Abstract
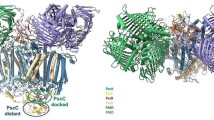
A Cu(I) metallochaperone, Atx1, interacts with the amino-terminal domain of a Cu(I)-transporting ATPase, PacSN, but not with a domain of related Zn-transporting ATPase, ZiaAN in Synechocystis PCC 6803. This is thought to prevent ZiaAN from acquiring Cu(I), which it binds more tightly than Zn. Solution structures of Atx1, PacSN, and the heterodimer were previously described. Here we report solution structural studies of the ZiaAN soluble domain. Apo-ZiaAN has a typical ferredoxin-like fold followed by an atypical 34 residues of unstructured polypeptide containing a His7 motif. ZiaAN competes with the metallochromic indicator 4-(2-pyridylazo)resorcinol for 1 equiv of Zn, which can be displaced by thiol-modifying p-mercuriphenylsulfonic acid, establishing that a high-affinity site involves thiols of the CXXC motif within the ferredoxin-like fold. A single equivalent of Zn affects nuclear magnetic resonance signals arising from the CXXC motif as well as all seven His residues. The presence of NMR-line broadening in both sites implies that Zn1-ZiaAN undergoes exchange phenomena, consistent with CXXC-bound Zn coincidentally sampling various His ligands. These Zn-dependent dynamic changes could either aid metal transfer or alter intramolecular interactions. No formation of Atx1–Cu(I)–ZiaAN heterodimers was observed, and in the presence of equimolar ZiaAN and PacSN, only Atx1–Cu(I)–PacSN complexes were detected. Residues flanking the CXXC motif of PacSN (R13-ASS20) differ in charge and bulk from those of ZiaAN (D18-KLK25) and make contacts in the Atx1–Cu(I)–PacSN complex. Crucially, swapping these residues flanking the CXXC motifs of ZiaAN and PacSN reciprocally swaps partner choice by Atx1. These few residues of the two ATPases have diverged during evolution to bias Atx1 interactions in favor of PacSN rather than ZiaAN.








Similar content being viewed by others
Abbreviations
- HSQC:
-
Heteronuclear single quantum coherence
- NMR:
-
Nuclear magnetic resonance
- NOE:
-
Nuclear Overhauser effect
- NOESY:
-
Nuclear Overhauser effect spectroscopy
- PAR:
-
4-(2-Pyridylazo)resorcinol
References
Lutsenko S, Kaplan JH (1995) Biochemistry 34:15607–15613
Solioz M, Vulpe C (1996) Trends Biochem Sci 21:237–241
Nucifora G, Chu L, Misra TK, Silver S (1989) Proc Natl Acad Sci USA 86:3544–3548
Silver S, Phung LT (1996) Annu Rev Microbiol 50:753–789
Gatti D, Mitra B, Rosen BP (2000) J Biol Chem 275:34009–34012
Arnesano F, Banci L, Bertini I, Ciofi-Baffoni S, Molteni E, Huffman DL, O’Halloran TV (2002) Genome Res 12:255–271
Rutherford JC, Cavet JS, Robinson NJ (1999) J Biol Chem 274:25827–25832
Williams LE, Mills RF (2005) Trends Plant Sci 10:491–502
Bartee MY, Lutsenko S (2007) Biometals 20:627–637
Argüello JM (2003) J Membr Biol 195:93–108
Mandal AK, Arguello JM (2003) Biochemistry 42:11040–11047
Banci L, Bertini I, Ciofi-Baffoni S, Finney LA, Outten CE, O’Halloran TV (2002) J Mol Biol 323:883–897
Banci L, Bertini I, Ciofi-Baffoni S, Su XC, Miras R, Bal N, Mintz E, Catty P, Shokes JE, Scott RA (2006) J Mol Biol 356:638–650
Liu T, Reyes-Caballero H, Li C, Scott RA, Giedroc DC (2007) Biochemistry 46:11057–11068
Okkeri J, Haltia T (2006) Biochim Biophys Acta 1757:1485–1495
Dutta SJ, Liu J, Hou Z, Mitra B (2006) Biochemistry 45:5923–5931
Liu J, Dutta SJ, Stemmler AJ, Mitra B (2006) Biochemistry 45:763–772
González-Guerrero M, Eren E, Rawat S, Stemmler TL, Argüello JM (2008) J Biol Chem 283:29753–29759
González-Guerrero M, Argüello JM (2008) Proc Natl Acad Sci USA 105:5992–5997
Borrelly GP, Rondet SA, Tottey S, Robinson NJ (2004) Mol Microbiol 53:217–227
Thelwell C, Robinson NJ, Turner-Cavet JS (1998) Proc Natl Acad Sci USA 85:10728–10733
Tottey S, Rich PR, Rondet SAM, Robinson NJ (2001) J Biol Chem 276:9999–20004
Tottey S, Rondet SAM, Borrelly GPM, Robinson PJ, Rich PR, Robinson NJ (2002) J Biol Chem 277:5490–5497
Pufahl RA, Singer CP, Peariso KL, Lin SJ, Schmidt PJ, Fahrni CJ, Culotta VC, Penner-Hahn JE, O’Halloran TV (1997) Science 278:853–856
Rosenzweig AC, O’Halloran TV (2000) Curr Opin Chem Biol 4:140–147
Wernimont AK, Huffman DL, Lamb AL, O’Halloran TV, Rosenzweig AC (2000) Nat Struct Biol 7:766–771
Walker JM, Tsivkovskii R, Lutsenko S (2002) J Biol Chem 277:27953–27959
Borrelly GPM, Blindauer CA, Schmid R, Butler CS, Cooper CE, Harvey I, Sadler PJ, Robinson NJ (2004) Biochem J 378:293–297
Banci L, Bertini I, Ciofi-Baffoni S, Su XC, Borrelly GP, Robinson NJ (2004) J Biol Chem 279:27502–27510
Banci L, Bertini I, Ciofi-Baffoni S, Kandias NG, Robinson NJ, Spyroulias GA, Su XC, Tottey S, Vanarotti M (2006) Proc Natl Acad Sci USA 103:8320–8325
Changela A, Chen K, Xue Y, Holschen J, Outten CE, O’Halloran TV, Mondragon A (2003) Science 301:1383–1387
Rae TD, Schmidt PJ, Pufahl RA, Culotta VC, O’Halloran TV (1999) Science 284:805–808
Arnesano F, Banci L, Bertini I, Cantini F, Ciofi-Baffoni S, Huffman DL, O’Halloran TV (2001) J Biol Chem 276:41365–41376
Beers J, Glerum DM, Tzagoloff A (1997) J Biol Chem 272:33191–33196
Culotta VC, Klomp LW, Strain J, Casareno RL, Krems B, Gitlin JD (1997) J Biol Chem 272:23469–23472
Klomp LW, Lin SJ, Yuan DS, Klausner RD, Culotta VC, Gitlin JD (1997) J Biol Chem 272:9221–9226
Valentine JS, Gralla EB (1997) Science 278:817–818
Carr S, Winge DR (2003) Acc Chem Res 36:309–316
Rosenzweig AC (2001) Acc Chem Res 34:119–128
O’Halloran TV, Finney LA (2003) Science 300:931–936
Banci L, Bertini I, Del Conte R, Markey J, Ruiz-Duenas FJ (2001) Biochemistry 25:15660–15668
Solioz M, Stoyanov JV (2003) FEMS Microbiol Rev 27:183–195
Keller R (2004) The computer aided resonance assignment tutorial. Cantina, Switzerland
Güntert P, Braun W, Wüthrich K (1991) J Mol Biol 217:517–530
Vuister GW, Bax A (1993) J Am Chem Soc 115:7772–7777
Gagné SM, Tsuda S, Li MX, Chandra M, Smillie LB, Sykes BD (1994) Protein Sci 3:1961–1974
Guntert P (2004) Methods Mol Biol 278:353–378
Case DA, Darden TA, Cheatham TE, Simmerling CE, Wang J, Duke RE, Luo R, Merz KM, Wang B, Pearlman DA, Crowley M, Brozell S, Tsui V, Gohlke H, Mongan J, Hornak V, Cui P, Beroza GP, Schafmeister CE, Caldwell JW, Ross WS, Kollman PA (2004) AMBER 8, version 8.0. University of California, San Francisco
Vriend G (1990) J Mol Graph 8:52–56
Laskowski RA, Rullmann JAC, MacArthur MW, Kaptein R, Thornton JM (1996) J Biomol NMR 8:477–486
Farrow NA, Muhandiram R, Singer AU, Pascal SM, Kay CM, Gish G, Shoelson SE, Pawson T, Forman-Kay JD, Kay LE (1994) Biochemistry 33:5984–6003
Grzesiek S, Bax A (1993) J Am Chem Soc 115:12593–12594
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) J Biomol NMR 6:277–293
Peng JW, Wagner G (1992) J Magn Reson 98:308–332
Lee LK, Rance M, Chazin WJ, Palmer AG (1997) J Biomol NMR 9:287–298
Hwang TL, Mori S, Shaka AJ, Van Zijl PCM (1997) J Am Chem Soc 119:6203–6204
Tjandra N, Kuboniwa H, Ren H, Bax A (1995) Eur J Biochem 230:1014–1024
Kroenke CD, Loria JP, Lee LK, Rance M, Palmer AG (1998) J Am Chem Soc 120:7905–7915
Banci L, Bertini I, Ciofi-Baffoni S, Huffman DL, O’Halloran TV (2001) J Biol Chem 276:8415–8426
Arnesano F, Banci L, Bertini I, Huffman DL, O’Halloran TV (2001) Biochemistry 40:1528–1539
Rosenzweig AC, Huffman DL, Hou MY, Wernimont AK, Pufahl RA, O’Halloran TV (1999) Structure 7:605–617
Achila D, Banci L, Bertini I, Bunce J, Ciofi-Baffoni S, Huffman DL (2006) Proc Natl Acad Sci USA 103:5729–5734
Banci L, Bertini I, Ciofi-Baffoni S, Del Conte R, Gonnelli L (2003) Biochemistry 42:1939–1949
VanZile ML, Cosper NJ, Scott RA, Giedroc DP (2000) Biochemistry 39:11818–11829
Dainty SJ, Patterson CJ, Waldron KJ, Robinson NJ (2009) J Biol Inorg Chem (in review)
Acknowledgments
This work was supported by research grants from the BBSRC (BB/E001688/1) and from the European Commission (“European Network of Research Infrastructures for Providing Access and Technological Advancements in Bio-NMR” contract no. 026145, SPINE II contract no. LSHG-CT-2006-031220 “From Receptor to Gene: Structures of Complexes from Signaling Pathways Linking Immunology, Neurobiology and Cancer,” and Marie Curie host fellowships for early stage research training no. MEST-CT-2004-504391 “NMR in Inorganic Structural Biology”).
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article will be printed in the upcoming Journal of Biological Inorganic Chemistry special issue CELL BIOLOGY OF COPPER.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Banci, L., Bertini, I., Ciofi-Baffoni, S. et al. NMR structural analysis of the soluble domain of ZiaA-ATPase and the basis of selective interactions with copper metallochaperone Atx1. J Biol Inorg Chem 15, 87–98 (2010). https://doi.org/10.1007/s00775-009-0568-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-009-0568-7