Abstract
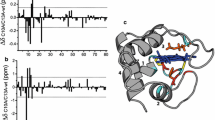
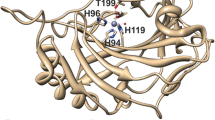
The structure of oxidized Rhodopseudomonas palustris cytochrome c 556 has been modeled after that of high-spin cytochrome c′ from the same bacterium, the latter being the protein with the greatest sequence identity (35%) among all sequenced proteins in the genomes. The two proteins differ in the number of ligands to iron and in spin state, the former being six-coordinate low-spin and the latter five-coordinate high-spin. In order to validate this modeled structure, several structural restraints were obtained by performing a restricted set of NMR experiments, without performing a complete assignment of the protein signals. The aim was to exploit the special restraints arising from the paramagnetism of the metal ion. A total of 43 residual-dipolar-coupling and 74 pseudocontact-shift restraints, which together sampled all regions of the protein, were used in conjunction with over 40 routinely obtained NOE distance restraints. A calculation procedure was undertaken combining the program MODELLER and the solution structure determination program PARAMAGNETIC DYANA, which includes paramagnetism-based restraints. The directions and magnitude of the magnetic susceptibility anisotropy tensor were also calculated. The approach readily provides useful results, especially for paramagnetic metalloproteins of moderate to large dimensions.






Similar content being viewed by others
References
Moore GR, McClune GJ, Clayden NJ, Williams RJ, Alsaadi BM, Angstrom J, Ambler RP, Van Beeumen J, Tempst P, Bartsch RG, Meyer TE, Kamen MD (1982) Eur J Biochem 123:73–80
La Mar GN, Jackson JT, Dugad LB, Cusanovich MA, Bartsch RG (1990) J Biol Chem 265:16173–16180
Banci L, Bertini I, Turano P, Vicens Oliver M (1992) Eur J Biochem 204:107–112
Tempst P, Van Beeumen J (1983) Eur J Biochem 129:603–614
Tempst P, Van Beeumen J (1983) Eur J Biochem 135:321–330
Ambler RP, Bartsch RG, Daniel M, Kamen MD, McLellan L, Meyer TE, Van Beeumen J (1981) Proc Natl Acad Sci USA 78:6854–6857
Ambler RP, Meyer TE, Bartsch RG, Cusanovich MA (2001) Arch Biochem Biophys 388:25–33
Hu W, De Smet L, Van Driessche G, Bartsch RG, Meyer TE, Cusanovich MA, Van Beeumen J (1998) Eur J Biochem 258:29–36
Shibata N, Iba S, Misaki S, Meyer TE, Bartsch RG, Cusanovich MA, Morimoto Y, Higuchi Y, Yasuoka N (1998) J Mol Biol 284:751
Sanchez R, Pieper U, Melo F, Marti-Renom MA, Mirkovic N, Sali A (2000) Nat Struct Biol 7:986–990
Fesik SW, Zuiderweg ERP (1988) J Magn Reson 78:588–593
Bertini I, Luchinat C, Parigi G (2002) Prog NMR Spectrosc 40:249–273
Kurland RJ, McGarvey BR (1970) J Magn Reson 2:286–301
Bertini I, Luchinat C, Parigi G (2001) Solution NMR of paramagnetic molecules. Elsevier, Amsterdam
Banci L, Bertini I, Huber JG, Luchinat C, Rosato A (1998) J Am Chem Soc 120:12903–12909
Sali A, Potterton L, Yuan F, Van Vlijmen H, Karplus M (1995) Proteins Struct Funct Genet 23:318–326
Huang X, Miller W (1991) Adv Appl Math 12:337–357
Güntert P, Mumenthaler C, Wüthrich K (1997) J Mol Biol 273:283–298
Banci L, Bertini I, Cremonini MA, Gori Savellini G, Luchinat C, Wüthrich K, Güntert P (1998) J Biomol NMR 12:553–557
Bertini I, Luchinat C, Parigi G (2002) Concepts Magn Reson 14:259–286
Emerson SD, La Mar GN (1990) Biochemistry 29:1545–1556
Bren KL, Gray HB, Banci L, Bertini I, Turano P (1995) J Am Chem Soc 117:8067–8073
Banci L, Bertini I, Pierattelli R, Vila AJ (1994) Inorg Chem 33:4338–4343
Johnson RD, Ramaprasad S, La Mar GN (1983) J Am Chem Soc 105:7205–7206
Banci L, Bertini I, Luchinat C, Piccioli M, Scozzafava A, Turano P (1989) Inorg Chem 28:4650–4656
Banci L, Bertini I, Luchinat C, Piccioli M (1991) In: Bertini I, Molinari H, Niccolai N (eds) NMR and biomolecular structure. VCH, Weinheim, pp 31–60
Arnesano F, Banci L, Bertini I, Faraone-Mennella J, Rosato A, Barker PD, Fersht AR (1999) Biochemistry 38:8657–8670
Arnesano F, Banci L, Bertini I, Ciofi-Baffoni S, de Lumley Woodyear T, Johnson CM, Barker PD (2000) Biochemistry 39:1499–1514
Jeener J, Meier BH, Bachmann P, Ernst RR (1979) J Chem Phys 71:4546–4553
Bertini I, Donaire A, Felli IC, Luchinat C, Rosato A (1996) Magn Reson Chem 34:948–950
Allegrozzi M, Bertini I, Janik MBL, Lee Y-M, Liu G, Luchinat C (2000) J Am Chem Soc 122:4154–4161
Banci L, Bertini I, Bren KL, Cremonini MA, Gray HB, Luchinat C, Turano P (1996) J Biol Inorg Chem 1:117–126
Shokhirev NV, Walker FA (1998) J Biol Inorg Chem 3:581–594
Banci L, Bertini I, Gray HB, Luchinat C, Reddig T, Rosato A, Turano P (1997) Biochemistry 36:9867–9877
Banci L, Bertini I, Bren KL, Gray HB, Sompornpisut P, Turano P (1997) Biochemistry 36:8992–9001
Bertini I, Luchinat C, Parigi G, Walker FA (1999) J Biol Inorg Chem 4:515–519
Arnesano F, Banci L, Bertini I, Karin van der Wetering, Czisch M, Kaptein R (2000) J Biomol NMR 17:295–304
Chou JJ, Li S, Klee CB, Bax A (2001) Nat Struct Biol 8:990–997
Bertini I, Kowalewski J, Luchinat C, Parigi G (2001) J Magn Reson 152:103–108
Acknowledgements
This work was supported by Murst ex 40%, Italy, the European Union, contracts HPRI-CT-2001-00147 and QLG2-CT-1999-01003, CNR, Italy, contract 99.00950.CT03, and the United States Department of Energy (grant no. DE-FG03-02ER15359 to Caltech).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Bertini, I., Faraone-Mennella, J., Gray, H.B. et al. NMR-validated structural model for oxidized Rhodopseudomonas palustris cytochrome c 556 . J Biol Inorg Chem 9, 224–230 (2004). https://doi.org/10.1007/s00775-003-0511-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-003-0511-2