Abstract
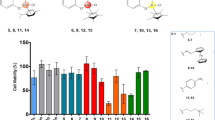
Novel ruthenium(III) complexes of general formula Na[RuCl2(L1−3-N,O)2] where L(1–3) denote deprotonated Schiff bases (HL1-HL3) derived from 5-substituted salicyladehyde and alkylamine (propyl- or butylamine) were prepared and characterized based on elemental analysis, mass spectra, infrared, electron spin/paramagnetic resonance (ESR/EPR) spectroscopy, and cyclovoltammetric study. Optimization of five isomers of complex C1 was done by DFT calculation. The interaction of C1–C3 complexes with DNA (Deoxyribonucleic acid) and BSA (Bovine serum albumin) was investigated by electron spectroscopy and fluorescence quenching. The cytotoxic activity of C1–C3 was investigated in a panel of four human cancer cell lines (K562, A549, EA.hy926, MDA-MB-231) and one human non-tumor cell line (MRC-5). Complexes displayed an apparent cytoselective profile, with IC50 values in the low micromolar range from 1.6 ± 0.3 to 23.0 ± 0.1 µM. Cisplatin-resistant triple-negative breast cancer cells MDA-MB-231 displayed the highest sensitivity to complexes, with Ru(III) compound containing two chlorides and two deprotonated N-propyl-5-chloro-salicylidenimine (hereinafter C1) as the most potent (IC50 = 1.6 µM), and approximately ten times more active than cisplatin (IC50 = 21.9 µM). MDA-MB-231 cells treated for 24 h with C1 presented with apoptotic morphology, as seen by acridine orange/ethidium bromide staining, while 48 h of treatment induced DNA fragmentation, and necrotic changes in cells, as seen by flow cytometry analysis. Drug-accumulation study by inductively coupled plasma mass spectrometry (ICP-MS) demonstrated markedly higher intracellular accumulation of C1 compared with cisplatin.
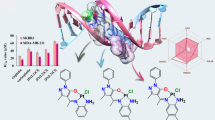
Graphical abstract



















Similar content being viewed by others
Data availability
The dataset generated and analyzed during the current study is available upon reasonable request.
References
Clarke MJ (2003) Ruthenium metallopharmaceuticals. Coord Chem Rev 236(1–2):209–233
Keppler B (1993) Metal complexes in cancer chemotherapy. VCH-Verlag, Weinheim, pp 1–42
Hartinger CG, Jakupec MA, Zorbas-Seifried S et al (2008) KP1019, a new redox-active anticancer agent—preclinical development and results of a clinical phase I study in tumor patients. Chem Biodivers 10:2140–2155. https://doi.org/10.1002/cbdv.200890195
Alessio E, Mestroni G, Bergamo A et al (2004) Ruthenium antimetastatic agents. Curr Top Med Chem 4(15):1525–1535. https://doi.org/10.2174/1568026043387421
Clarke MJ, Bitler S, Rennert D et al (1980) Reduction and subsequent binding of ruthenium ions catalyzed by subcellular components. J Inotg Biochem 12(1):79–87. https://doi.org/10.1016/S0162-0134(00)80045-8
Kahrović E, Zahirović A, Turkušić E et al (2016) A dinuclear ruthenium(II) Schiff base complex with dissimilar coordination: synthesis, characterization, and biological activity. Z Anorg Allg Chem 642(6):480–485. https://doi.org/10.1002/zaac.201600008
Ljubijankić N, Zahirović A, Turkušić E et al (2013) DNA binding properties of two ruthenium(III) complexes containing Schiff bases derived from salicylaldehyde: spectroscopic and electrochemical evidence of CT DNA intercalation. Croat Chem Acta 86(2):215–222. https://doi.org/10.5562/cca2216
Kahrović E, Zahirović A, Kraljević Pavelić S et al (2017) In vitro anticancer activity of binuclear Ru(II) complexes with Schiff bases derived from 5-substituted salicylaldehyde and 2-aminopyridine with notably low IC50 values. J Coord Chem 70(10):1683–1697. https://doi.org/10.1080/00958972.2017.1308503
Muzika V, Custovic S, Alicelebic S et al (2019) Dinuclear ruthenium(II) Schiff base complex: a first in vivo study in Swiss albino mice. Bratisl Med J 120(01):26–34. https://doi.org/10.4149/BLL_2019_004
Perdew JP (1991) Electronic Structure of Solids ‘91,. In: Ziesche P, Eschrig H (eds) 11, and subsequent references. Akademie Verlag, Berlin
Adamo C, Barone V (1998) Exchange functionals with improved long-range behavior and adiabatic connection methods without adjustable parameters: the mPW and mPW1PW models. J Chem Phys 108:664–675. https://doi.org/10.1063/1.475428
Hay PJ, Wadt WR (1985) Ab initio effective core potentials for molecular calculations—potentials for the transition-metal atoms Sc to Hg. J Chem Phys 82:270–283. https://doi.org/10.1063/1.448799
Petersson GA, Al-Laham MA (1991) A complete basis set model chemistry. II. Open-shell systems and the total energies of the first-row atoms. J Chem Phys 94:6081–6090. https://doi.org/10.1063/1.460447
Cossi M, Rega N, Scalmani G et al (2003) Energies, structures, and electronic properties of molecules in solution with the C-PCM solvation model. J Comput Chem 24:669–681. https://doi.org/10.1002/jcc.10189
Miertuš S, Scrocco E, Tomasi J (1981) Electrostatic interaction of a solute with a Continuum. A direct utilization of ab initio molecular potentials for the prevision of solvent effects. Chem Phys 55:117–129. https://doi.org/10.1016/0301-0104(81)85090-2. (And subsequent references)
Frisch MJ et al. (2019) Gaussian 16, Revision C.01. Gaussian, Inc., Wallingford
https://www.sdsc.edu/services/hpc/expanse/index.html. Accessed Dec 2022
Rosenberg B, VanCamp L, Trosko JE et al (1969) Platinum compounds: a new class of potent antitumour agents. Nature 222:385–386. https://doi.org/10.1038/222385a0
Supino R (1995) Methods in molecular biology. In: O’Hare S, Atterwill S (eds) In vitro toxicity testing protocols, vol 43. C. K. Humana Press, Clifton, pp 137–149
Ormerod MG (1994) Analysis of DNA-general methods. In: Ormerod MG (ed) In flow cytometry, a practical approach. Oxford University Press, New York, pp 119–125
Spector DL, Goldman RD, Leinwand LA (1998) Culture and biochemical analysis of cells. In: Cells: A Laboratory Manual, vol 1. Cold Spring Harbor Laboratory Press, New York
Slee EA, Zhu H, Chow SC et al (1996) Benzyloxycarbonyl-Val-Ala-Asp (OMe) fluoromethylketone (Z-VAD.FMK) inhibits apoptosis by blocking the processing of CPP32. Biochem J 315(1):21–24. https://doi.org/10.1042/bj3150021
Dodo K, Katoh M, Shimizu T (2005) Inhibition of hydrogen peroxide-induced necrotic cell death with 3-amino-2-indolylmaleimide derivatives. Bioorg Med Chem Lett 15(12):3114–3118. https://doi.org/10.1016/j.bmcl.2005.04.016
Pavlović M, Nikolić S, Gligorijević N et al (2019) New organoruthenium compounds with pyrido[2′,3′:5,6]pyrazino[2,3-f][1, 10]phenanthroline: synthesis, characterization, cytotoxicity, and investigation of mechanism of action. J Biol Inorg Chem 24(2):297–310. https://doi.org/10.1007/s00775-019-01647-4
Eruslanov E, Kusmartsev S (2009) Identification of ROS using oxidized DCFDA and flow-cytometry. In: Armstrong D (ed) Advanced protocols in oxidative stress II, methods in molecular biology, vol 594. D. Humana Press, Totowa, pp 57–72. https://doi.org/10.1007/978-1-60761-411-1_4
Kahrović E (2014) Ruthenium compounds with Schiff bases: design and promising application of salicylideneimine complexes. In: Keller GP (ed) Ruthenium synthesis, physicochemical properties and applications. NOVA Science Publishers, New York, pp 269–284
Vargiu AV, Robertazzi A, Magistrato A et al (2008) The hydrolysis mechanism of the anticancer ruthenium drugs NAMI-A and ICR investigated by DFT−PCM calculations. J Phys Chem B 112(14):4401–4409. https://doi.org/10.1021/jp710078y
Kannan S, Ramesh R (2006) Synthesis, characterization, catalytic oxidation and biological activity of ruthenium(III) Schiff base complexes derived from 3-acetyl-6-methyl-2H-pyran-2,4(3H)-dione. Polyhedron 25(16):3095–3103. https://doi.org/10.1016/j.poly.2006.05.042
Sivagamasundari M, Ramesh R (2007) Luminescent property and catalytic activity of Ru(II) carbonyl complexes containing N, O donor of 2-hydroxy-1-naphthylideneimines. Spectrochim Acta A Mol Biomol Spectrosc 67(1):256–262. https://doi.org/10.1016/j.saa.2006.03.017
Biswas N, Saha S, Zangrando E et al (2021) Catecholase-like activity and theoretical study in solid state of a new Ru(III)-Schiff base complex. Acta Chim Slov 68(1):212–221. https://doi.org/10.17344/acsi.2020.6379
Chen C, Ji J, Wang CJ (2020) Synthesis, characterization and crystal structures of half-sandwich ruthenium complexes with bidentate chiral Schiff-base ligands. J Organomet Chem 910:121129. https://doi.org/10.1016/j.jorganchem.2020.121129
Wu F, Wang CJ, Lin H et al (2018) Syntheses, characterization and crystal structures of ruthenium(II)/(III) complexes with tridentate salicylaldiminato ligands. J Coord Chem 71(2):219–230. https://doi.org/10.1080/00958972.2017.1423476
Zahirović A, Žilić D, Kraljević Pavelić S et al (2019) Type of complex-BSA binding forces affected by different coordination modes of alliin in novel water-soluble ruthenium complexes. New J Chem 43(15):5791–5804. https://doi.org/10.1039/C9NJ00826H
Pushkar Y, Moonshiram D, Purohit V et al (2014) Spectroscopic analysis of catalytic water oxidation by [RuII(bpy)(tpy)H2O]2+ suggests that RuV=O is not a rate-limiting intermediate. J Am Chem Soc 136(34):11938–11945. https://doi.org/10.1021/ja506586b
Planas N, Vigara L, Cady C et al (2011) Electronic structure of oxidized complexes derived from cis-[RuII(bpy)2(H2O)2]2+and its photoisomerization mechanism. Inorg Chem 50(21):11134–11142. https://doi.org/10.1021/ic201686c
Daniel Q, Huang P, Fan T et al (2017) Rearranging from 6- to 7-coordination initiates the catalytic activity: an EPR study on a Ru-bda water oxidation catalyst. Coord Chem Rev 346:206–215. https://doi.org/10.1016/j.ccr.2017.02.019
Stoll S, Schweige A (2006) EasySpin, a comprehensive software package for spectral simulation and analysis in EPR. J Magn Reson 178(1):42–55. https://doi.org/10.1016/j.jmr.2005.08.013
Szymańska B, Skrzypek D, Kovala-Demertzi D et al (2006) Synthesis and spectroscopic study of copper(II) and manganese(II) complexes with pipemidic acid. Spectrochim Acta A Mol Biomol Spectrosc 63(3):518–523. https://doi.org/10.1016/j.saa.2005.05.038
Munshi P, Samanta R, Kumar LG (1999) Paramagnetic ruthenium(III) ortho-metallated complexes. Synthesis, spectroscopic and redox properties. J Organomet 586(2):176–183. https://doi.org/10.1016/S0022-328X(99)00261-2
Webb MI, Walsby CJ (2013) EPR as a probe of the intracellular speciation of ruthenium(III) anticancer compounds. Metallomics 5(12):1624–1633. https://doi.org/10.1039/C3MT00090G
Canevali C, Chiodini N, Morazzoni F et al (2000) Electron paramagnetic resonance characterization of ruthenium-dispersed tin oxide obtained by sol–gel and impregnation methods. J Mater Chem 10(3):773–778. https://doi.org/10.1039/A907947E
Schluga P, Hartinger CG, Egger A et al (2006) Redox behavior of tumor-inhibiting ruthenium(III) complexes and effects of physiological reductants on their binding to GMP. Dalton Trans 6(14):1796–1802. https://doi.org/10.1039/b511792e
Mandler D (2010) Anal bioanal chem. In: Scholz F (ed) Electroanalytical methods Guide to experiments and applications, vol 398(7-8), 2nd edn. Springer, Berlin, pp 2771–2772. https://doi.org/10.1007/s00216-010-4195-5
Jabłońska-Wawrzycka A, Rogala P, Michałkiewicz S et al (2013) Ruthenium complexes in different oxidation states: synthesis, crystal structure, spectra and redox properties. Dalton Trans 42(17):6092–6101. https://doi.org/10.1039/C3DT32214A
Reisner E, Arion VB, Guedes da Silva MFC et al (2004) Tuning of redox potentials for the design of ruthenium anticancer drugs—an electrochemical study of [trans-RuCl4L(DMSO)]-and [trans-RuCl4L2]-complexes, where L = Imidazole, 1,2,4-Triazole, Indazole. Inorg Chem 43(22):7083–7093. https://doi.org/10.1021/ic049479c
Mestroni G, Alessio E, Sava G et al (1994) Water-soluble ruthenium(III)-dimethyl sulfoxide complexes: chemical behaviour and pharmaceutical properties. Met Based Drugs 1(1):41–63. https://doi.org/10.1155/MBD.1994.41
Wiśniewska J, Fandzloch M, Muzioł T et al (2019) The hydrolysis of a ruthenium(III) complex with triazolopyrimidine ligands and mechanistic insights into its anticancer activity. Inorg Chem Commun 109:107567. https://doi.org/10.1016/j.inoche.2019.107567
Azam M, Hussain Z, Warad I et al (2012) Novel Pd(II)–salen complexes showing high in vitro anti-proliferative effects against human hepatoma cancer by modulating specific regulatory genes. Dalton Trans 41(35):10854–10864. https://doi.org/10.1039/C2DT31143G
Long EC, Barton JK (1990) On demonstrating DNA intercalation. Acc Chem Res 23(9):271–273. https://doi.org/10.1021/ar00177a001
Cory M, McKee DD, Kagan J et al (1985) Design, synthesis, and DNA binding properties of bifunctional intercalators. Comparison of polymethylene and diphenyl ether chains connecting phenanthridine. J Am Chem Soc 107(8):2528–2536. https://doi.org/10.1021/ja00294a054
Brabec V, Kasparkova J (2018) Ruthenium coordination compounds of biological and biomedical significance. DNA binding agents. Coord Chem Rev 376:75–94. https://doi.org/10.1016/j.ccr.2018.07.012
Patel MN, Parmar PA, Gandhi DS (2011) Synthesis, characterization and DNA binding and cleavage properties of ruthenium(II) complexes with various polypyridyls. J Enzyme Inhib Med Chem 26(5):734–741. https://doi.org/10.3109/14756366.2011.570007
Jiang CW, Chao H, Li H, Ji LN (2003) Syntheses, characterization and DNA-binding studies of ruthenium(II) terpyridine complexes: [Ru(tpy)(PHBI)]2+ and [Ru(tpy)(PHNI)]2+. J Inorg Biochem 93(3–4):247–255. https://doi.org/10.1016/s0162-0134(02)00577-9
Yin HJ, Zhang AG, Gao LH et al (2019) (2019) DNA groove-binding and acid-base properties of a Ru(II) complex containing anthryl moieties. Nucleosides Nucleotides Nucleic Acids. https://doi.org/10.1080/15257770.2019.1669804
Ramezanpour A, Karami K, Kharaziha M et al (2021) A mononuclear PdII complex with Naphcon; crystal structure, experimental and computational studies of the interaction with DNA/BSA and evaluation of anticancer activity. Polyhedron 206:115333. https://doi.org/10.1016/j.poly.2021.115333
Li Y, Yang Z, Zhou M et al (2017) Synthesis and crystal structure of new monometallic Ni(ii) and Co(ii) complexes with an asymmetrical aroylhydrazone: effects of the complexes on DNA/protein binding property, molecular docking, and in vitro anticancer activity. RSC Adv 7(78):49404–49422. https://doi.org/10.1039/C7RA10283F
Senthil RD, Bhuvanesh NSP, Natarajan K (2011) Effect of N(4)-phenyl substitution in 2-Oxo-1,2-dihydroquinoline-3-carbaldehyde semicarbazones on the structure, DNA/protein interaction, and antioxidative and cytotoxic activity of Cu(II) complexes. Inorg Chem 50(24):12852–12866. https://doi.org/10.1021/ic2020308
Grigoryan KR, Aznauryan AG, Bagramyan NA et al (2008) Spectroscopic determination of binding between human serum albumin and a platinum(II) dimethylsulfoxide complex. J Appl Spectrosc 75(4):593–596. https://doi.org/10.1007/s10812-008-9070-1
Grigoryan KR, Kazaryan AG (2009) Interaction of dimethylsulfoxide and diethylsulfoxide complexes of platinum(II) with bovine serum albumin. JAppl Spectrosc 76(4):607–610. https://doi.org/10.1007/s10812-009-9241-8
Maikoo S, Chakraborty A, Vukea N et al (2021) Ruthenium complexes with mono- or bis-heterocyclic chelates: DNA/BSA binding, antioxidant and anticancer studies. J Biomol Struct Dyn 39:4077–4088. https://doi.org/10.1080/07391102.2020.1775126
Sauer M, Hofkens J, Enderlein J (2011) Basic principles of fluorescence spectroscopy. Wiley, New York. https://doi.org/10.1002/9783527633500.ch1
Lakowicz JR (2006) Principles of fluorescence spectroscopy. Springer, Berlin
Pauzi AZ, Yeap SK, Abu N et al (2016) Combination of cisplatin and bromelain exerts synergistic cytotoxic effects against breast cancer cell line MDA-MB-231 in vitro. Chin Med 15(11):46. https://doi.org/10.1186/s13020-016-0118-5
Nikolic S, Opsenica D, Filipović V et al (2015) Strong in vitro cytotoxic potential of new ruthenium−cymene complexes. Organometallics 34:3464–3473. https://doi.org/10.1021/acs.organomet.5b00041
Barr MP, Gray SG, Hoffmann AC et al (2013) Generation and characterisation of cisplatin-resistant non-small cell lung cancer cell lines displaying a stem-like signature. PLoS One 8(1):e54193. https://doi.org/10.1371/journal.pone.0054193
Baharuddin P, Satar N, Fakiruddin KS et al (2016) Curcumin improves the efficacy of cisplatin by targeting cancer stem-like cells through p21 and cyclin D1-mediated tumour cell inhibition in non-small cell lung cancer cell lines. Oncol Rep 35:13–25. https://doi.org/10.3892/or.2015.4371
Ban KA, Godellas CV (2014) Epidemiology of breast cancer. Surg Oncol Clin N Am 23(3):409–422. https://doi.org/10.1016/j.soc.2014.03.011
Hongthong K, Ratanaphan A (2016) BRCA1-associated triple-negative breast cancer and potential treatment for ruthenium-based compounds. Curr Cancer Drug Target 16(7):606–617. https://doi.org/10.2174/1568009616666160203113957
Pavlović M, Tadić A, Gligorijević N et al (2020) Synthesis, chemical characterization, PARP inhibition, DNA binding and cellular uptake of novel ruthenium(II)-arene complexes bearing benzamide derivatives in human breast cancer cells. J Inorg Biochem. https://doi.org/10.1016/j.jinorgbio.2020.111155
Tadić A, Poljarević J, Krstić M et al (2018) Ruthenium-arene complexes with NSAIDs: synthesis, characterization and bioactivity. New J Chem 42(4):3001–3019. https://doi.org/10.1039/c7nj04416j
Galluzzi L, Senovilla L, Vitale I et al (2012) Molecular mechanisms of cisplatin resistance. Oncogene 31(15):1869–1883. https://doi.org/10.1038/onc.2011.384
Huang X, Halicka HD, Traganos F et al (2005) Cytometric assessment of DNA damage in relation to cell cycle phase and apoptosis. Cell Prolif 38(4):223–243. https://doi.org/10.1111/j.1365-2184.2005.00344.x
Kumari S, Badana AK, Murali Mohan G et al (2018) Reactive oxygen species: a key constituent in cancer survival. Biomark Insights 13:1–9. https://doi.org/10.1177/1177271918755
Acknowledgements
This work was supported by the Federal Ministry of Education and Science of Bosnia and Herzegovina (Grant no. 05-39-2619-1/18) and by the Ministry of Education, Science and Technological Development of the Republic of Serbia (Grant no. III 41026 and Agreement no. 451-03-68/2022-14/200043). Computational resources in this study were procured through the NSF XSEDE grant (MCB140083) awarded to P.I. We thank Biljana Dojčinović for the ICP-MS measurements, Adnan Zahirović and Irnesa Osmanković for their support in measuring BSA and DNA interactions.
Author information
Authors and Affiliations
Contributions
EK: design and development of compounds, interactions, supervision, writing—original draft, visualization, writing—review of chemical part. NL: syntheses. DŽ and JJ: X-band ESR spectroscopy, measurements and writing. P-PI: electronic structure calculations. MP: methodology, formal analysis and investigation, visualization, writing—original draft, writing—review and editing (biological part). SA: conceptualization, writing—review and editing (biological part). SR: conceptualization, funding acquisition, supervision. SG-Š: conceptualization, writing—review and editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pavlović, M., Kahrović, E., Aranđelović, S. et al. Tumor selective Ru(III) Schiff bases complexes with strong in vitro activity toward cisplatin-resistant MDA-MB-231 breast cancer cells. J Biol Inorg Chem 28, 263–284 (2023). https://doi.org/10.1007/s00775-023-01989-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-023-01989-0