Abstract
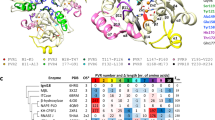
PqqB is an enzyme involved in the biosynthesis of pyrroloquinoline quinone and a distal member of the metallo-β-lactamase (MBL) superfamily. PqqB lacks two residues in the conserved signature motif HxHxDH that makes up the key metal-chelating elements that can bind up to two metal ions at the active site of MBLs and other members of its superfamily. Here, we report crystal structures of PqqB bound to Mn2+, Mg2+, Cu2+, and Zn2+. These structures demonstrate that PqqB can still bind metal ions at the canonical MBL active site. The fact that PqqB can adapt its side chains to chelate a wide spectrum of metal ions with different coordination features on a uniform main chain scaffold demonstrates its metal-binding plasticity. This plasticity may provide insights into the structural basis of promiscuous activities found in ensembles of metal complexes within this superfamily. Furthermore, PqqB belongs to a small subclass of MBLs that contain an additional CxCxxC motif that binds a structural Zn2+. Our data support a key role for this motif in dimerization.





Similar content being viewed by others
Abbreviations
- PQQ:
-
Pyrroloquinoline quinone
- MBL:
-
Metallo-β-lactamase
- Pp:
-
Pseudomonas putida
- IPTG:
-
Isopropyl β-d-1-thiogalactopyranoside
- SEC:
-
Size exclusion chromatography
- PDB:
-
Protein data bank
- Ec:
-
Escherichia coli
References
Hartmann C, Klinman JP (1988) BioFactors 1:41–49
Magnusson OT, Toyama H, Saeki M, Schwarzenbacher R, Klinman JP (2004) J Am Chem Soc 126:5342–5343
Chowanadisai W, Bauerly KA, Tchaparian E, Wong A, Cortopassi GA, Rucker RB (2010) J Biol Chem 285:142–152
Singh AK, Pandey SK, Saha G, Gattupalli NK (2015) Exp Gerontol 66:1–9
Ohwada K, Takeda H, Yamazaki M, Isogai H, Nakano M, Shimomura M, Fukui K, Urano S (2008) J Clin Biochem Nutr 42:29–34
Yang C, Yu L, Kong L, Ma R, Zhang J, Zhu Q, Zhu J, Hao D (2014) PLoS One 9:e109502
Shen YQ, Bonnot F, Imsand EM, RoseFigura JM, Sjolander K, Klinman JP (2012) Biochemistry 51:2265–2275
Metlitzky M, Puehringer S, Fisher SJ (2012) J Biophys Chem 3:206–210
Bebrone C (2007) Biochem Pharmacol 74:1686–1701
Baier F, Tokuriki N (2014) J Mol Biol 426:2442–2456
Podzelinska K, He SM, Wathier M, Yakunin A, Proudfoot M, Hove-Jensen B, Zechel DL, Jia ZC (2009) J Biol Chem 284:17216–17226
He SM, Wathier M, Podzelinska K, Wong M, McSorley FR, Asfaw A, Hove-Jensen B, Jia ZC, Zechel DL (2011) Biochemistry 50:8603–8615
Kal S, Que L (2017) J Biol Inorg Chem 22:339–365
King D, Strynadka N (2011) Protein Sci 20:1484–1491
Velterop JS, Sellink E, Meulenberg JJ, David S, Bulder I, Postma PW (1995) J Bacteriol 177:5088–5098
Klinman JP, Bonnot F (2014) Chem Rev 114:4343–4365
Kabsch W (2010) Acta Crystallogr D Biol Crystallogr 66:125–132
Otwinowski Z, Minor W (1997) Method Enzymol 276:307–326
McCoy AJ, Grosse-Kunstleve RW, Adams PD, Winn MD, Storoni LC, Read RJ (2007) J Appl Crystallogr 40:658–674
Emsley P, Cowtan K (2004) Acta Crystallogr D Biol Crystallogr 60:2126–2132
Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung LW, Kapral GJ, Grosse-Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, Zwart PH (2010) Acta Crystallogr D Biol Crystallogr 66:213–221
Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AG, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS (2011) Acta Crystallogr D Biol Crystallogr 67:235–242
Brunger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW, Jiang JS, Kuszewski J, Nilges M, Pannu NS, Read RJ, Rice LM, Simonson T, Warren GL (1998) Acta Crystallogr D Biol Crystallogr 54:905–921
Chen VB, Arendall WB 3rd, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, Murray LW, Richardson JS, Richardson DC (2010) Acta Crystallogr D Biol Crystallogr 66:12–21
Krissinel E, Henrick K (2007) J Mol Biol 372:774–797
Schrodinger LLC (2015) The PyMOL molecular graphics system, version 1.8.05
Valladares MH, Felici A, Weber G, Adolph HW, Zeppezauer M, Rossolini GM, Amicosante G, Frere JM, Galleni M (1997) Biochemistry 36:11534–11541
Bebrone C, Delbruck H, Kupper MB, Schlomer P, Willmann C, Frere JM, Fischer R, Galleni M, Hoffmann KMV (2009) Antimicrob Agents Ch 53:4464–4471
Garau G, Bebrone C, Anne C, Galleni M, Frere JM, Dideberg O (2005) J Mol Biol 345:785–795
Baier F, Copp JN, Tokuriki N (2016) Biochemistry 55:6375–6388
Green VL, Verma A, Owens RJ, Phillips SEV, Carr SB (2011) Acta Crystallogr F 67:1160–1164
Guo Y, Wang J, Niu GJ, Shui WQ, Sun YN, Zhou HG, Zhang YZ, Yang C, Lou ZY, Rao ZH (2011) Protein Cell 2:384–394
Zhang HM, Hao Q (2011) FASEB J 25:2574–2582
Kim Y, Tesar C, Mire J, Jedrzejczak R, Binkowski A, Babnigg G, Sacchettini J, Joachimiak A (2011) PLoS One 6:e24621
Baier F, Chen J, Solomonson M, Strynadka NCJ, Tokuriki N (2015) ACS Chem Biol 10:1684–1693
Acknowledgements
This research was supported in part by Grants GM-66569 (C.M.W.) and GM-118117 (J.P.K.) from the National Institutes of Health. The work used Advanced Photon Source beamlines 19-BM, 19-ID, and GM/CA, the latter funded in whole or in part with Federal funds from the National Cancer Institute (ACB-12002) and the National Institute of General Medical Sciences (AGM-12006). The Advanced Photon Source is a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tu, X., Latham, J.A., Klema, V.J. et al. Crystal structures reveal metal-binding plasticity at the metallo-β-lactamase active site of PqqB from Pseudomonas putida . J Biol Inorg Chem 22, 1089–1097 (2017). https://doi.org/10.1007/s00775-017-1486-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-017-1486-8