Abstract
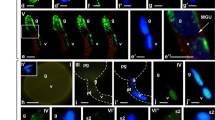
The transcriptional and posttranscriptional AGO-mediated control of gene expression may play important roles during male monocot gametophyte development. In this report, we demonstrated dynamic changes in the spatiotemporal distribution of AGO1 and AGO4, which are key proteins of the RNA-induced silencing complex (RISC) in Hyacinthus orientalis male gametophyte development. During maturation of the bicellular pollen grains and in vitro pollen tube growth, the pattern of AGO1 localization was correlated with previously observed transcriptional activity of the cells. During the period of high transcriptional activity, AGO1 is associated with chromatin while the clustered distribution of AGO1 in the interchromatin areas is accompanied by condensation of chromatin and the gradual transcriptional silencing of both cells in mature, dehydrated pollen. During pollen tube growth and the restarting of RNA synthesis in the vegetative nucleus, AGO1 is dispersed in the chromatin. Additionally, the gradual increase in the cytoplasmic pool of AGO1 in the elongating pollen tube indicates the activation of the posttranscriptional gene silencing (PTGS) pathway. During pollen tube growth in the generative cell and in the sperm cells, AGO1 is present mainly in the areas between highly condensed chromatin clusters. Changes in the distribution of AGO4 that indicated the possibility of spatiotemporal organization in the RNA-directed DNA methylation (RdDM) process (cytoplasmic and nuclear steps) were also observed during hyacinth male gametophyte development. Based on our findings, we propose that in the germinating pollen tube, the cytoplasmic assembly of AGO4/siRNA takes place and that the mature complexes could be transported to the nucleus to carry out their function during the next steps of pollen tube growth.






Similar content being viewed by others
Abbreviations
- 5mC:
-
5-Methylcytosine
- acH4:
-
Acetylated histone H4
- AGO1:
-
Argonaute 1
- AGO4:
-
Argonaute 4
- BMA:
-
Butyl methacrylate
- GC:
-
Generative cytoplasm
- GN:
-
Generative nucleus
- MGU:
-
Male germ unit
- miRNA:
-
Micro RNA
- MMA:
-
Methyl methacrylate
- PAb:
-
Polyclonal antibody
- PTGS:
-
Posttranscriptional gene silencing
- RdDM:
-
RNA-directed DNA methylation
- RISC:
-
RNA-induced silencing complex
- SC:
-
Sperm cell
- siRNA:
-
Small interfering RNA
- sRNA:
-
Small noncoding RNA
- TE:
-
Transposable element
- TGS:
-
Transcriptional gene silencing
- VC:
-
Vegetative cytoplasm
- VN:
-
Vegetative nucleus
References
Arribas-Hernández L, Kielpinski LJ, Brodersen P (2016) mRNA decay of Most Arabidopsis miRNA targets requires slicer activity of AGO1. Plant Physiol 171:2620–2632. https://doi.org/10.1104/pp.16.00231
Baumberger N, Baulcombe DC (2005) Arabidopsis ARGONAUTE1 is an RNA slicer that selectively recruits microRNAs and short interfering RNAs. Proc Natl Acad Sci U S A 102:11928–11933. https://doi.org/10.1073/pnas.0505461102
Bednarska E, Górska-Brylass A (1987) Ultrastructural transformations of nuclei differentiating Hyacinthus orientalis L. pollen grain cells. Acta Soc Bot Pol 56:667–685. https://doi.org/10.5586/asbp.1987.057
Bednarska E (1988) Ultrastructural transformations in the cytoplasm of differentiating Hyacinthus orientalis L. pollen cells. Acta Soc Bot Pol 57:235–245. https://doi.org/10.5586/asbp.1988.024
Borges F, Gomes G, Gardner R, Moreno N, McCormick S, Feijó JA, Becker JD (2008) Comparative transcriptomics of Arabidopsis sperm cells. Plant Physiol 148:1168–1181. https://doi.org/10.1104/pp.108.125229
Borges F, Pereira PA, Slotkin RK, Martienssen RA, Becker JD (2011) MicroRNA activity in the Arabidopsis male germline. J Exp Bot 62:1611–1620. https://doi.org/10.1093/jxb/erq452
Borges F, Martienssen RA (2013) Establishing epigenetic variation during genome reprogramming. RNA Biol 10:490–494. https://doi.org/10.4161/rna.24085
Brewbaker JL, Kwack BH (1963) The essential role of calcium ions in pollen germination and pollen tube growth. Am J Bot 50:859–865 https://www.jstor.org/stable/2439772
Calarco JP, Borges F, Donoghue MT, Van Ex F, Jullien PE, Lopes T, Gardner R, Berger F, Feijó JA, Becker JD, Martienssen RA (2012) Reprogramming of DNA methylation in pollen guides epigenetic inheritance via small RNA. Cell 151:194–205. https://doi.org/10.1016/j.cell.2012.09.001
Carbonell A, Fahlgren N, Garcia-Ruiz H, Gilbert KB, Montgomery TA, Nguyen T, Cuperus JT, Carrington JC (2012) Functional analysis of three Arabidopsis ARGONAUTES using slicer-defective mutants. Plant Cell 24:3613–3629. https://doi.org/10.1105/tpc.112.099945
Chambers C, Shuai B (2009) Profiling microRNA expression in Arabidopsis pollen using microRNA array and real-time PCR. BMC Plant Biol 10:87. https://doi.org/10.1186/1471-2229-9-87
Chan SW, Zilberman D, Xie Z, Johansen LK, Carrington JC, Jacobsen SE (2004) RNA silencing genes control de novo DNA methylation. Science 303:1336 http://science.sciencemag.org/content/303/5662/1336
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 25:21–44. https://doi.org/10.1146/annurev.cellbio.042308.113417
Dolata J, Bajczyk M, Bielewicz D, Niedojadło K, Niedojadło J, Pietrykowska H, Walczak W, Szweykowska-Kulinska Z, Jarmolowski A (2016) Salt stress reveals a new role for ARGONAUTE1 in miRNA biogenesis at the transcriptional and posttranscriptional levels. Plant Physiol 172:297–312. https://doi.org/10.1104/pp.16.00830
Duan CG, Fang YY, Zhou BJ, Zhao JH, Hou WN, Zhu H, Ding SW, Guo HS (2012) Suppression of Arabidopsis ARGONAUTE1-mediated slicing, transgene-induced RNA silencing, and DNA methylation by distinct domains of the Cucumber mosaic virus 2b protein. Plant Cell 24:259–274 http://www.plantcell.org/cgi/doi/10.1105/tpc.111.092718
Dunoyer P, Voinnet O (2005) The complex interplay between plant viruses and host RNA-silencing pathways. Curr Opin Plant Biol 8:415–423. https://doi.org/10.1016/j.pbi.2005.05.012
Grant-Downton R, Hafidh S, Twell D, Dickinson HG (2009a) Small RNA pathways are present and functional in the angiosperm male gametophyte. Mol Plant 2:500–512. https://doi.org/10.1093/mp/ssp003
Grant-Downton R, Le Trionnaire G, Schmid R, Rodriguez-Enriquez J, Hafidh S, Mehdi S, Twell D, Dickinson H (2009b) MicroRNA and tasiRNA diversity in mature pollen of Arabidopsis thaliana. BMC Genomics 10:643. https://doi.org/10.1186/1471-2164-10-643
Grant-Downton R, Rodriguez-Enriquez J (2012) Emerging roles for non-coding RNAs in male reproductive development in flowering plants. Biomolecules 2:608–621. https://doi.org/10.3390/biom2040608
Grant-Downton R, Kourmpetli S, Hafidh S, Khatab H, Le Trionnaire G, Dickinson H, Twell D (2013) Artificial microRNAs reveal cell-specific differences in small RNA activity in pollen. Curr Biol 23:R599–R601. https://doi.org/10.1016/j.cub.2013.05.055
Hafidh S, Fíla J, Honys D (2016) Male gametophyte development and function in angiosperms: a general concept. Plant Reprod 29:31–51. https://doi.org/10.1007/s00497-015-0272-4
Hutvagner G, Simard MJ (2008) Argonaute proteins: key players in RNA silencing. Nat Rev Mol Cell Biol 9:22–32. https://doi.org/10.1038/nrm2321
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAs and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53. https://doi.org/10.1146/annurev.arplant.57.032905.105218
Kapoor M, Arora R, Lama T, Nijhawan A, Khurana JP, Tyagi AK, Kapoor S (2008) Genome-wide identification, organization and phylogenetic analysis of dicer-like, Argonaute and RNA-dependent RNA polymerase gene families and their expression analysis during reproductive development and stress in rice. BMC Genomics 9:451. https://doi.org/10.1186/1471-2164-9-451
Kozłowska M, Niedojadło K, Brzostek M, Bednarska-Kozakiewicz E (2016) Epigenetic marks in the Hyacinthus orientalis L. mature pollen grain and during in vitro pollen tube growth. Plant Reprod 29:251–263 https://doi.org/10.1007/s00497-016-0289-3
Lenartowski R, Suwińska A, Lenartowska M (2015) Calreticulin expression in relation to exchangeable Ca(2+) level that changes dynamically during anthesis, progamic phase, and double fertilization in Petunia. Planta 241:209–227. https://doi.org/10.1007/s00425-014-2178-z
Le Trionnaire G, Grant-Downton RT, Kourmpetli S, Dickinson HG, Twell D (2011) Small RNA activity and function in angiosperm gametophytes. J Exp Bot 62:1601–1610. https://doi.org/10.1093/jxb/erq399
Liu X, Lu T, Dou Y, Yu B, Zhang C (2014) Identification of RNA silencing components in soybean and sorghum. BMC bioinformatics 15:4. https://doi.org/10.1186/1471-2105-15-4
Liu C, Xin Y, Xu L, Cai Z, Xue Y, Liu Y, Xie D, Liu Y, Qi Y (2017) Arabidopsis ARGONAUTE1 binds chromatin to promote gene transcription in response to hormones and stresses. Dev Cell 44:348–361. https://doi.org/10.1016/j.devcel.2017.12.002
Ma Z, Zhang X (2018) Actions of plant Argonautes: predictable or unpredictable? Curr Opin Plant Biol 45:59–67. https://doi.org/10.1016/j.pbi.2018.05.007
Mallory AC, Vaucheret H (2006) Functions of microRNAs and related small RNAs in plants. Nat Genet 38(Suppl):S31–S36. https://doi.org/10.1038/ng1791
Mallory A, Vaucheret H (2010) Form, function, and regulation of ARGONAUTE proteins. Plant Cell 22:3879–3889. https://doi.org/10.1105/tpc.110.080671
Matzke MA, Kanno T, Matzke AJ (2015) RNA-directed DNA methylation: the evolution of a complex epigenetic pathway in flowering plants. Annu Rev Plant Biol 66:243–267. https://doi.org/10.1146/annurev-arplant-043014-114633
Movahedi A, Sun W, Zhang J, Wu X, Mousavi M, Mohammadi K, Yin T, Zhuge Q (2015) RNA-directed DNA methylation in plants. Plant Cell Rep 34:1857–1862. https://doi.org/10.1007/s00299-015-1839-0
Niedojadło J, Dełeńko K, Niedojadło K (2016) Regulation of poly(a) RNA retention in the nucleus as a survival strategy of plants during hypoxia. RNA Biol 3:531–543. https://doi.org/10.1080/15476286.2016.1166331
Peng H, Chun J, Ai TB, Tong YA, Zhang R, Zhao MM, Chen F, Wang SH (2012) MicroRNA profiles and their control of male gametophyte development in rice. Plant Mol Biol 80:85–102. https://doi.org/10.1007/s11103-012-9898-x
Poulsen C, Vaucheret H, Brodersen P (2013) Lessons on RNA silencing mechanisms in plants from eukaryotic argonaute structures. Plant Cell 25:22–37. https://doi.org/10.1105/tpc.112.105643
Qian Y, Cheng Y, Cheng X, Jiang H, Zhu S, Cheng B (2011) Identification and characterization of dicer-like, Argonaute and RNA-dependent RNA polymerase gene families in maize. Plant Cell Rep 30:1347–1363. https://doi.org/10.1007/s00299-011-1046-6
Russell SD, Gou X, Wong CE, Wang X, Yuan T, Wei X, Bhalla PL, Singh MB (2012) Genomic profiling of rice sperm cell transcripts reveals conserved and distinct elements in the flowering plant male germ lineage. New Phytol 195:560–573. https://doi.org/10.1111/j.1469-8137.2012.04199.x
Slotkin RK, Vaughn M, Borges F, Tanurdzić M, Becker JD, Feijó JA, Martienssen RA (2009) Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136:461–472. https://doi.org/10.1016/j.cell.2008.12.038
Song X, Li P, Zhai J, Zhou M, Ma L, Liu B, Jeong DH, Nakano M, Cao S, Liu C, Chu C, Wang XJ, Green PJ, Meyers BC, Cao X (2012) Roles of DCL4 and DCL3b in rice phased small RNA biogenesis. Plant J 69:462–474. https://doi.org/10.1111/j.1365-313X.2011.04805.x
Tolia NH, Joshua-Tor L (2007) Slicer and the argonautes. Nat Chem Biol 3:36–43. https://doi.org/10.1038/nchembio848
Van Ex F, Jacob Y, Martienssen RA (2011) Multiple roles for small RNAs during plant reproduction. Curr Opin Plant Biol 14:588–593. https://doi.org/10.1016/j.pbi.2011.07.003
Vaucheret H (2008) Plant ARGONAUTES. Trends Plant Sci 13:350–358. https://doi.org/10.1016/j.tplants.2008.04.007
Wang H, Zhang X, Liu J, Kiba T, Woo J, Ojo T, Hafner M, Tuschl T, Chua NH, Wang XJ (2011) Deep sequencing of small RNAs specifically associated with Arabidopsis AGO1 and AGO4 uncovers new AGO functions. Plant J 67:292–304. https://doi.org/10.1111/j.1365-313X.2011.04594.x
Wei LQ, Yan LF, Wang T (2011) Deep sequencing on genome-wide scale reveals the unique composition and expression patterns of microRNAs in developing pollen of Oryza sativa. Genome Biol 12:R53. https://doi.org/10.1186/gb-2011-12-6-r53
Wendte JM, Pikaard CS (2017) The RNAs of RNA-directed DNA methylation. Biochim Biophys Acta 1860:140–148. https://doi.org/10.1016/j.bbagrm.2016.08.004
Ye R, Wang W, Iki T, Liu C, Wu Y, Ishikawa M, Zhou X, Qi Y (2012) Cytoplasmic assembly and selective nuclear import of Arabidopsis Argonaute4/siRNA complexes. Mol Cell 46:859–870. https://doi.org/10.1016/j.molcel.2012.04.013
Zhai L, Sun W, Zhang K, Jia H, Liu L, Liu Z, Teng F, Zhang Z (2014) Identification and characterization of Argonaute gene family and meiosis-enriched Argonaute during sporogenesis in maize. J Integr Plant Biol 56:1042–1052. https://doi.org/10.1111/jipb.12205
Zhang H, Xia R, Meyers BC, Walbot V (2015) Evolution, functions, and mysteries of plant ARGONAUTE proteins. Curr Opin Plant Biol 27:84–90. https://doi.org/10.1016/j.pbi.2015.06.011
Zienkiewicz K, Smoliński DJ, Bednarska E (2006) Distribution of poly(a) RNA and splicing machinery elements in mature Hyacinthus orientalis L. pollen grains and pollen tubes growing in vitro. Protoplasma 227:95–103. https://doi.org/10.1007/s00709-005-0153-z
Zienkiewicz K, Zienkiewicz A, Smoliński DJ, Rafińska K, Świdziński M, Bednarska E (2008a) Transcriptional state and distribution of poly(a) RNA and RNA polymerase II in differentiating Hyacinthus orientalis L. pollen grains. Sex Plant Reprod 21:233–245. https://doi.org/10.1007/s00497-008-0085-9
Zienkiewicz K, Zienkiewicz A, Rodriguez-Garcia MI, Smoliński DJ, Świdziński M, Bednarska E (2008b) Transcriptional activity and distribution of splicing machinery elements during Hyacinthus orientalis L. pollen tube growth. Protoplasma 233:129–139. https://doi.org/10.1007/s00709-008-0298-7
Zienkiewicz K, Suwińska A, Niedojadło K, Zienkiewicz A, Bednarska E (2011) Nuclear activity of sperm cells during Hyacinthus orientalis L. in vitro pollen tube growth. J Exp Bot 62:1255–1269. https://doi.org/10.1093/jxb/erq354
Zilberman D, Cao X, Jacobsen SE (2003) ARGONAUTE4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299:716–719. https://doi.org/10.1126/science.1079695
Zilberman D, Cao X, Johansen LK, Xie Z, Carrington JC, Jacobsen SE (2004) Role of Arabidopsis ARGONAUTE4 in RNA-directed DNA methylation triggered by inverted repeats. Curr Biol 14:1214–1220. https://doi.org/10.1016/j.cub.2004.06.055
Funding
This project was supported by Polish National Science Center (NCN) grant no. 2011/03/D/NZ3/00603 (to KN).
Author information
Authors and Affiliations
Contributions
KN conceived and designed research. KN, MK, AK-L, RL conducted the experiments. KN and AK-L performed the quantitative analysis. KN, JN, and EB-K analyzed the experimental and statistical data. KN wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Burkhard Becker
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Fig. S1
(PNG 282 kb)
Supplementary Fig. S2
(PNG 485 kb)
Supplementary Fig. S3
(PNG 466 kb)
Supplementary Fig. S4
(PNG 1007 kb)
Supplementary Fig. S5
(PNG 837 kb)
Supplementary Fig. S6
(PNG 463 kb)
Supplementary Fig. S7
(PNG 557 kb)
Supplementary Fig. S8
(PNG 396 kb)
Supplementary Fig. S9
(PNG 389 kb)
Supplementary Fig. S10
(PNG 276 kb)
Supplementary Fig. S11
(PNG 955 kb)
Rights and permissions
About this article
Cite this article
Niedojadło, K., Kupiecka, M., Kołowerzo-Lubnau, A. et al. Dynamic distribution of ARGONAUTE1 (AGO1) and ARGONAUTE4 (AGO4) in Hyacinthus orientalis L. pollen grains and pollen tubes growing in vitro. Protoplasma 257, 793–805 (2020). https://doi.org/10.1007/s00709-019-01463-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-019-01463-2