Abstract
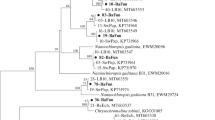
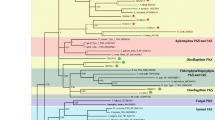
The modular polyketide synthase (PKS) and nonribosomal peptide synthetase (NRPS) have been found to be involved in natural product synthesis in many microorganisms. Study on their diversities in natural environment may provide important ecological insights, in addition to opportunities for antibacterial drugs development. In this study, the PKS and NRPS gene diversities in two coast sediments near China Zhongshan Station were studied. The phylogenetic analysis of amino acid (AA) sequences indicated that the identified ketosynthase (KS) domains were clustered with those from diverse bacterial groups, including Proteobacteria, Firmicutes, Planctomycetes, Cyanobacteria, Actinobacteria, and some uncultured symbiotic bacteria. One new branch belonging to hybrid PKS/NRPS enzyme complexes and five independent clades were found on the phylogenetic tree. The obtained adenylation (A) domains were mainly clustered within the Cyanobacteria and Proteobacteria group. Most of the identified KS and A domains showed below 80 and 60% identities at the AA level to their closest matches in GenBank, respectively. The diversities of both KS and A domains in natural environmental sample were different from those in sewage-contaminated sample. These results revealed the great diversity and novelty of both PKS and NRPS genes in Antarctic sediment.



Similar content being viewed by others
References
Baker JT, Borris RP, Carte B, Cordell GA, Soejarto DD, Cragg GM, Gupta MP, Iwu MM, Madulid DR, Tyler VE (1995) Natural product drug discovery and development: new perspectives on international collaboration. J Nat Prod 58:1325–1357
Beja O, Suzuki MT, Koonin EV, Aravind L, Hadd A, Nguyen LP, Villacorta R, Amjadi M, Garrigues C, Jovanovich SB, Feldman RA, DeLong EF (2000) Construction and analysis of bacterial artificial chromosome libraries from a marine microbial assemblage. Environ Microbiol 2:516–529
Bowman JP, McCuaig RD (2003) Biodiversity, community structural shifts, and biogeography of prokaryotes within Antarctic continental shelf sediment. Appl Environ Microbiol 69:2463–2483
Bowman JP, Rea SM, McCammon SA, McMeekin TA (2000) Diversity and community structure within anoxic sediment from marine salinity meromictic lakes and a coastal meromictic marine basin, Vestfold Hilds, Eastern Antarctica. Environ Microbiol 2:227–237
Burns BP, Seifert A, Goh F, Pomati F, Jungblut AD, Serhat A, Neilan BA (2005) Genetic potential for secondary metabolite production in stromatolite communities. FEMS Microbiol Lett 243:293–301
Cane DE, Walsh CT (1999) The parallel and convergent universes of polyketide synthases and nonribosomal peptide synthetases. Chem Biol 6:R319–R325
Cane DE, Walsh CT, Khosla C (1998) Harnessing the biosynthetic code: combinations, permutations, and mutations. Science 282:63–68
Courtois S, Cappellano CM, Ball M, Francou FX, Normand P, Helynck G, Martinez A, Kolvek SJ, Hopke J, Osburne MS, August PR, Nalin R, Guerineau M, Jeannin P, Simonet P, Pernodet JL (2003) Recombinant environmental libraries provide access to microbial diversity for drug discovery from natural products. Appl Environ Microbiol 69:49–55
Crosa JH, Walsh CT (2002) Genetics and assembly line enzymology of siderophore biosynthesis in bacteria. Microbiol Mol Biol Rev 66:223–249
Edwards DJ, Marquez BL, Nogle LM, McPhail K, Goeger DE, Roberts MA, Gerwick WH (2004) Structure and biosynthesis of the jamaicamides, new mixed polyketide-peptide neurotoxins from the marine cyanobacterium Lyngbya majuscula. Chem Biol 11:817–833
Ehrenreich IM, Waterbury JB, Webb EA (2005) Distribution and diversity of natural product genes in marine and freshwater cyanobacterial cultures and genomes. Appl Environ Microbiol 71:7401–7413
Faulkner DJ (2000) Marine pharmacology. Antonie Van Leeuwenhoek 77:135–145
Faulkner DJ (2002) Marine natural products. Nat Prod Rep 19:1–48
Firn RD, Jones CG (2000) The evolution of secondary metabolism-a unifying model. Mol Microbiol 37:989–994
Franzmann PD, Patterson BM, Power TR, Nichols PD, Davis GB (1996) Microbial biomass in a shallow, urban aquifer contaminated with aromatic hydrocarbons: analysis by phospholipid fatty acid content and composition. J Appl Bacteriol 80:617–625
Gillespie DE, Brady SF, Bettermann AD, Cianciotto NP, Liles MR, Rondon MR, Clardy J, Goodman RM, Handelsman J (2002) Isolation of antibiotics turbomycin A and B from a metagenomic library of soil microbial DNA. Appl Environ Microbiol 68:4301–4306
Ginolhac A, Jarrin C, Gillet B, Robe P, Pujic P, Tuphile K, Bertrand H, Vogel TM, Perriere G, Simonet P, Nalin R (2004) Phylogenetic analysis of polyketide synthase I domains from soil metagenomic libraries allows selection of promising clones. Appl Environ Microbiol 70:5522–5527
Gokhale RS, Tsuji SY, Cane DE, Khosla C (1999) Dissecting and exploiting intermodular communication in polyketide synthases. Science 284:482–485
Hoffmann D, Hevel JM, Moore RE, Moore BS (2003) Sequence analysis and biochemical characterization of the nostopeptolide A biosynthetic gene cluster from Nostoc sp. GSV224. Gene 311:171–180
Jimeno J, Lopez-Martin JA, Ruiz-Casado A, Izquierdo MA, Scheuer PJ, Rinehart K (2004) Progress in the clinical development of new marine derived anticancer compounds. Anticancer Drugs 15:321–329
Jungblut AD, Hawes I, Mountfort D, Hitzfeld B, Dietrich DR, Burns BP, Neilan BA (2005) Diversity within cyanobacterial mat communities in variable salinity meltwater ponds of McMurdo Ice Shelf. Antarctica Environ Microbiol 7:519–529
Larimer FW, Chain P, Hauser L, Lamerdin J, Malfatti S, Do L, Land ML, Pelletier DA, Beatty JT, Lang AS, Tabita FR, Gibson JL, Hanson TE, Bobst C, Torres JL, Peres C, Harrison FH, Gibson J, Harwood CS (2004) Complete genome sequence of the metabolically versatile photosynthetic bacterium Rhodopseudomonas palustris. Nat Biotechnol 22:55–61
Lin X, Zeng R (2007) Denaturing gradient gel electrophoresis (DGGE) analysis of microbial community in sewage contaminated soil from Zhongshan Station, Antarctic. Chin J Appl Environ Biol (in Chinese) (in press)
Luesch H, Hoffmann D, Hevel JM, Becker JE, Golakoti T, Moore RE (2003) Biosynthesis of 4-methylproline in cyanobacteria: cloning of nosE and nosF genes and biochemical characterization of the encoded dehydrogenase and reductase activities. J Org Chem 68:83–91
McDaniel R, Thamchaipenet A, Gustafsson C, Fu H, Betlach M, Ashley G (1999) Multiple genetic modifications of the erythromycin polyketide synthase to produce a library of novel “unnatural” natural products. Proc Natl Acad Sci USA 96:1846–1851
Moffitt MC, Neilan BA (2001) On the presence of peptide synthetase and polyketide synthase genes in the cyanobacterial genus Nodularia. FEMS Microbiol Lett 196:207–214
Moffitt MC, Neilan BA (2003) Evolutionary affiliations within the superfamily of ketosynthases reflect complex pathway associations. J Mol Evol 56:446–457
Moore BS, Kalaitzis JA, Xiang L (2005) Exploiting marine actinomycete biosynthetic pathways for drug discovery. Antonie Van Leeuwenhoek 87:49–57
Neilan BA, Dittmann E, Rouhiainen L, Bass RA, Schaub V, Sivonen K, Borner T (1999) Nonribosomal peptide synthesis and toxigenicity of cyanobacteria. J Bacteriol 181:4089–4097
Nicholson TP, Rudd BA, Dawson M, Lazarus CM, Simpson TJ, Cox RJ (2001) Design and utility of oligonucleotide gene probes for fungal polyketide synthases. Chem Biol 8:157–178
Nishizawa T, Asayama M, Fujii K, Harada K, Shirai M (1999) Genetic analysis of the peptide synthetase genes for a cyclic heptapeptide microcystin in Microcystis spp. J Biochem (Tokyo) 126:520–529
Pathom-Aree W, Stach JE, Ward AC, Horikoshi K, Bull AT, Goodfellow M (2006) Diversity of actinomycetes isolated from Challenger Deep sediment (10,898 m) from the Mariana Trench. Extremophiles 10:181–189
Paulsen IT, Press C, Ravel J, Kobayashi D, Myers GS, Mavrodi D, DeBoy RT, Seshadri R, Ren Q, Madupu R, Dodson RJ, Durkin S, Brinkac LM, Daugherty SC, Sullivan SA, Rosovitz M, Gwinn ML, Zhou L, Nelson WC, Weidman J, Watkins K, Tran K, Khouri HM, Pierson E, Pierson L III, Thomashow L, Loper J (2005) Complete genome sequence of the plant commensal Pseudomonas fluorescens Pf-5. Nat Biotechnol 23:873–878
Piel J, Hofer I, Hui D (2004) Evidence for a symbiosis island involved in horizontal acquisition of pederin biosynthetic capabilities by the bacterial symbiont of Paederus fuscipes beetles. J Bacteriol 186:1280–1286
Schirmer A, Gadkari R, Reeves CD, Ibrahim F, DeLong EF, Hutchinson CR (2005) Metagenomic analysis reveals diverse polyketide synthase gene clusters in microorganisms associated with the marine sponge Discodermia dissoluta. Appl Environ Microbiol 71:4840–4849
Schwarzer D, Marahiel MA (2001) Multimodular biocatalysts for natural product assembly. Naturwissenschaften 88:93–101
Sjöling S, Cowan DA (2003) High 16S rDNA bacterial diversity in glacial meltwater lake sediment, Bratina Island, Antarctica. Extremophiles 7:275–282
Stachelhaus T, Mootz HD, Marahiel MA (1999) The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem Biol 6:493–505
Staunton J, Weissman KJ (2001) Polyketide biosynthesis: a millennium review. Nat Prod Rep 18:380–416
Turgay K, Marahiel MA (1994) A general approach for identifying and cloning peptide synthetase genes. Pept Res 7:238–241
Watve MG, Tickoo R, Jog MM, Bhole BD (2001) How many antibiotics are produced by the genus Streptomyces? Arch Microbiol 176:386–390
Zeng R, Zhao J, Zhang R, Lin N (2005) Bacterial community in sediment from the Western Pacific “Warm Pool” and its relationship to environment. Sci China Ser D 48:282–290
Acknowledgments
This work was supported by Hi-tech Research and Development Program of China (2007AA091407) and National Natural Science Funds of China (No. 40406029).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhao, J., Yang, N. & Zeng, R. Phylogenetic analysis of type I polyketide synthase and nonribosomal peptide synthetase genes in Antarctic sediment. Extremophiles 12, 97–105 (2008). https://doi.org/10.1007/s00792-007-0107-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-007-0107-9