Abstract
Dipeptidyl peptidase IV (DPP-IV) deactivates the incretin hormones GLP-1 and GIP by cleaving the penultimate proline or alanine from the N-terminal (P1-position) of the peptide. Inhibition of this enzyme will prevent the degradation of the incretin hormones and maintain glucose homeostasis; this makes it an attractive target for the development of drugs for diabetes. This paper reports 3D-QSAR analysis of several DPP-IV inhibitors, which were aligned by the receptor-based technique. The conformation of the molecules in the active site was obtained through docking methods. The QSAR models were generated on two training sets composed of 74 and 25 molecules which included phenylalanine, thiazolidine, and fluorinated pyrrolidine analogs. The 3D-QSAR models are robust with statistically significant r2, q2, and \({\text{r}}^{2}_{{{\text{pred}}}} \) values. The CoMFA and CoMSIA models were used to design some new inhibitors with several fold higher binding affinity.

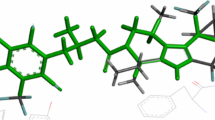
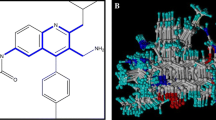
The CoMFA contours around molecule D1T155 (a) steric contours - favored (green); disfavored (yellow) (b) electrostatic contours - electropositive (blue); electronegative (red)











Similar content being viewed by others
References
Ogata S, Misumi Y, Ikehara YS (1959) J Biol Chem 264:596–601
Bjelke RJ, Christensen J, Branner S, Wagtmann N, Olsen C, Kanstrup AB, Rasmussen HB (2004) J Biol Chem 279:34691–34697
Abbott CA, Gorrell MD (2002) Ectopeptidases: CD13/Aminopeptidase N and CD26/Dipeptidylpeptidase IV. In: Langner J, Ansorge S (eds) Medicine and biology. Kulwer/Plenum, New York, pp 171–184
Lambeir AM, Diaz Pereira JF, Chacon P, Vermeulen G, Heremans K, Devreese B, Van Beeumen J, De Meester I, Scharpe S (1997) Biochim Biophys Acta 1340:215
Duke-Cohan JS, Morimoto C, Rocker JA, Schlossman SF (1996) J Immunol 156:1714–1721
Hartel S, Gossrau R, Hanski C, Reutter W (1988) Histochemistry 89:151–161
Engel M, Hoffmann T, Wagner L, Wermann M, Heiser U, Keifersauer R, Huber R, Bode W, Demuth H-U, Brandstetter H (2003) Proc Natl Acc Sci 100:5063–5068
Pederson RA, White HA, Schlenzig D, Pauly RP, McIntosh CH, Demuth H-U (1998) Diabetes 47:1253–1258
Pospisilik JA, Stafford SG, Demuth HU, McIntosh CH, Pederson RA (2002) Diabetes 51:2677–2683
Cheng JD, Dunbrack RL, Valianou M, Rogatko A, Alpaugh RK, Weiner LM (2002) Cancer Res 62:4767–4772
Kajiyama H, Kikkawa F, Suzuki T, Shibata K, Ino K, Mizutani S (2002) Cancer Res 62:2753–2757
Ho L, Aytac U, Stephens LC, Ohnuma K, Mills GB, McKee KS, Neumann C, LaPushin R, Cabanillas F, Abbruzzese JL, Morimoto C, Dang NH (2001) Clinic Cancer Res 7:2031–2040
Frohman LA, Downs TR, Heimer EP, Felix AM (1989) J Clinic Invest 83:1533–1540
Nausch I, Heymann E (1985) J Neurochem 44:1354–1357
Ahmad S, Wang L, Ward PE (1992) J Pharmacol Exp Ther 260:1257–1261
Nauck MA, Niedereichholz U, Ettler R, Holst JJ, Ørskov C, Ritzel R, Schmiegel WH (1997) Am J Physiol 273:E981–E988
Ahrén B (1998) BioEssays 20:642–651
Meier JJ, Nauck MA, Schmidt WE, Gallwitz B (2002) Regul Pept 107:1–13
Jörnvall H, Carlquist M, Kwauk S, Otte SC, McIntosh CH, Brown JC, Mutt V (1981) FEBS Lett 123:205–210
Adelhorst K, Hedegaard BB, Knudsen LB, Kirk O (1994) J Biol Chem 269:6275–6278
Rolin B, Deacon CF, Carr RD, Ahrén B (2004) Eur J Pharmacol 494:283–288
Green BD, Flatt PR, Bailey CJ (2006) Expert Opin Emerg Drugs 11:525–539
Kendall DM, Kim D, Maggs D (2006) Diabetes Technol Ther 8:385–396
Ahren B, Landin-Olsson M, Jansson PA, Svensson M, Holmes D, Schweizer A (2004) J Clin Endocrinol Metab 89:2078–2084
Pratley R, Galbreath E (2004) Diabetes 53:A83
Herman GA, Zhao P-L, Dietrich G (2004) Diabetologia 47:794–805
Triplitt C, Wright A, Chiquette E (2006) Pharmacotherapy 26:360–374
Brandt W, Lehmann T, Barth A, Fittkaui S (1993) J Mol Graphics 11:277–278
Brandt W, Lehmann T, Thondor I, Born I, Schutkowski M, Rahfeld JU, Neubert K, Barth A (1995) Int J Peptide Prot Res 46:494–507
Rasmussen HB, Branner S, Wiberg FC, Wagtmann NR (2003) Nat Struct Biol 10:19–25
Qiao L, Baumann CA, Crysler CS, Ninan NS, Abad MC, Spurlino JC, Desjarlais RL, Kervinen J, Neeper MP, Bayoumy SS, Williams R, Deckman IC, Dasgupta M, Reed RL, Huebert ND, Tomczuk BE, Moriarty KJ (2006) Bioorg Med Chem Lett 16:123–128
Kim D, Wang L, Beconi M, Eiermann GJ, Fisher MH, He H, Hickey GJ, Kowalchick JE, Leiting B, Lyons K, Marsilio F, Mccann ME, Patel RA, Petrov A, Scapin G, Patel SB, Roy RS, Wu JK, Wyvratt MJ, Zhang BB, Zhu L, Thornberry NA, Weber AE (2005) J Med Chem 48:141–151
Nordhoff S, Cerezo-Galvez S, Feurer A, Hill O, Matassa VG, Metz G, Rummey C, Thiemann M, Edwards PJ (2006) Bioorg Med Chem Lett 16:1744–1748
GOLD 3.0.1 CCDC Ltd., UK
Sybyl version 7.1 Tripos Inc., USA
Cerius2 version 4.8, Accelrys Inc., USA
Nayyar A, Malde A, Jain R, Coutinho EC (2006) Bioorg Med Chem 14:847–856
Tanimoto TT, Trans NY (1961) Acad Sci Ser 2(23):576–578
Nikolova N, Jaworska J (2003) QSAR Comb Sci 22:1006–1026
Xu J, Ok HO, Gonzalez EJ, Colwell LF Jr, Habulihaz B, He H, Leiting B, Lyons KA, Marsilio F, Patel RA, Wu JK, Thornberry NA, Weber AE, Parmee ER (2004) Bioorg Med Chem Lett 14:4759–4762
Edmondson SD, Mastracchio A, Duffy JL, Eiermann GJ, He H, Ita I, Leiting B, Leone JF, Lyons KA, Makarewicz AM, Patel RA, Petrov A, Wu JK, Thornberry NA, Weber AE (2005) Bioorg Med Chem Lett 15:3048–3052
Xu J, Wei L, Mathvink R, He J, Park Y-J, He H, Leiting B, Lyons KA, Marsilio F, Patel RA, Wu JK, Thornberry NA, Weber AE (2005) Bioorg Med Chem Lett 15:2533–2536
Hulin B, Cabral S, Lopaze MG, Van Volkenburg MA, Andrews KM, Parker JC (2005) Bioorg Med Chem Lett 15:4770–4773
InsightII version 2005L, Accelrys Inc, USA
Datar PA, Coutinho EC (2004) J Mol Graph Model 23:239–251
Cramer RD, Patterson DE, Bunce JD (1988) J Am Chem Soc 110:5959–5967
Klebe G, Abraham U, Mietzner T (1994) J Med Chem 37:4130–4146
Klebe G (1998) Drug Discov Des 12:87–104
Böhm M, Stürzebecher J, Klebe G (1999) J Med Chem 42:458–477
Wold S, Johansson E, Cocchi M (1993) PLS-partial least squares projections to latent structures in 3D-QSAR. In: Kubinyi H (ed) Drug design; theory methods and applications. ESCOM Lieden, The Netherlands, pp 523–550
Bush BL, Nachbar RB (1993) J Comput Aided Mol Des 7:587–619
Legar C, Politis DN, Romano JP (1992) Technometrics 34:378–399
Edmondson SD, Mastracchio A, Beconi M, Colwell LF, Habulihaz B, He H, Kumar S, Leiting B, Lyons KA, Mao A, Marsilio F, Patel RA, Wu JK, Zhu L, Thornberry NA, Weber AE, Parmee ER (2004) Bioorg Med Chem Lett 14:5151–5155
Ludwig K, Yan S, Fan H, Reutter W, Bottcher C (2003) Biochem Biophys Res Commun 304:73–77
Mentlein R (1999) Regul Pept 85:9–24
Ogata S, Misumi Y, Tsuji E, Takami N, Oda K, Ikehara Y (1992) Biochemistry 31:2582–2587
Abbott CA, McCaughan GW, Gorrell MD (1999) FEBS Letters 458:278–284
Datar PA, Khedkar SA, Malde AK, Coutinho EC (2006) J Comput Aided Mol Des 20:343–360
Acknowledgements
The computational facilities were jointly provided by the All India Council of Technical Education through grant (F. No. 8022/RID/NPROJ/RPS-5/2003-04) and the Department of Science and Technology through their FIST program (SR/FST/LSI-163/2003). R.R.S. Pissurlenkar thanks the Amrut Mody Research Foundation (AMRF) and M.S. Shaikh, the University Grants Commission (UGC) for financial support [Grant /F.No.7-16/2003(SR)].
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pissurlenkar, R.R.S., Shaikh, M.S. & Coutinho, E.C. 3D-QSAR studies of Dipeptidyl peptidase IV inhibitors using a docking based alignment. J Mol Model 13, 1047–1071 (2007). https://doi.org/10.1007/s00894-007-0227-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00894-007-0227-2