Abstract
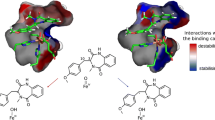
The 3D structure of a novel epoxide hydrolase from Aspergillus niger SQ-6 (sqEH) was constructed by using homology modeling and molecular dynamics simulations. Based on the 3D model, Asp191, His369 and Glu343 were predicted as catalytic triad. The putative active pocket is a hydrophobic environment and is rich in some important non—polar residues (Pro318, Trp282, Pro319, Pro317 and Phe242). Using three sets of epoxide inhibitors for docking study, the interaction energies of sqEH with each inhibitor are consistent with their inhibitory effects in previous experiments. Moreover, a critical water molecule which closes to the His369 was identified to be an ideal position for the hydrolysis step of the reaction. Two tyrosine residues (Tyr249 and Tyr312) are able to form hydrogen bonds with the epoxide oxygen atom to maintain the initial binding and positioning of the substrate in the active pocket. These docked complex models can well interpret the substrate specificity of sqEH, which could be relevant for the structural—based design of specific epoxide inhibitors.









Similar content being viewed by others
References
Arand M, Oesch F (2002) Wiley, New York, pp 459-483
Armstrong RN (1999) Drug Metab Rev 31:71–86
Oesch F (1973) Xenobiotica 3:305–340
Arand M, Wagner H, Oesch F (1996) J Biol Chem 271:4223–4229
Lacourciere GM, Armstrong RN (1993) J Am Chem Soc 115:10466–10467
Weijers CAGM, de Bont JAM (1999) J Mol Catal B Enzym 6:199–214
Armstrong RN, Cassidy CS (2000) Drug Metab Rev 32:327–338
Rink R, Fennema F, Smids M, Dehmel U, Janssen DB (1997) J Biol Chem 272:14650–14657
Arand M, Hemmer H, DSrk H, Baratti J, Archelas A, Furstoss R, Oesch F (1999) Biochem J 344:273–280
Wojtasek H, Prestwich GD (1996) Biochem Biophys Res Commun 220:323–329
Ready JM, Jacobsen EN (2001) J Am Chem Soc 123:2687–2688
Bell PA, Kasper CB (1993) J Biol Chem 268:14011–14017
Laughlin LT, Tzeng HF, Lin S, Armstrong RN (1998) Biochemistry 37:2897–2904
Moussou P, Archelas A, Furstoss R (1998) J Mol Catal B Enzym 5:447–458
Moussou P, Archelas A, Baratti J, Furstoss R (1998) Tetrahedron: Asymmetry 9:1539–1547
Cleij M, Archelas A, Furstoss R (1998) Tetrahedron: Asymmetry 9:1839–1842
Liu Y, Wu S, Wang J, Yang L, Sun W (2007) Protein Expr Purif 53:239–246
Zou J, Hallberg BM, Bergfors T, Oesch F, Arand M, Mowbray SL, Jones TA Structure 8:111–122
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Nucleic Acids Res 25:3389–3402
Higgins DG, Bleasby AJ, Fuchs R (1992) Comput Appl Biosci 8:189–191
Ke YY, Chen YC, Lin TH (2006) J Comput Chem 27:1556–1570
Gui C, Zhu W, Chen G, Luo X, Liew OW, Puah CM, Chen K, Jiang H (2007) Proteins 67:41–52
Accelrys. InsightII, Version 2000, Homology User Guide. San Diego: Biosym/MSI
Case DA, Darden TA, Cheatham TE III, Simmerling CL, Wang J, Duke RE, Luo R, Merz KM, Pearlman DA, Crowley M, Walker RC, Zhang W, Wang B, Hayik S, Roitberg A, Seabra G, Wong KF, Paesani F, Wu X, Brozell S, Tsui V, Gohlke H, Yang L, Tan C, Mongan J, Hornak V, Cui G, Beroza P, Mathews DH, Schafmeister C, Ross WS, Kollman PA (2006) AMBER 9 University of California San Francisco
Case DA, Cheatham TE, Darden T, Gohlke H, Luo R, Merz KM, Onufriev A, Simmerling C, Wang B, Woods R (2005) The Amber biomolecular simulation programs. J Comput Chem 26:1668–1688
Ponder JW, Case DA (2003) Adv Prot Chem 66:27–85
Simmerling C, Strockbine B, Roitberg AE (2002) J Am Chem Soc 124:11258–11259
Pastor RW, Brooks BR, Szabo A (1988) Mol Phys 65:1409–1419
Tuccinardi T, Manetti F, Schenone S, Martinelli A, Botta M (2007) J Chem Inf Model 47:644–655
Accelrys. InsightII, Version 2000, Profiles-3D. San Diego: Biosym/MSI
Lüthy R, Bowie JU, Eisenberg D (1992) Nature 356:83–85
Laskowski RA, Macarthur MW, Moss DS, Thornton JM (1993) J Appl Crystallogr 26:283–291
Binding site analysis user guide. (2000) San Diego, USA: Accelrys, Inc
Affinity San Diego Molecular Simulations Inc (2000)
Bartlett PA, Shea GT, Telfer SJ, Waterman S (1989) R Soc Chem 182–196
Shoichet BK, Kuntz ID, Bodian DL (1992) J Comput Chem 13:380–397
Nardini M, Dijkstra BW (1999) Curr Opin Struct Biol 9:732–737
Acknowledgements
This work was supported by the National Science Foundation of China (20333050, 20673044), PCSIRT (IRT0625), Key subject of Science and Technology by Jilin Province and 985 Graduate Innovation Program of Jilin University (20080220). We also thank professor David A. Case et al for providing us the Amber 9 software as a freeware.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luo, Q., Yao, Y., Han, WW. et al. Homology modeling of a novel epoxide hydrolase (EH) from Aspergillus niger SQ-6: structure-activity relationship in expoxides inhibiting EH activity. J Mol Model 15, 1125–1132 (2009). https://doi.org/10.1007/s00894-009-0466-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-009-0466-5