Abstract
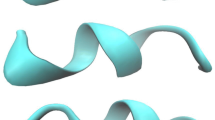
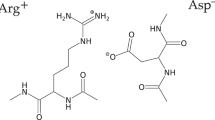
Peptide side chain interactions were studied by molecular dynamics simulation using explicit solvent on a peptide with the sequence AAARAAAAEAAEAAAARA. Three different protonation states of the glutamic acid side chains were simulated for four 20 ns runs each, a total simulation time of 240 ns. Two different salt bridge geometries were observed and the preferred geometry was found to depend on Glu — Arg residue spacing. Stable charge clusters were also observed, particularly in the fully charged peptide. Salt bridges were selectively interrupted upon protonation, with concomitant changes in secondary structure. The fully charged peptide was highly helical between residues 9 and 13, although protonation increased helicity near the N-terminus. The contribution of salt bridges to helix stability therefore depends on both position and relative position of charged residues within a sequence.




Similar content being viewed by others
References
Frauenfelder H, McMahon BH (2000) Ann Phys 9:655–667
Fenimore PW, Frauenfelder H, McMahon BH, Parak FG (2002) Proc Natl Acad Sci USA 99:16047–16501
Zimm BH, Bragg JK (1959) J Chem Phys 31:526–535
Lifson S, Roig A (1961) J Chem Phys 34:1963–1974
Schwarz G (1968) Biopolymers 6:873–897
Poland D, Scheraga HA (1966) J Chem Phys 45:2071–2090
Snow CD, Nguyen H, Pande VS, Gruebele M (2002) Nature 420:102–106
Gnanakaran S, Garcia AE (2003) J Phys Chem B 107:12555–12557
Sorin EJ, Pande VS (2005) Biophys J 88:2472–2493
Shoemaker KR, Kim PS, York EJ, Stewart JM, Baldwin RL (1987) Nature 326:563–567
Fairman R, Shoemaker KR, York EJ, Stewart JM, Baldwin RL (1990) Biophys Chem 37:107–119
Pozo Ramajo A, Petty SA, Volk M (2006) Chem Phys 323:11–20
Garcia AE, Sanbonmatsu KY (2002) Proc Natl Acad Sci USA 99:2782–2787
Bierzynski A, Kim PS, Baldwin RL (1982) Proc Natl Acad Sci USA 79:2470–2474
Wlodawer A, Svensson LA, Sjölin L, Gilliland GL (1988) Biochemistry 27:2705–2717
Rico M, Gallego E, Santoro J, Bermejo FJ, Nieto JL, Herranz J (1984) Biochem Biophys Res Commun 123:757–763
Ghosh T, Garde S, Garcia AE (2003) Biophys J 85:3187–3193
Wang WZ, Lin T, Sun YC (2007) J Phys Chem B 111:3508–3514
Okur A, Wickstrom L, Layten M, Geney R, Song K, Hornak V, Simmerling C (2006) J Chem Theory Comput 2:420–433
Sugita Y, Okamoto Y (2005) Biophys J 88:3180–3190
Monticelli L, Tieleman DP, Colombo G (2005) J Phys Chem B 109:20064–20067
Kalé L, Skeel R, Bhandarkar M, Brunner R, Gursoy A, Krawetz N, Phillips J, Shinozaki A, Varadarajan K, Schulten K (1999) J Comput Phys 151:283–312
Humphrey W, Dalke A, Schulten K (1996) J Mol Graphics 14:33–38
Feig M, Karanicolas J, Charles L, Brooks I (2004) J Mol Graphics Modell 22:377–395
Zhang W, Lei H, Chowdhury S, Duan Y (2004) J Phys Chem B 108:7479–7489
Williams S, Causgrove TP, Gilmanshin R, Fang KS, Callender RH, Woodruff WH, Dyer RB (1996) Biochemistry 35:691–697
Okur A, Wickstrom L, Simmerling C (2008) J Chem Theory Comput 4:488–498
Acknowledgments
Simulations were performed on the High-Performance Computing Cluster at Texas A&M University-Corpus Christi, which is supported by grant #0321218 from the National Science Foundation. This work was supported by a grant from Texas Research Development Fund.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Walker, K.D., Causgrove, T.P. Contribution of arginine-glutamate salt bridges to helix stability. J Mol Model 15, 1213–1219 (2009). https://doi.org/10.1007/s00894-009-0482-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-009-0482-5