Abstract
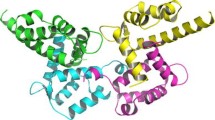
The homodimeric structure of human S100A16 in the apo state has been obtained both in the solid state and in solution, resulting in good agreement between the structures with the exception of two loop regions. The homodimeric solution structure of human S100A16 was also calculated in the calcium(II)-bound form. Differently from most S100 proteins, the conformational rearrangement upon calcium binding is minor. This characteristic is likely to be related to the weak binding affinity of the protein for the calcium(II) ions. In turn, this is ascribed to the lack of the glutamate residue at the end of the S100-specific N-domain binding site, which in most S100 proteins provides two important side chain oxygen atoms as calcium(II) ligands. Furthermore, the presence of hydrophobic interactions stronger than for other S100 proteins, present in the closed form of S100A16 between the third and fourth helices, likely make the closed structure of the second EF-hand particularly stable, so even upon calcium(II) binding such a conformation is not disrupted.










Similar content being viewed by others
References
Santamaria-Kisiel L, Rintala-Dempsey AC, Shaw GS (2006) Biochem J 396:201–214
Donato R (2003) Microsc Res Tech 60:540–551
Donato R (1986) Cell Calcium 7:123–145
Nelson MR, Chazin WJ (1998) Biometals 11:297–318
Brodersen DE, Etzerodt M, Madsen P, Celis JE, Thøgersen HC, Nyborg J, Kjeldgaard M (1998) Structure 6:477–489
Wilder PT, Baldisseri DM, Udan R, Vallely KM, Weber DJ (2003) Biochemistry 42:13410–13421
Randazzo A, Acklin C, Schafer BW, Heizmann CW, Chazin WJ (2001) Biochem Biophys Res Commun 288:462–467
Brodersen DE, Nyborg J, Kjeldgaard M (1999) Biochemistry 38:1695–1704
Moroz OV, Burkitt W, Wittkowski H, He W, Ianoul A, Novitskaya V, Xie J, Polyakova O, Lednev IK, Shekhtman A, Derrick PJ, Bjoerk P, Foell D, Bronstein IB (2009) BMC Biochem 10:11
Hwang JJ, Park MH, Choi SY, Koh JY (2005) J Biol Chem 280:11995–12001
Yu WH, Fraser PE (2001) J Neurosci 21:2240–2246
Marenholz I, Heizmann CW (2004) Biochem Biophys Res Commun 313:237–244
Sturchler E, Cox JA, Durussel I, Weibel M, Heizmann CW (2006) J Biol Chem 281:38905–38917
Ridinger K, Ilg EC, Niggli FK, Heizmann CW, Schäfer BW (1998) Biochim Biophys Acta 1448:254–263
Leslie AGW (1991) In: Moras D, Podjarny AD, Thierry J-C (eds) Molecular data processing. Oxford University Press, Oxford
Evans PR (1993) Proceedings of the CCP4 study weekend. In: Sawyer L, Isaacs N, Bailey S (eds) Data collection and processing, pp 114–122
Collaborative Computational Project N (1994) Acta Crystallogr D50:760–763
Schneider TR, Sheldrick GM (2002) Acta Crystallogr D 58:1772–1779
Sheldrick GM (2008) Acta Crystallogr A 64:112–122
Vonrhein C, Blanc E, Roversi P, Bricogne G (2007) Methods Mol Biol 364:215–230
Bricogne G, Vonrhein C, Flensburg C, Schiltz M, Paciorek W (2003) Acta Crystallogr D 59:2023–2030
Perrakis A, Morris RJH, Lamzin VS (1999) Nat Struct Biol 6:458–463
Cowtan K (2006) Acta Crystallogr D 62:1002–1011
McRee DE (1999) J Struct Biol 125:156–165
Murshudov GN, Vagin AA, Dodson EJ (1997) Acta Crystallogr D53:240–255
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) J Appl Crystallogr 26:283–291
Keller R (2004) The computer aided resonance assignment tutorial. CANTINA, Goldau
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) J Biomol NMR 44:213–223
Guntert P (2004) Methods Mol Biol 278:353–378
Herrmann T, Güntert P, Wüthrich K (2002) J Mol Biol 319:209–227
Case DA, Darden TA, Cheatham TE, Simmerling CL, Wang J, Duke RE, Luo R, Merz KM, Wang B, Pearlman DA, Crowley M, Brozell S, Tsui V, Gohlke H, Mongan J, Hornak V, Cui G, Beroza P, Schafmeister CE, Caldwell JW, Ross WS, Kollman PA (2008) AMBER 10. University of California, San Francisco
Laskowski RA, Rullmann JAC, MacArthur MW, Kaptein R, Thornton JM (1996) J Biomol NMR 8:477–486
Vriend G (1990) J Mol Graphics 8:52–56
Kay LE, Torchia DA, Bax A (1989) Biochemistry 28:8972–8979
Barbato G, Ikura M, Kay LE, Pastor RW, Bax A (1992) Biochemistry 31:5269–5278
Goddard TD, Kneller DG (2000) SPARKY 3. University of California, San Francisco
Lipari G, Szabo A (1982) J Am Chem Soc 104:4546–4559
Dosset P, Hus JC, Marion D, Blackledge M (2001) J Biomol NMR 20:223–231
Garcia de la Torre JG, Huertas ML, Carrasco B (2000) J Magn Reson 147:138–146
Inman KG, Baldisseri DM, Miller KE, Weber DJ (2001) Biochemistry 40:3439–3448
Zhukov I, Ejchart A, Bierzynski A (2008) Biochemistry 47:640–650
Dutta K, Cox CJ, Basavappa R, Pascal SM (2008) Biochemistry 47:7637–7647
Bertini I, Fragai M, Luchinat C, Parigi G (2000) Magn Reson Chem 38:543–550
Arnesano F, Banci L, Bertini I, Fantoni A, Tenori L, Viezzoli MS (2005) Angew Chem Int Ed 44:6341–6344
Bertini I, Fragai M, Luchinat C, Melikian M, Mylonas E, Sarti N, Svergun D (2009) J Biol Chem 284:12821–12828
Bertini I, Calderone V, Fragai M, Jaiswal R, Luchinat C, Melikian M, Mylonas E, Svergun D (2008) J Am Chem Soc 130:7011–7021
Bertini I, Gupta YK, Luchinat C, Parigi G, Schlörb C, Schwalbe H (2005) Angew Chem Int Ed 44:2223–2225
Luchinat C, Parigi G (2007) J Am Chem Soc 129:1055–1064
Bertini I, Dasgupta S, Hu X, Karavelas T, Luchinat C, Parigi G, Yuan J (2009) J Biol Inorg Chem 14:1097–1107
Brüschweiler R (2003) Curr Opin Struct Biol 13:175–183
Bernadò P, Garcìa de la Torre J, Pons M (2002) J Biomol NMR 23:139–150
Smith SP, Shaw GS (1998) Structure 6:211–222
Otterbein L, Kordowska J, Witte-Hoffmann C, Wang CL, Dominguez R (2002) Structure 10:557–567
Drohat AC, Baldisseri DM, Rustandi RR, Weber DJ (1998) Biochemistry 37:2729–2740
Babini E, Bertini I, Capozzi F, Luchinat C, Quattrone A, Turano M (2005) J Proteome Res 4:1961–1971
Marenholz I, Heizmann CW, Fritz G (2004) Biochem Biophys Res Commun 322:1111–1122
Maler L, Sastry M, Chazin WJ (2002) J Mol Biol 317:279–290
Bhattacharya S, Chazin WJ (2003) Structure 11:738–739
Gerke V, Weber K (1985) EMBO J 4:2917–2920
Rety S, Sopkova J, Renouard M, Osterloh D, Gerke V, Tabaries S, Russo-Marie F, Lewit-Bentley A (1999) Nat Struct Biol 6:89–95
Kube E, Becker T, Weber K, Gerke V (1992) J Biol Chem 267:14175–14182
Rescher U, Gerke V (2008) Pflugers Arch J Physiol 455:575–582
Smith SP, Shaw GS (1998) Structure 6:211–222
Kilby PM, Van Eldik LJ, Roberts GC (1996) Structure 4:1041–1052
Jacob J, Duclohier H, Cafiso DS (1999) Biophys J 76:1367–1376
Richardson JS (1981) Adv Protein Chem 34:167–339
MacArthur MW, Thornton JM (1991) J Mol Biol 218:397–412
Blaber M, Zhang XJ, Matthews BW (1993) Science 260:1637–1640
Koradi R, Billeter M, Wüthrich K (1996) J Mol Graphics 14:51–55
Acknowledgments
This work was supported by MIUR-FIRB contracts RBLA032ZM7 and RBIP06LSS2, and by European Commission contracts EU-NMR 026145 and SPINE2-COMPLEXES 031220.
Author information
Authors and Affiliations
Corresponding author
Additional information
An interactive 3D complement page in Proteopedia is available at: http://proteopedia.org/wiki/index.php/Journal:JBIC:3.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Babini, E., Bertini, I., Borsi, V. et al. Structural characterization of human S100A16, a low-affinity calcium binder. J Biol Inorg Chem 16, 243–256 (2011). https://doi.org/10.1007/s00775-010-0721-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-010-0721-3