scope of data availability; these data are used under the license of this study and are not disclosed, but are available from the author (sedate@stu.shzu.edu.cn (Dongdong Zhang)) on reasonable request.
We confirm that all methods in the manuscript are carried out in accordance with the relevant guidelines.
Abbreviations
- GBM:
-
Glioblastoma
- LGG:
-
Low-grade glioma
- PD-1:
-
Programmed cell death-1
- PDL-1:
-
Programmed cell death-ligand 1
- NF1:
-
Neurofibromas 1
- PN:
-
Plexiform neurofibroma
- BP:
-
Biology progress
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PPI:
-
Protein-protein interaction
- MD:
-
Molecule dynamics
- RMSD:
-
Root mean squared error
- RMSF:
-
Root mean square float
- GDA:
-
Human Genetic Disease Association
- k-NN:
-
K-nearest neighbor
References
Zhang Q, Xiang W, Yi DY, Xue BZ, Wen WW, Abdelmaksoud A et al (2018) Current status and potential challenges of mesenchymal stem cell-based therapy for malignant gliomas. Stem Cell Res Ther 9(1):228
Zhao W, Lin J (2021) Nan fang yi ke da xue xue bao = Journal of Southern Medical University 41(8):1171–1176
Yang Z, Yang Z, Hu Z, Li B, Liu D, Chen X et al (2021) UAP1L1 plays an oncogene-like role in glioma through promoting proliferation and inhibiting apoptosis. Ann Transl Med 9(7):542
Jänne PA, van den Heuvel MM, Barlesi F, Cobo M, Mazieres J, Crinò L et al (2017) Selumetinib plus docetaxel compared with docetaxel alone and progression-free survival in patients with KRAS-mutant advanced non-small cell lung cancer: the SELECT-1 randomized clinical trial. JAMA 317(18):1844–1853
Oxnard GR, Yang JC, Yu H, Kim SW, Saka H, Horn L et al (2020) TATTON: a multi-arm, phase Ib trial of osimertinib combined with selumetinib, savolitinib, or durvalumab in EGFR-mutant lung cancer. Ann Oncol 31(4):507–516
Daina A, Michielin O, Zoete V (2019) SwissTargetPrediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res 47(W1):W357–W364
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y et al (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Tang Z, Kang B, Li C, Chen T, Zhang Z (2019) GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res 47(W1):W556–W560
Orre LM, Vesterlund M, Pan Y, Arslan T, Zhu Y, Fernandez Woodbridge A et al (2019) SubCellBarCode: proteome-wide mapping of protein localization and relocalization. Mol Cell 73(1):166-182.e7
Ji JW, Zhang YD, Lai YJ, Huang CG (2020) Mettl3 regulates the proliferation, migration and invasion of glioma cells by inhibiting PI3K/Akt signaling pathway. Eur Rev Med Pharmacol Sci 24(7):3818–3828
Jiang L, Wang C, Lei F, Zhang L, Zhang X, Liu A et al (2015) miR-93 promotes cell proliferation in gliomas through activation of PI3K/Akt signaling pathway. Oncotarget 6(10):8286–8299
Funding
“Shihezi University high-level talent scientific research start-up project” prevention and control of recombinant endophytes
“Technical research on” branch blight “of fragrant pear,” Project No.: RCZK202047
Author information
Authors and Affiliations
Contributions
Dongdong Zhang and Tieying Zhang contributed equally to this research and Dongdong Zhang and Tieying Zhang wrote the main manuscript text. Jin Li and Jianbo Zhu are the corresponding authors. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, D., Zhang, T., Zhu, J. et al. Discovery of novel targets and mechanisms of MEK inhibitor Selumetinib for LGG treatment based on molecular docking and molecular dynamics simulation. J Mol Model 28, 138 (2022). https://doi.org/10.1007/s00894-022-05132-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-022-05132-9