Abstract
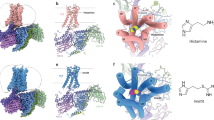
It is widely acknowledged that the H1 receptor antagonists have important therapeutic significance in the treatment of various allergic disorders, but little was known about the binding mode between the receptor and antagonists since the crystal structure of G-protein coupling receptors (GPCRs) were hard to obtain. In this paper, a theoretical three-dimensional model of human histamine H1 receptor (HHR1) was developed on the basis of recently reported high resolution structures of human A2A adenosine receptor, human β2-adrenoceptor and turkey β1-adrenoceptor. Furthermore, three representative H1 receptor antagonists were chosen for docking studies. Subsequently, a qualitative pharmacophore model was developed by Hiphop algorithm based on the docking conformations of these three antagonists. In this paper, active environment, certain key residues, and the corresponding pharmacophore features of H1 receptor were identified by such combinations of receptor-based and ligand-based approaches, which would give sufficient guidance for the rational design of novel antihistamine agents.

Active environment, certain key residues and the corresponding pharmacophore features of H1 receptor were identified by such a combination of receptor-based and ligand-based approaches, which would give sufficient guidance for the rational design of novel antihistamine agents.










Similar content being viewed by others
References
Hill SJ (1990) Pharmacol Rev 42:45–83
Hill SJ, Ganellin CR, Timmerman H, Schwartz JC, Shankley NP, Young JM, Schunack W, Levi R, Haas HL (1997) Pharmacol Rev 49:253–278
Fukui H, Fujimoto K, Mizuguchi H, Sakamoto K, Horio Y, Takai S, Yamada K, Ito S (1994) Biochem Biophys Res Commun 201:894–901
Gantz I, Schaffer M, DelValle J, Logsdon C, Campbell V, Uhler M, Yamada T (1991) Proc Natl Acad Sci USA 88:429–433
Lovenberg TW, Roland BL, Wilson SJ, Jiang X, Pyati J, Huvar A, Jackson MR, Erlander MG (1999) Mol Pharmacol 55:1101–1107
Zhu Y, Michalovich D, Wu H, Tan KB, Dytko GM, Mannan IJ, Boyce R, Alston J, Tierney LA, Li X, Herrity NC, Vawter L, Sarau HM, Ames RS, Davenport CM, Hieble JP, Wilson S, Bergsma DJ, Fitzgerald LR (2001) Mol Pharmacol 59:434–441
Ohta K, Hayashi H, Mizuguchi H, Kagamiyama H, Fujimoto K, Fukui H (1994) Biochem Biophys Res Commun 203:1096–1101
Nonaka H, Otaki S, Ohshima E, Kono M, Kase H, Ohta K, Fukui H, Ichimura M (1998) Eur J Pharmacol 345:111–117
Wieland K, Laak AM, Smit MJ, Kuhne R, Timmerman H, Leurs R (1999) J Biol Chem 274:29994–30000
Gillard M, Van Der Perren C, Moguilevsky N, Massingham R, Chatelain P (2002) Mol Pharmacol 61:391–399
Leurs R, Smit MJ, Timmerman H (1995) Pharmacol Ther 66:413–463
Delgado LF, Pferferman A, Sole D, Naspitz CK (1998) Ann Allergy Asthma Immunol 80:333–337
Ter Laak AM, Timmerman H, Leurs R, Nederkoorn PH, Smit MJ, Donne-Op den Kelder GM (1995) J Comput Aided Mol Des 9:319–330
Kiss R, Kovari Z, Keseru GM (2004) Eur J Med Chem 39:959–967
Bruysters M, Jongejan A, Akdemir A, Bakker RA, Leurs R (2005) J Biol Chem 280:34741–34746
Strasser A, Wittmann HJ (2007) J Comput Aided Mol Des 21:499–509
Palczewski K, Kumasaka T, Hori T, Behnke CA, Motoshima H, Fox BA, Le Trong I, Teller DC, Okada T, Stenkamp RE, Yamamoto M, Miyano M (2000) Science 289:739–745
Teller DC, Okada T, Behnke CA, Palczewski K, Stenkamp RE (2001) Biochemistry 40:7761–7772
Yeagle PL, Choi G, Albert AD (2001) Biochemistry 40:11932–11937
Du LP, Li MY, Tsai KC, You QD, Xia L (2005) Biochem Biophys Res Commun 332:677–687
Yang Q, Du L, Wang X, Li M, You Q (2008) J Mol Graph Model 27:178–187
Du L, Li M, You Q, Xia L (2007) Biochem Biophys Res Commun 355:889–894
Li M, Fang H, Du L, Xia L, Wang B (2008) J Mol Model 14:957–966
Yang Q, Du L, Tsai K-C, Wang X, Li M, You Q (2009) QSAR Comb Sci 28:59–71
Jaakola VP, Griffith MT, Hanson MA, Cherezov V, Chien EY, Lane JR, Ijzerman AP, Stevens RC (2008) Science 322:1211–1217
Rasmussen SG, Choi HJ, Rosenbaum DM, Kobilka TS, Thian FS, Edwards PC, Burghammer M, Ratnala VR, Sanishvili R, Fischetti RF, Schertler GF, Weis WI, Kobilka BK (2007) Nature 450:383–387
Warne T, Serrano-Vega MJ, Baker JG, Moukhametzianov R, Edwards PC, Henderson R, Leslie AG, Tate CG, Schertler GF (2008) Nature 454:486–491
Rosenbaum DM, Rasmussen SG, Kobilka BK (2009) Nature 459:356–363
Mobarec JC, Sanchez R, Filizola M (2009) J Med Chem 52:5207–5216
Notredame C, Higgins DG, Heringa J (2000) J Mol Biol 302:205–217
Gouet P, Robert X, Courcelle E (2003) Nucleic Acids Res 31:3320–3323
Sali A, Blundell TL (1993) J Mol Biol 234:779–815
Im W, Feig M, Brooks CL 3rd (2003) Biophys J 85:2900–2918
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) J Comput Chem 4:187–217
MacKerell AD, Bashford D, Bellott M, Dunbrack RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiorkiewicz-Kuczera J, Yin D, Karplus M (1998) J Phys Chem B 102:3586–3616
Jo S, Kim T, Iyer VG, Im W (2008) J Comput Chem 29:1859–1865
Ryckaert J-P, Ciccotti G, Berendsen HJC (1977) J Comput Phys 23:327–341
Kleywegt GJ (2000) Acta Crystallogr D Biol Crystallogr 56:249–265
Laskowski RA, Rullmannn JA, MacArthur MW, Kaptein R, Thornton JM (1996) J Biomol NMR 8:477–486
Colovos C, Yeates TO (1993) Protein Sci 2:1511–1519
Eisenberg D, Luthy R, Bowie JU (1997) Methods Enzymol 277:396–404
Vriend G, Sander C (1993) J Appl Crystallogr 26:47–60
Tomii K, Hirokawa T, Motono C (2005) Proteins 7(61 Suppl):114–121
Shen MY, Sali A (2006) Protein Sci 15:2507–2524
Stewart JJ (1990) J Comput Aided Mol Des 4:1–105
Jakalian A, Bush BL, Jack DB, Bayly CI (2000) J Comput Chem 21:132–146
Jakalian A, Jack DB, Bayly CI (2002) J Comput Chem 23:1623–1641
Tsai K-C, Wang S-H, Hsiao N-W, Li M, Wang B (2008) Bioorg Med Chem Lett 18:3509–3512
OpenEye Scientific Software, Inc: Santa Fe, NM, 2007; pp. Quality Atomic Charges, Proton Assignment and Canonicalization
Moustakas DT, Lang PT, Pegg S, Pettersen E, Kuntz ID, Brooijmans N, Rizzo RC (2006) J Comput Aided Mol Des 20:601–619
McDonald IK, Thornton JM (1994) J Mol Biol 238:777–793
Wallace AC, Laskowski RA, Thornton JM (1995) Protein Eng 8:127–134
DeLano WL, DeLano (2006) Scientific, San Carlos, CA, USA. http://www.pymol.org
Clement OO, Mehl AT (2000) HipHop: pharmacophores based on multiple common-feature alignments. In: Pharmacophore perception, development & use in drug design, International University Line, pp 69–84
Barnum D, Greene J, Smellie A, Sprague P (1996) J Chem Inf Comput Sci 36:563–571
Rayan A (2010) J Mol Model 16:183–191
Mehler EL, Periole X, Hassan SA, Weinstein H (2002) J Comput Aided Mol Des 16:841–853
Anthes JC, Gilchrest H, Richard C, Eckel S, Hesk D, West RE Jr, Williams SM, Greenfeder S, Billah M, Kreutner W, Egan RE (2002) Eur J Pharmacol 449:229–237
Ghoneim OM, Legere JA, Golbraikh A, Tropsha A, Booth RG (2006) Bioorg Med Chem 14:6640–6658
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Tables S1-2
For pharmacophore external validation (DOC 463 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Yang, Q., Li, M. et al. In silico binding characteristics between human histamine H1 receptor and antagonists. J Mol Model 16, 1529–1537 (2010). https://doi.org/10.1007/s00894-010-0666-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-010-0666-z