Abstract
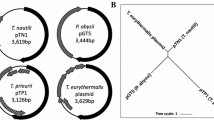
In this study, a transformation system enabling large-scale gene recombination was developed for the hyperthermophilic archaeon Thermococcus kodakarensis. Using the uracil auxotroph T. kodakarensis KU216 (∆pyrF) as a parent strain, we constructed multiple host strains harboring two 1-kbp DNA regions from the genomes of either the hyperthermophilic archaeon Pyrococcus furiosus or Methanocaldococcus jannaschii. The two regions were selected so that the regions between them on the respective genomes would include pyrF genes, which can potentially be used for selection. Transformation using these host strains and genomic DNA from P. furiosus or M. jannaschii were carried out. Transformants with exogenous pyrF were obtained only using host strains with regions from P. furiosus, and only when the distances between the two regions were relatively short (2–5 kbp) on the P. furiosus genome. To insert longer DNA fragments, we examined the possibilities of using P. furiosus cells to provide intact genomic DNA. A cell pellet of P. furiosus was overlaid with that of T. kodakarensis so that cells were in direct contact. As a result, we were able to isolate T. kodakarensis strains harboring DNA fragments from P. furiosus with lengths of up to 75 kbp in a single transformation step.






Similar content being viewed by others
Abbreviations
- PyrF:
-
Orotidine-5′-monophosphate decarboxylase
- ASW:
-
Artificial seawater
- YT:
-
Yeast extract and tryptone
- AA:
-
20 Amino acids
- 5-FOA:
-
5-Fluoroorotic acid
- CFE:
-
Cell-free extract
- GAP:
-
Glyceraldehyde 3-phosphate
References
Ajon M, Fröls S, van Wolferen M, Stoecker K, Teichmann D, Driessen AJ, Grogan DW, Albers SV, Schleper C (2011) UV-inducible DNA exchange in hyperthermophilic archaea mediated by type IV pili. Mol Microbiol 82:807–817
Atomi H, Fukui T, Kanai T, Morikawa M, Imanaka T (2004) Description of Thermococcus kodakaraensis sp. nov., a well studied hyperthermophilic archaeon previously reported as Pyrococcus sp. KOD1. Archaea 1:263–267
Bertani G, Baresi L (1987) Genetic transformation in the methanogen Methanococcus voltae PS. J Bacteriol 169:2730–2738
Bult CJ, White O, Olsen GJ, Zhou L, Fleischmann RD, Sutton GG, Blake JA, FitzGerald LM, Clayton RA, Gocayne JD et al (1996) Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 273:1058–1073
Burke DT, Carle GF, Olson MV (1987) Cloning of large segments of exogenous DNA into yeast by means of artificial chromosome vectors. Science 236:806–812
Fiala G, Stetter KO (1986) Pyrococcus furiosus sp. nov. represents a novel genus of marine heterotrophic archaebacteria growing optimally at 100 °C. Arch Microbiol 145:56–61
Fuke T, Sato T, Jha S, Tansengco ML, Atomi H (2018) Phytoene production utilizing the isoprenoid biosynthesis capacity of Thermococcus kodakarensis. Extremophiles 22:301–313
Fukui T, Atomi H, Kanai T, Matsumi R, Fujiwara S, Imanaka T (2005) Complete genome sequence of the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1 and comparison with Pyrococcus genomes. Genome Res 15:352–363
Gibson DG, Benders GA, Andrews-Pfannkoch C, Denisova EA, Baden-Tillson H, Zaveri J, Stockwell TB, Brownley A, Thomas DW, Algire MA et al (2008a) Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science 319:1215–1220
Gibson DG, Benders GA, Axelrod KC, Zaveri J, Algire MA, Moodie M, Montague MG, Venter JC, Smith HO, Hutchison CA 3rd (2008b) One-step assembly in yeast of 25 overlapping DNA fragments to form a complete synthetic Mycoplasma genitalium genome. Proc Natl Acad Sci USA 105:20404–20409
Gibson DG, Glass JI, Lartigue C, Noskov VN, Chuang RY, Algire MA, Benders GA, Montague MG, Ma L, Moodie MM et al (2010) Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329:52–56
Grogan DW (1996) Exchange of genetic markers at extremely high temperatures in the archaeon Sulfolobus acidocaldarius. J Bacteriol 178:3207–3211
Hutchison CA 3rd, Chuang RY, Noskov VN, Assad-Garcia N, Deerinck TJ, Ellisman MH, Gill J, Kannan K, Karas BJ, Ma L et al (2016) Design and synthesis of a minimal bacterial genome. Science 351:aad6253
Itaya M, Tsuge K, Koizumi M, Fujita K (2005) Combining two genomes in one cell: stable cloning of the Synechocystis PCC6803 genome in the Bacillus subtilis 168 genome. Proc Natl Acad Sci USA 102:15971–15976
Itaya M, Fujita K, Kuroki A, Tsuge K (2008) Bottom-up genome assembly using the Bacillus subtilis genome vector. Nat Methods 5:41–43
Itaya M, Sato M, Hasegawa M, Kono N, Tomita M, Kaneko S (2018) Far rapid synthesis of giant DNA in the Bacillus subtilis genome by a conjugation transfer system. Sci Rep 8:8792
Jones WJ, Leigh JA, Mayer F, Woese CR, Wolfe RS (1983) Methanococcus jannaschii sp. nov., an extremely thermophilic methanogen from a submarine hydrothermal vent. Arch Microbiol 136:254–261
Kanai T, Matsuoka R, Beppu H, Nakajima A, Okada Y, Atomi H, Imanaka T (2011) Distinct physiological roles of the three [NiFe]-hydrogenase orthologs in the hyperthermophilic archaeon Thermococcus kodakarensis. J Bacteriol 193:3109–3116
Kaneko S, Fukushima H, Nakahama M, Asano S, Miyazaki Y, Aizawa Y, Itaya M (2018) DNA synthesis by fragment assembly using extra-cellular DNA delivered by artificial controlled horizontal transfer. J Biochem 163:305–312
Keller MW, Lipscomb GL, Loder AJ, Schut GJ, Kelly RM, Adams MW (2015) A hybrid synthetic pathway for butanol production by a hyperthermophilic microbe. Metab Eng 27:101–106
Keller MW, Lipscomb GL, Nguyen DM, Crowley AT, Schut GJ, Scott I, Kelly RM, Adams MW (2017) Ethanol production by the hyperthermophilic archaeon Pyrococcus furiosus by expression of bacterial bifunctional alcohol dehydrogenases. Microb Biotechnol 10:1535–1545
Lipscomb GL, Stirrett K, Schut GJ, Yang F, Jenney FE Jr, Scott RA, Adams MW, Westpheling J (2011) Natural competence in the hyperthermophilic archaeon Pyrococcus furiosus facilitates genetic manipulation: construction of markerless deletions of genes encoding the two cytoplasmic hydrogenases. Appl Environ Microbiol 77:2232–2238
Lyu Z, Jain R, Smith P, Fetchko T, Yan Y, Whitman WB (2016) Engineering the autotroph Methanococcus maripaludis for geraniol production. ACS Synth Biol 5:577–581
Maeder DL, Weiss RB, Dunn DM, Cherry JL, González JM, DiRuggiero J, Robb FT (1999) Divergence of the hyperthermophilic archaea Pyrococcus furiosus and P. horikoshii inferred from complete genomic sequences. Genetics 152:1299–1305
Matsubara K, Yokooji Y, Atomi H, Imanaka T (2011) Biochemical and genetic characterization of the three metabolic routes in Thermococcus kodakarensis linking glyceraldehyde 3-phosphate and 3-phosphoglycerate. Mol Microbiol 81:1300–1312
Matsumi R, Manabe K, Fukui T, Atomi H, Imanaka T (2007) Disruption of a sugar transporter gene cluster in a hyperthermophilic archaeon using a host-marker system based on antibiotic resistance. J Bacteriol 189:2683–2691
Mevarech M, Werczberger R (1985) Genetic transfer in Halobacterium volcanii. J Bacteriol 162:461–462
Morikawa M, Izawa Y, Rashid N, Hoaki T, Imanaka T (1994) Purification and characterization of a thermostable thiol protease from a newly isolated hyperthermophilic Pyrococcus sp. Appl Environ Microbiol 60:4559–4566
Naor A, Lapierre P, Mevarech M, Papke RT, Gophna U (2012) Low species barriers in halophilic archaea and the formation of recombinant hybrids. Curr Biol 22:1444–1448
Patel GB, Nash JH, Agnew BJ, Sprott GD (1994) Natural and electroporation-mediated transformation of Methanococcus voltae protoplasts. Appl Environ Microbiol 60:903–907
Robb FT, Place AR (1995) Media for Thermophiles. In: Robb FT, Place AR (eds) Archaea: a laboratory manual-thermophiles. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp 167–168
Rosenshine I, Tchelet R, Mevarech M (1989) The mechanism of DNA transfer in the mating system of an archaebacterium. Science 245:1387–1389
Santangelo TJ, Čuboňová L, Reeve JN (2008) Shuttle vector expression in Thermococcus kodakaraensis: contributions of cis elements to protein synthesis in a hyperthermophilic archaeon. Appl Environ Microbiol 74:3099–3104
Santangelo TJ, Čuboňová L, Reeve JN (2010) Thermococcus kodakarensis genetics: TK1827-encoded β-glycosidase, new positive-selection protocol, and targeted and repetitive deletion technology. Appl Environ Microbiol 76:1044–1052
Sato T, Fukui T, Atomi H, Imanaka T (2003) Targeted gene disruption by homologous recombination in the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J Bacteriol 185:210–220
Sato T, Imanaka H, Rashid N, Fukui T, Atomi H, Imanaka T (2004) Genetic evidence identifying the true gluconeogenic fructose-1,6-bisphosphatase in Thermococcus kodakaraensis and other hyperthermophiles. J Bacteriol 186:5799–5807
Sato T, Fukui T, Atomi H, Imanaka T (2005) Improved and versatile transformation system allowing multiple genetic manipulations of the hyperthermophilic archaeon Thermococcus kodakaraensis. Appl Environ Microbiol 71:3889–3899
Schmidt KJ, Beck KE, Grogan DW (1999) UV stimulation of chromosomal marker exchange in Sulfolobus acidocaldarius: implications for DNA repair, conjugation and homologous recombination at extremely high temperatures. Genetics 152:1407–1415
Shalev Y, Turgeman-Grott I, Tamir A, Eichler J, Gophna U (2017) Cell surface glycosylation is required for efficient mating of Haloferax volcanii. Front Microbiol 8:1253
Shimosaka T, Makarova KS, Koonin EV, Atomi H (2019) Identification of dephospho-coenzyme A (dephospho-CoA) kinase in Thermococcus kodakarensis and elucidation of the entire CoA biosynthesis pathway in Archaea. mBio 10:e01146–19
Shizuya H, Birren B, Kim UJ, Mancino V, Slepak T, Tachiiri Y, Simon M (1992) Cloning and stable maintenance of 300-kilobase-pair fragments of human DNA in Escherichia coli using an F-factor-based vector. Proc Natl Acad Sci USA 89:8794–8797
Thorgersen MP, Lipscomb GL, Schut GJ, Kelly RM, Adams MW (2014) Deletion of acetyl-CoA synthetases I and II increases production of 3-hydroxypropionate by the metabolically-engineered hyperthermophile Pyrococcus furiosus. Metab Eng 22:83–88
Tomita H, Yokooji Y, Ishibashi T, Imanaka T, Atomi H (2012) Biochemical characterization of pantoate kinase, a novel enzyme necessary for coenzyme A biosynthesis in the Archaea. J Bacteriol 194:5434–5443
Tsuge K, Itaya M (2001) Recombinational transfer of 100-kilobase genomic DNA to plasmid in Bacillus subtilis 168. J Bacteriol 183:5453–5458
Worrell VE, Nagle DP Jr, McCarthy D, Eisenbraun A (1988) Genetic transformation system in the archaebacterium Methanobacterium thermoautotrophicum Marburg. J Bacteriol 170:653–656
Yokooji Y, Tomita H, Atomi H, Imanaka T (2009) Pantoate kinase and phosphopantothenate synthetase, two novel enzymes necessary for CoA biosynthesis in the Archaea. J Biol Chem 284:28137–28145
Acknowledgements
The authors are grateful to Ms. Rikako Fujimoto for constructing the expression plasmid pPcsgChiA1. This work was partially supported by JSPS KAKENHI Grant Number JP19H05679 (Post-Koch Ecology), JP19H05684 to H.A. and JP19H05689 to M.O., and initiated with the support of the CREST program of the Japan Science and Technology Agency (Grant No.: 10104047) to H.A. This work was also partially supported by JSPS KAKENHI to T.S. (Grant No.: 18K05411).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Moracci.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sato, T., Takada, D., Itoh, T. et al. Integration of large heterologous DNA fragments into the genome of Thermococcus kodakarensis. Extremophiles 24, 339–353 (2020). https://doi.org/10.1007/s00792-020-01159-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-020-01159-z