Abstract
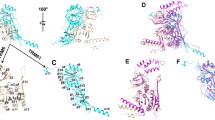
2′-O-ribose methylation is one of the most common posttranscriptional modifications in RNA. Methylations at different positions are introduced by enzymes from at least two unrelated superfamilies. Recently, a new family of eukaryotic RNA methyltransferases (MTases) has been identified, and its representative from yeast (Yol125w, renamed as Trm13p) has been shown to 2′-O-methylate position 4 of tRNA. Trm13 is conserved in Eukaryota, but exhibits no sequence similarity to other known MTases. Here, I present the results of bioinformatics analysis which suggest that Trm13 is a strongly diverged member of the Rossmann-fold MTase (RFM) superfamily, and therefore is evolutionarily related to 2′-O-MTases such as Trm7 and fibrillarin. However, the character of conserved residues in the predicted active site of the Trm13 family suggests it may use a different mechanism of ribose methylation than its relatives. A molecular model of the Trm13p structure has been constructed and evaluated for potential accuracy using model quality assessment methods. The predicted structure will facilitate experimental analyses of the Trm13p mechanism of action.



Similar content being viewed by others
Abbreviations
- aa:
-
Amino acid(s)
- e:
-
Expectation
- MTase:
-
Methyltransferase
- RFM:
-
Rossmann-fold MTase
- SAM:
-
AdoMet, S-adenosyl-L-methionine
References
Grosjean H (2005) Fine-tuning of RNA functions by modification and editing, vol 12. Springer, Berlin-Heidelberg
Dunin-Horkawicz S, Czerwoniec A, Gajda MJ, Feder M, Grosjean H, Bujnicki JM (2006) MODOMICS: a database of RNA modification pathways. Nucleic Acids Res 34(Database issue):D145–D149. doi:10.1093/nar/gkj084
Tollervey D, Lehtonen H, Jansen R, Kern H, Hurt EC (1993) Temperature-sensitive mutations demonstrate roles for yeast fibrillarin in pre-rRNA processing, pre-rRNA methylation, and ribosome assembly. Cell 72(3):443–457. doi:10.1016/0092-8674(93)90120-F
Feder M, Pas J, Wyrwicz LS, Bujnicki JM (2003) Molecular phylogenetics of the RrmJ/fibrillarin superfamily of ribose 2′-O-methyltransferases. Gene 302(1–2):129–138. doi:10.1016/S0378-1119(02)01097-1
Pintard L, Lecointe F, Bujnicki JM, Bonnerot C, Grosjean H, Lapeyre B (2002) Trm7p catalyses the formation of two 2′-O-methylriboses in yeast tRNA anticodon loop. EMBO J 21(7):1811–1820. doi:10.1093/emboj/21.7.1811
Pintard L, Bujnicki JM, Lapeyre B, Bonnerot C (2002) MRM2 encodes a novel yeast mitochondrial 21 S rRNA methyltransferase. EMBO J 21(5):1139–1147. doi:10.1093/emboj/21.5.1139
Tkaczuk KL, Dunin-Horkawicz S, Purta E, Bujnicki JM (2007) Structural and evolutionary bioinformatics of the SPOUT superfamily of methyltransferases. BMC Bioinformatics 8:73. doi:10.1186/1471-2105-8-73
Cavaille J, Chetouani F, Bachellerie JP (1999) The yeast Saccharomyces cerevisiae YDL112w ORF encodes the putative 2′-O-ribose methyltransferase catalyzing the formation of Gm18 in tRNAs. RNA 5(1):66–81
Lapeyre B, Purushothaman SK (2004) Spb1p-directed formation of Gm2922 in the ribosome catalytic center occurs at a late processing stage. Mol Cell 16:663–669. doi:10.1016/j.molcel.2004.10.022
Clouet-d'Orval B, Gaspin C, Mougin A (2005) Two different mechanisms for tRNA ribose methylation in Archaea: a short survey. Biochimie 87:889–895. doi:10.1016/j.biochi.2005.02.004
Purta E, van Vliet F, Tkaczuk KL, Dunin-Horkawicz S, Mori H, Droogmans L, Bujnicki JM (2006) The yfhQ gene of Escherichia coli encodes a tRNA:Cm32/Um32 methyltransferase. BMC Mol Biol 7:23. doi:10.1186/1471-2199-7-23
Wilkinson ML, Crary SM, Jackman JE, Grayhack EJ, Phizicky EM (2007) The 2′-O-methyltransferase responsible for modification of yeast tRNA at position 4. RNA 13:404–413. doi:10.1261/rna.399607
Schubert HL, Blumenthal RM, Cheng X (2003) Many paths to methyltransfer: a chronicle of convergence. Trends Biochem Sci 28:329–335. doi:10.1016/S0968-0004(03)00090-2
Kozbial PZ, Mushegian AR (2005) Natural history of S-adenosylmethionine-binding proteins. BMC Struct Biol 5:19. doi:10.1186/1472-6807-5-19
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. doi:10.1093/nar/25.17.3389
Kurowski MA, Bujnicki JM (2003) GeneSilico protein structure prediction meta-server. Nucleic Acids Res 31:3305–3307. doi:10.1093/nar/gkg557
Lundstrom J, Rychlewski L, Bujnicki J, Elofsson A (2001) Pcons: a neural-network-based consensus predictor that improves fold recognition. Protein Sci 10:2354–2362. doi:10.1110/ps.08501
Kosinski J, Gajda MJ, Cymerman IA, Kurowski MA, Pawlowski M, Boniecki M, Obarska A, Papaj G, Sroczynska-Obuchowicz P, Tkaczuk KL et al (2005) FRankenstein becomes a cyborg: the automatic recombination and realignment of fold recognition models in CASP6. Proteins 61(Suppl 7):106–113. doi:10.1002/prot.20726
Simons KT, Kooperberg C, Huang E, Baker D (1997) Assembly of protein tertiary structures from fragments with similar local sequences using simulated annealing and Bayesian scoring functions. J Mol Biol 268(1):209–225. doi:0022-2836/97/160209-17 $25.00/0/mb970959
Tramontano A, Morea V (2003) Assessment of homology-based predictions in CASP5. Proteins 53(Suppl 6):352–368. doi:10.1002/prot.20187
Wang G, Jin Y, Dunbrack RL Jr (2005) Assessment of fold recognition predictions in CASP6. Proteins 61(Suppl 7):46–66. doi:10.1002/prot.20721
Bujnicki JM, Rychlewski L (2002) RNA:(guanine-N2) methyltransferases RsmC/RsmD and their homologs revisited-bioinformatic analysis and prediction of the active site based on the uncharacterized Mj0882 protein structure. BMC Bioinformatics 3(1):10. doi:10.1186/1471-2105-3-10
Purta E, van Vliet F, Tricot C, De Bie LG, Feder M, Skowronek K, Droogmans L, Bujnicki JM (2005) Sequence-structure-function relationships of a tRNA (m(7)G46) methyltransferase studied by homology modeling and site-directed mutagenesis. Proteins 59(3):482–488. doi:10.1002/prot.20454
Fabrega C, Hausmann S, Shen V, Shuman S, Lima CD (2004) Structure and mechanism of mRNA cap (guanine-N7) methyltransferase. Mol Cell 13(1):77–89. doi:10.1016/S1097-2765(03)00522-7
Sunita S, Purta E, Durawa M, Tkaczuk KL, Swaathi J, Bujnicki JM, Sivaraman J (2007) Functional specialization of domains tandemly duplicated within 16 S rRNA methyltransferase RsmC. Nucleic Acids Res 35(13):4264–4274. doi:10.1093/nar/gkm411
Zegers I, Gigot D, van Vliet F, Tricot C, Aymerich S, Bujnicki JM, Kosinski J, Droogmans L (2006) Crystal structure of Bacillus subtilis TrmB, the tRNA (m7G46) methyltransferase. Nucleic Acids Res 34(6):1925–1934. doi:10.1093/nar/gkl116
Bradley P, Malmstrom L, Qian B, Schonbrun J, Chivian D, Kim DE, Meiler J, Misura KM, Baker D (2005) Free modeling with Rosetta in CASP6. Proteins 61:128–134. doi:10.1002/prot.20729
Wallner B, Elofsson A (2006) Identification of correct regions in protein models using structural, alignment, and consensus information. Protein Sci 15(4):900–913. doi:10.1110/ps.051799606
Wallner B, Fang H, Elofsson A (2003) Automatic consensus-based fold recognition using Pcons, ProQ, and Pmodeller. Proteins 53(Suppl 6):534–541. doi:10.1002/prot.10536
Pawlowski M, Gajda MJ, Matlak R, Bujnicki JM (2008) MetaMQAP: a meta-server for the quality assessment of protein models BMC. Bioinformatics 9:403. doi:10.1186/1471-2105-9-403
Armon A, Graur D, Ben-Tal N (2001) ConSurf: an algorithmic tool for the identification of functional regions in proteins by surface mapping of phylogenetic information. J Mol Biol 307(1):447–463. doi:10.1006/jmbi.2001.4474
Glaser F, Pupko T, Paz I, Bell RE, Bechor-Shental D, Martz E, Ben-Tal N (2003) ConSurf: identification of functional regions in proteins by surface-mapping of phylogenetic information. Bioinformatics 19(1):163–164. doi:10.1093/bioinformatics/19.1.163
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci USA 98(18):10037–10041. doi:10.1073/pnas.181342398
DeLano WL (2002) The PyMol Molecular Graphics System. on World Wide Web http://www.pymol.org, DeLano Scientific, Palo Alto, CA, USA:1.
Froimowitz M (1993) HyperChem: a software package for computational chemistry and molecular modeling. Biotechniques 14(6):1010–1013
Hager J, Staker BL, Jakob U (2004) Substrate binding analysis of the 23 S rRNA methyltransferase RrmJ. J Bacteriol 186:6634–6642. doi:10.1128/JB.186.19.6634-6642.2004
Kealey JT, Santi DV (1991) Identification of the catalytic nucleophile of tRNA (m5U54)methyltransferase. Biochemistry 30:9724–9728. doi:10.1021/bi00104a022
Wu JC, Santi DV (1987) Kinetic and catalytic mechanism of HhaI methyltransferase. J Biol Chem 262(10):4778–4786
Liu Y, Santi DV (2000) m5C RNA and m5C DNA methyl transferases use different cysteine residues as catalysts. PNAS 97(15):8263–8265
Bujnicki JM, Feder M, Ayres CL, Redman KL (2004) Sequence-structure-function studies of tRNA:m5C methyltransferase Trm4p and its relationship to DNA:m5C and RNA:m5U methyltransferases. Nucleic Acids Res 32:2453–2463. doi:10.1093/nar/gkh564
Tkaczuk KL, Obarska A, Bujnicki JM (2006) Molecular phylogenetics and comparative modeling of HEN1, a methyltransferase involved in plant microRNA biogenesis. BMC Evol Biol 6:6. doi:10.1186/1471-2148-6-100
Andreeva A, Tidow H (2008) A novel CHHC Zn-finger domain found in spliceosomal proteins and tRNA modifying enzymes. Bioinformatics 24(20):2277–2280. doi:10.1093/bioinformatics/btn431
O'Farrell HC, Scarsdale JN, Rife JP (2004) Crystal structure of KsgA, a universally conserved rRNA adenine dimethyltransferase in Escherichia coli. J Mol Biol 339(2):337–353. doi:10.1016/j.jmb.2004.02.068
Zhou H, Zhou Y (2004) Single-body residue-level knowledge-based energy score combined with sequence-profile and secondary structure information for fold recognition. Proteins 55(4):1005–1013. doi:10.1002/prot.20007
Rychlewski L, Jaroszewski L, Li W, Godzik A (2000) Comparison of sequence profiles. Strategies for structural predictions using sequence information. Protein Sci 9(2):232–241. doi::10.1110/ps.9.2.232
Soding J, Biegert A, Lupas AN (2005) The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res 33(Web Server issue):W244–W248. doi:10.1093/nar/gki408
Schluckebier G, Zhong P, Stewart KD, Kavanaugh TJ, Abad-Zapatero C (1999) The 2.2 A structure of the rRNA methyltransferase ErmC' and its complexes with cofactor and cofactor analogs: implications for the reaction mechanism. J Mol Biol 289(2):277–291. doi:10.1006/jmbi.1999.2788
Fischer D (2003) 3D-SHOTGUN: a novel, cooperative, fold-recognition meta-predictor. Proteins 51(3):434–441. doi:10.1002/prot.10357
Kelley LA, MacCallum RM, Sternberg MJ (2000) Enhanced genome annotation using structural profiles in the program 3D-PSSM. J Mol Biol 299(2):499–520. doi:10.1006/jmbi.2000.3741
Bussiere DE, Muchmore SW, Dealwis CG, Schluckebier G, Nienaber VL, Edalji RP, Walter KA, Ladror US, Holzman TF, Abad-Zapatero C (1998) Crystal structure of ErmC', an rRNA methyltransferase which mediates antibiotic resistance in bacteria. Biochemistry 37(20):7103–7112. doi:10.1021/bi973113c
Jones DT (1999) GenTHREADER: an efficient and reliable protein fold recognition method for genomic sequences. J Mol Biol 287(4):797–815. doi:10.1006/jmbi.1999.2583
Yu L, Petros AM, Schnuchel A, Zhong P, Severin JM, Walter K, Holzman TF, Fesik SW (1997) Solution structure of an rRNA methyltransferase (ErmAM) that confers macrolide-lincosamide-streptogramin antibiotic resistance. Nat Struct Biol 4(6):483–489. doi:10.1038/nsb0697-483
Lim K, Zhang H, Tempczyk A, Bonander N, Toedt J, Howard A, Eisenstein E, Herzberg O (2001) Crystal structure of YecO from Haemophilus influenzae (HI0319) reveals a methyltransferase fold and a bound S-adenosylhomocysteine. Proteins 45(4):397–407. doi:10.1002/prot.10004
Shi J, Blundell TL, Mizuguchi K (2001) FUGUE: sequence-structure homology recognition using environment-specific substitution tables and structure-dependent gap penalties. J Mol Biol 310(1):243–257. doi:10.1006/jmbi.2001.4762
Acknowledgments
KLT was supported by two grants from the Polish Ministry of Science (grant number N301 2396 33 and doctoral grant number N301 105 32/3599) and the START fellowship from Foundation for Polish Science. KLT would like to thank Dr. Janusz M. Bujnicki for the discussions and comments concerning the presented material and this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tkaczuk, K.L. Trm13p, the tRNA:Xm4 modification enzyme from Saccharomyces cerevisiae is a member of the Rossmann-fold MTase superfamily: prediction of structure and active site. J Mol Model 16, 599–606 (2010). https://doi.org/10.1007/s00894-009-0570-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-009-0570-6