Abstract
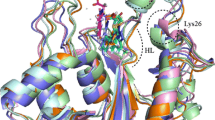
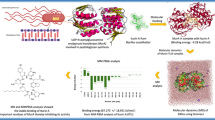
Leprosy is an infectious disease caused by Mycobacterium leprae. The increasing drug and multi-drug resistance of M. leprae enforce the importance of finding new drug targets. Mycobacterium has unusually impermeable cell wall that contributes to considerable resistance to many drugs. Peptidoglycan is an important component of the cell wall of M. leprae. UDP-N-acetylmuramoyl-glycyl-D-glutamate-2, 6-diaminopimelate ligase (MurE) plays a crucial role in the peptidoglycan biosynthesis and hence it could be considered as a potential drug target for leprosy. Structure of this enzyme for M. leprae has not yet been elucidated. We modeled the three-dimensional structure of MurE from M. leprae using comparative modeling methods based on the X-ray crystal structure of MurE from E. coli and validated. The 3D-structure of M. leprae MurE enzyme was docked with its substrates meso-diaminopimelic acid (A2pm) and UDP-N-acetyl muramoyl-glycyl-D- glutamate (UMGG) and its product UDP-N-acetyl muramoyl-glycyl-D-glu-meso-A2pm (UTP) and also with ATP. The docked complexes reveal the amino acids responsible for binding the substrates. Superposition of these complex structures suggests that carboxylic acid group of UMGG is positioned in proximity to γ-phosphate of the ATP to facilitate the formation of acylphosphate intermediate. The orientation of an amino group of A2pm facilitates the nucleophilic attack to form the product. Overall, the proposed model together with its binding features gained from docking studies could help to design a truly selective ligand inhibitor specific to MurE for the treatment of leprosy.









Similar content being viewed by others
References
World Health Organization (2000) Leprosy-global situation. Wkly Epidemiol Rec 75:226–231
Brosch R, Gordon SV, Eiglmeier K, Garnier T, Cole ST (2000) Comparative genomics of leprosy and tubercle bacilli. Res Microbiol 151:135–142
Cole ST, Eiglmeier K, Parkhill J et al. (2001) Massive gene decay in the leprosy bacillus. Nature 409:1007–1011
Levy L, Shepard CC, Fasal P (1976) The bactericidal effect of rifampicin on M. leprae in man: a) single doses of 600, 900 and 1200 mg; and b) daily doses of 300 mg. Int J Lepr Other Mycobact Dis 44:183–187
Chemotherapy of leprosy. Report of a WHO study group. World Health Organ Tech Rep Ser 847:1–24 (1994)
Norman G, Joseph G, Ebenezer G, Rao SP, Job CK (2003) Secondary rifampin resistance following multi-drug therapy—a case report. Int J Lepr Other Mycobact Dis 71:18–21
Guelpa-Lauras CC, Grosset JH, Constant-Desportes M, Brucker G (1984) Nine cases of rifampin-resistant leprosy. Int J Lepr Other Mycobact Dis 52:101–102
Ji BH (1985) Drug resistance in leprosy—a review. Lepr Rev 56:265–278
Ji B (2002) Rifampin-resistant leprosy: a review and a research proposal of a pilot study. Lepr Rev 73:2–8
Matsuoka M, Suzuki Y, Garcia IE, Fafutis-Morris M, Vargas-González A, Carreño-Martinez C, Fukushima Y, Nakajima C (2010) Possible mode of emergence for drug-resistant leprosy is revealed by an analysis of samples from Mexico. Jpn J Infect Dis 63:412–416
Brennan PJ (2003) Structure, function and biogenesis of the cell wall of Mycobacterium tuberculosis. Tuberculosis (Edinb) 83:91–97
Draper P, Kandler O, Darbre A (1987) Peptidoglycan and arabinogalactan of Mycobacterium leprae. J Gen Microbiol 133:1187–1194
Mahapatra S, Crick DC, Brennan PJ (2000) Comparison of the UDP-N-acetylmuramate: L-alanine ligase enzymes from Mycobacterium tuberculosis and Mycobacterium leprae. J Bacteriol 182:6827–6830
Shanmugam A, Natarajan J (2010) Computational genome analyses of metabolic enzymes in Mycobacterium leprae for drug target identification. Bioinformation 4:392–395
Gordon E, Flouret B, Chantalat L, van Heijenoort J, Mengin-Lecreulx D, Dideberg O (2001) Crystal structure of UDP-N-acetylmuramoyl-L-alanyl-D-glutamate: meso-diaminopimelate ligase from Escherichia coli. J Biol Chem 276:10999–11006
Schleifer KH, Kandler O (1972) Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol Rev 36:407–477
Gowthaman R, Silvester AJ, Saranya K, Kanya KS, Archana NR (2006) Modeling of the potential coiled-coil structure of snapin protein and its interaction with SNARE complex. Bioinformation 1:269–275
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bernstein FC, Koetzle TF, Williams GJ, Meyer EF Jr, Brice MD, Rogers JR, Kennard O, Shimanouchi T, Tasumi M (1978) The Protein Data Bank: a computer-based archival file for macromolecular structures. Arch Biochem Biophys 185:584–591
Kelley LA, Sternberg MJ (2009) Protein structure prediction on the web: a case study using the Phyre server. Nat Protoc 4:363–371
Jones DT, Taylor WR, Thornton JM (1992) A new approach to protein fold recognition. Nature 358:86–89
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) ClustalW and ClustalX version 2.0. Bioinformatics 23:2947–2948
Sali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234:779–815
Qadri YJ, Berdiev BK, Song Y, Lippton HL, Fuller CM, Benos DJ (2009) Psalmotoxin-1 docking to human acid-sensing ion channel-1. J Biol Chem 284:17625–17633
Maiti R, Van Domselaar GH, Zhang H, Wishart DS (2004) SuperPose: a simple server for sophisticated structural superposition. Nucleic Acids Res 32 (Web Server issue):W590-594
ACD/ChemSketch Freeware, version 10.00, Advanced Chemistry Development, Inc, Toronto, ON, Canada, www.acdlabs.com, 2006
Weininger D (1988) SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J Chem Inf Comput Sci 28:31–36
Wishart DS, Knox C, Guo AC, Shrivastava S, Hassanali M, Stothard P, Chang Z, Woolsey J (2006) DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res 34 (Database issue): D668-672
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, Olson AJ (1998) Automated docking using a Lamarckian Genetic algorithm and empirical binding free energy function. J Comput Chem 19:1639–1662
Rayan A (2009) New tips for structure prediction by comparative modeling. Bioinformation 3:263–267
Basavannacharya C, Robertson G, Munshi T, Keep NH, Bhakta S (2010) ATP-dependent MurE ligase in Mycobacterium tuberculosis: biochemical and structural characterization. Tuberculosis (Edinb) 90:16–24
Bertrand JA, Auger G, Martin L, Fanchon E, Blanot D, Le Beller D, van Heijenoort J, Dideberg O (1999) Determination of the MurD mechanism through crystallographic analysis of enzyme complexes. J Mol Biol 289:579–590
Basavannacharya C, Moody PR, Munshi T, Cronin N, Keep NH, Bhakta S (2010) Essential residues for the enzyme activity of ATP-dependent MurE ligase from Mycobacterium tuberculosis. Protein Cell 1:1011–1022
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shanmugam, A., Natarajan, J. Comparative modeling of UDP-N-acetylmuramoyl-glycyl-D-glutamate-2, 6-diaminopimelate ligase from Mycobacterium leprae and analysis of its binding features through molecular docking studies. J Mol Model 18, 115–125 (2012). https://doi.org/10.1007/s00894-011-1039-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-011-1039-y