Abstract
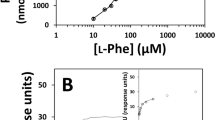
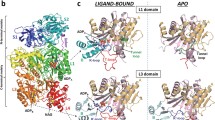
Phenylalanine hydroxylase (PAH) catalyzes the hydroxylation of L-Phe to L-Tyr. Dysfunctional PAH results in phenylketonuria and mammalian PAH is therefore highly regulated and displays positive cooperativity for L-Phe (Hill coefficient (h) = 2). L-Phe does not bind to the regulatory ACT domain in full-length tetrameric human PAH and cooperativity is elicited by homotropic binding to the catalytic site (Thórólfsson et al. in Biochemistry 41:7573–7585, 2002). PAH from Caenorhabditis elegans (cePAH) is devoid of cooperativity for L-Phe (h = 0.9), and, as shown in this work, structural analysis reveal an additional L-Phe binding site at the regulatory domain of full-length cePAH. This site involves the GA(S)L/ISRP motifs, which are also found in ACT domains of other L-Phe binding proteins, such as prephenate dehydratase. Isothermal titration calorimetry further demonstrated 2 binding sites per subunit for cePAH versus ~1 for hPAH. Steric occlusion of the regulatory site, notably by residues Lys215/Tyr216 from the adjacent catalytic domain, appears to hinder regulatory binding in full-length hPAH. Accordingly, the humanized mutant Q215K/N216Y of cePAH binds ~1.4 L-Phe/subunit. This mutant also displays high catalytic activity and certain positive cooperativity for L-Phe (h = 1.4). Our results support that the acquisition of positive cooperativity in mammalian forms of PAH is accompanied by a closure of the regulatory L-Phe binding site. Concomitantly, the function of the regulatory ACT domain appears to be adapted from amino acid binding to serving the communication of conformational changes among catalytic subunits.








Similar content being viewed by others
Abbreviations
- AAAH:
-
Aromatic amino acid hydroxylases
- BH4 :
-
(6R)-L-erythro-5,6,7,8-tetrahydrobiopterin
- cePAH:
-
Phenylalanine hydroxylase from C. elegans
- hPAH:
-
Human phenylalanine hydroxylase
- ITC:
-
Isothermal titration calorimetry
- MD:
-
Molecular dynamics
- PAH:
-
Phenylalanine hydroxylase
- PDT:
-
Prephenate dehydratase
- TH:
-
Tyrosine hydroxylase
- T m :
-
Midpoint melting temperature
- TPH:
-
Tryptophan hydroxylase
- wt:
-
Wild type
References
An J, Totrov M, Abagyan R (2005) Pocketome via comprehensive identification and classification of ligand binding envelopes. Mol Cell Proteomics 4:752–761
Bel Y, Jacobson KB, Ferre J (1992) A comparative study of Drosophila phenylalanine hydroxylase with a natural and a synthetic tetrahydropterin as cofactor. Comp Biochem Physiol B 103:557–562
Calvo AC, Pey AL, Ying M, Loer CM, Martinez A (2008) Anabolic function of phenylalanine hydroxylase in Caenorhabditis elegans. FASEB J 22:3046–3058
Case DA, Darden TA, Cheatham TE I, Simmerling CL, Wang J, Duke RE et al (2008) AMBER 10. University of California, San Francisco
Clamp M, Cuff J, Searle SM, Barton GJ (2004) The Jalview Java alignment editor. Bioinformatics 20:426–427
Døskeland AP, Døskeland SO, Øgreid D, Flatmark T (1984) The effect of ligands of phenylalanine 4-monooxygenase on the cAMP-dependent phosphorylation of the enzyme. J Biol Chem 259:11242–11248
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797
Erlandsen H, Pey AL, Gamez A, Perez B, Desviat LR, Aguado C et al (2004) Correction of kinetic and stability defects by tetrahydrobiopterin in phenylketonuria patients with certain phenylalanine hydroxylase mutations. Proc Natl Acad Sci USA 101:16903–16908
Fisher AL, Page KE, Lithgow GJ, Nash L (2008) The Caenorhabditis elegans K10C2.4 gene encodes a member of the fumarylacetoacetate hydrolase family: a Caenorhabditis elegans model of type I tyrosinemia. J Biol Chem 283:9127–9135
Fitzpatrick PF (1999) Tetrahydropterin-dependent amino acid hydroxylases. Annu Rev Biochem 68:355–381
Fitzpatrick PF (2003) Mechanism of aromatic amino acid hydroxylation. Biochemistry 42:14083–14091
Flatmark T, Stevens RC (1999) Structural insight into the aromatic amino acid hydroxylases and their disease-related mutant forms. Chem Rev 99:2137–2160
Gersting SW, Kemter KF, Staudigl M, Messing DD, Danecka MK, Lagler FB et al (2008) Loss of function in phenylketonuria is caused by impaired molecular motions and conformational instability. Am J Hum Genet 83:5–17
Gjetting T, Petersen M, Guldberg P, Guttler F (2001) Missense mutations in the N-terminal domain of human phenylalanine hydroxylase interfere with binding of regulatory phenylalanine. Am J Hum Genet 68:1353–1360
Gu X (2001) Maximum-likelihood approach for gene family evolution under functional divergence. Mol Biol Evol 18:453–464
Gu X (2006) A simple statistical method for estimating type-II (cluster-specific) functional divergence of protein sequences. Mol Biol Evol 23:1937–1945
Hooft RW, Vriend G, Sander C, Abola EE (1996) Errors in protein structures. Nature 381:272
Hufton SE, Jennings IG, Cotton RG (1995) Structure and function of the aromatic amino acid hydroxylases. Biochem J 311:353–366
Infanger LC, Rocheleau TA, Bartholomay LC, Johnson JK, Fuchs J, Higgs S et al (2004) The role of phenylalanine hydroxylase in melanotic encapsulation of filarial worms in two species of mosquitoes. Insect Biochem Mol Biol 34:1329–1338
Kappock TJ, Caradonna JP (1996) Pterin-dependent amino acid hydroxylases. Chem Rev 96:2659–2756
Kappock TJ, Harkins PC, Friedenberg S, Caradonna JP (1995) Spectroscopic and kinetic properties of unphosphorylated rat hepatic phenylalanine hydroxylase expressed in Escherichia coli—comparison of resting and activated states. J Biol Chem 270:30532–30544
Kaufman S (1993) The phenylalanine hydroxylating system. Adv Enzymol Relat Areas Mol Biol 67:77–264
Kleppe R, Uhlemann K, Knappskog PM, Haavik J (1999) Urea-induced denaturation of human phenylalanine hydroxylase. J Biol Chem 274:33251–33258
Kobe B, Jennings IG, House CM, Michell BJ, Goodwill KE, Santarsiero BD et al (1999) Structural basis of autoregulation of phenylalanine hydroxylase. Nat Struct Biol 6:442–448
Leiros HK, Pey AL, Innselset M, Moe E, Leiros I, Steen IH et al (2007) Structure of phenylalanine hydroxylase from Colwellia psychrerythraea 34H, a monomeric cold active enzyme with local flexibility around the active site and high overall stability. J Biol Chem 282:21973–21986
Loer CM, Davidson B, McKerrow J (1999) A phenylalanine hydroxylase gene from the nematode C. elegans is expressed in the hypodermis. J Neurogenet 13:157–180
Martínez A, Haavik J, Flatmark T (1990) Cooperative homotropic interaction of L-noradrenaline with the catalytic site of phenylalanine 4-monooxygenase. Eur J Biochem 193:211–219
Martínez A, Olafsdottir S, Flatmark T (1993) The cooperative binding of phenylalanine to phenylalanine 4-monooxygenase studied by 1H-NMR paramagnetic relaxation. Changes in water accessibility to the iron at the active site upon substrate binding. Eur J Biochem 211:259–266
Pey AL, Thórólfsson M, Teigen K, Ugarte M, Martínez A (2004) Thermodynamic characterization of the binding of tetrahydropterins to phenylalanine hydroxylase. J Am Chem Soc 126:13670–13678
Phillips RS, Kaufman S (1984) Ligand effects on the phosphorylation state of hepatic phenylalanine hydroxylase. J Biol Chem 259:2474–2479
Schwede T, Kopp J, Guex N, Peitsch MC (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucl Acids Res 31:3381–3385
Scriver CR, Kaufman S (2001) Hyperphenylalaninemia:phenylalanine hydroxylase deficiency. In: Scriver CR, Beaudet AL, Valle D, Sly WS (eds) The metabolic and molecular bases of inherited disease, 8th edn. McGraw-Hill, New York, pp 1667–1724
Shiman R, Gray DW (1980) Substrate activation of phenylalanine hydroxylase. A kinetic characterization. J Biol Chem 255:4793–4800
Shiman R, Gray DW, Pater A (1979) A simple purification of phenylalanine hydroxylase by substrate-induced hydrophobic chromatography. J Biol Chem 254:11300–11306
Shiman R, Xia T, Hill MA, Gray DW (1994) Regulation of rat liver phenylalanine hydroxylase. II. Substrate binding and the role of activation in the control of enzymatic activity. J Biol Chem 269:24647–24656
Siltberg-Liberles J, Martinez A (2009) Searching distant homologs of the regulatory ACT domain in phenylalanine hydroxylase. Amino Acids 36:235–249
Siltberg-Liberles J, Steen IH, Svebak RM, Martinez A (2008) The phylogeny of the aromatic amino acid hydroxylases revisited by characterizing phenylalanine hydroxylase from Dictyostelium discoideum. Gene 427:86–92
Solstad T, Stokka AJ, Andersen OA, Flatmark T (2003) Studies on the regulatory properties of the pterin cofactor and dopamine bound at the active site of human phenylalanine hydroxylase. Eur J Biochem 270:981–990
Stokka AJ, Flatmark T (2003) Substrate-induced conformational transition in human phenylalanine hydroxylase as studied by surface plasmon resonance analyses: the effect of terminal deletions, substrate analogues and phosphorylation. Biochem J 369:509–518
Stokka AJ, Carvalho RN, Barroso JF, Flatmark T (2004) Probing the role of crystallographically defined/predicted hinge-bending regions in the substrate-induced global conformational transition and catalytic activation of human phenylalanine hydroxylase by single-site mutagenesis. J Biol Chem 279:26571–26580
Tan K, Li H, Zhang R, Gu M, Clancy ST, Joachimiak A (2008) Structures of open (R) and close (T) states of prephenate dehydratase (PDT)–implication of allosteric regulation by L-phenylalanine. J Struct Biol 162:94–107
Teigen K, McKinney JA, Haavik J, Martinez A (2007) Selectivity and affinity determinants for ligand binding to the aromatic amino Acid hydroxylases. Curr Med Chem 14:455–467
Thöny B, Auerbach G, Blau N (2000) Tetrahydrobiopterin biosynthesis, regeneration and functions. Biochem J 347(Pt 1):1–16
Thórólfsson M, Ibarra-Molero B, Fojan P, Petersen SB, Sanchez-Ruiz JM, Martínez A (2002) L-phenylalanine binding and domain organization in human phenylalanine hydroxylase: a differential scanning calorimetry study. Biochemistry 41:7573–7585
Thórólfsson M, Teigen K, Martínez A (2003) Activation of phenylalanine hydroxylase: effect of substitutions at Arg68 and Cys237. Biochemistry 42:3419–3428
Vriend G (1990) WHAT IF: a molecular modeling and drug design program. J Mol Graph 8:52–56
Wiens M, Koziol C, Batel R, Muller WE (1998) Phenylalanine hydroxylase from the sponge Geodia cydonium: implication for allorecognition and evolution of aromatic amino acid hydroxylases. Dev Comp Immunol 22:469–478
Acknowledgments
The expert assistance of Randi M. Svebak and Ali J. Muñoz Sepulveda is greatly appreciated. We are grateful to Anne Wiemhoefer and Thies Nolte for preliminary DSC experiments. This work received support from the Research Council of Norway and Meltzer L. Høyskolefond.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Flydal, M.I., Mohn, T.C., Pey, A.L. et al. Superstoichiometric binding of L-Phe to phenylalanine hydroxylase from Caenorhabditis elegans: evolutionary implications. Amino Acids 39, 1463–1475 (2010). https://doi.org/10.1007/s00726-010-0611-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-010-0611-6