Abstract
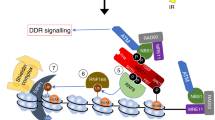
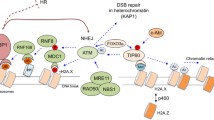
This review focuses on the function of heterochromatin protein HP1 in response to DNA damage. We specifically outline the regulatory mechanisms in which HP1 and its interacting partners are involved. HP1 protein subtypes (HP1α, HP1β, and HP1γ) are the main components of constitutive heterochromatin, and HP1α and HP1β in particular are responsible for heterochromatin maintenance. The recruitment of these proteins to DNA lesions is also important from the perspective of proper DNA repair mechanisms. For example, HP1α is necessary for the binding of the main DNA damage-related protein 53BP1 at DNA repair foci, which are positive not only for the HP1α protein but also for the RAD51 protein, a component of DNA repair machinery. The HP1β protein also appears in monomeric form in DNA lesions together with the evolutionarily well-conserved protein called proliferating cell nuclear antigen (PCNA). The role of HP1 in DNA lesions is also mediated via the Kap1 transcription repressor. Taken together, these results indicate that the function of HP1 after DNA injury depends strongly on the kinetics of other DNA repair-related factors and their post-translational modifications, such as the phosphorylation of Kap-1.



Similar content being viewed by others
Notes
FRAP shows, via fluorescence protein technology (for example, green fluorescent protein (GFP)-tagged protein of interest), how a fluorescence probe diffuses throughout the sample. The fluorescence of fluorochrome (for example, GFP or mCherry) is destroyed by laser photobleaching, and the fraction that is not recovered represents the so-called immobile fraction (Bastiaens and Pepperkok 2000). The proteins that especially accumulate at nuclear bodies or chromocenters (clusters of centromeric heterochromatin) are characterized by a striking immobile fraction (Cheutin et al. 2003).
References
Alagoz M, Katsuki Y, Ogiwara H et al (2015) SETDB1, HP1 and SUV39 promote repositioning of 53BP1 to extend resection during homologous recombination in G 2 cells. Nucleic Acids Res 43:7931–7944. doi:10.1093/nar/gkv722
Aucott R, Bullwinkel J, Yu Y et al (2008) HP1-beta is required for development of the cerebral neocortex and neuromuscular junctions. J Cell Biol 183:597–606. doi:10.1083/jcb.200804041
Ayoub N, Jeyasekharan AD, Bernal JA, Venkitaraman AR (2008) HP1-beta mobilization promotes chromatin changes that initiate the DNA damage response. Nature 453:682–686. doi:10.1038/nature06875
Ayoub N, Jeyasekharan AD, Venkitaraman AR (2009) Mobilization and recruitment of HP1: a bimodal response to DNA breakage. Cell Cycle 8:2945–2950
Baldeyron C, Soria G, Roche D et al (2011) HP1alpha recruitment to DNA damage by p150CAF-1 promotes homologous recombination repair. J Cell Biol 193:81–95. doi:10.1083/jcb.201101030
Ball AR, Yokomori K (2009) Revisiting the role of heterochromatin protein 1 in DNA repair. J Cell Biol 185:573–575. doi:10.1083/jcb.200904033
Bartova E, Galiova G, Legartova S et al (2010) Genome instability in the context of chromatin structure and fragile sites. Crit Rev Eukaryot Gene Expr 20:181–194
Bastiaens PI, Pepperkok R (2000) Observing proteins in their natural habitat: the living cell. Trends Biochem Sci 25:631–637
Cheutin T, McNairn AJ, Jenuwein T et al (2003) Maintenance of stable heterochromatin domains by dynamic HP1 binding. Science 299:721–725
Ciccia A, Elledge SJ (2010) The DNA damage response: making it safe to play with knives. Mol Cell 40:179–204. doi:10.1016/j.molcel.2010.09.019
Dinant C, Luijsterburg MS (2009) The emerging role of HP1 in the DNA damage response. Mol Cell Biol 29:6335–6340. doi:10.1128/MCB.01048-09
Dundr M (2012) Nuclear bodies: multifunctional companions of the genome. Curr Opin Cell Biol 24:415–422. doi:10.1016/j.ceb.2012.03.010
Fanti L, Pimpinelli S (2008) HP1: a functionally multifaceted protein. Curr Opin Genet Dev 18:169–174. doi:10.1016/j.gde.2008.01.009
Geuting V, Reul C, Löbrich M (2013) ATM release at resected double-strand breaks provides heterochromatin reconstitution to facilitate homologous recombination. PLoS Genet 9:e1003667. doi:10.1371/journal.pgen.1003667
Goodarzi AA, Noon AT, Deckbar D et al (2008) ATM signaling facilitates repair of DNA double-strand breaks associated with heterochromatin. Mol Cell 31:167–177. doi:10.1016/j.molcel.2008.05.017
Hartlerode AJ, Scully R (2009) Mechanisms of double-strand break repair in somatic mammalian cells. Biochem J 423:157–168. doi:10.1042/BJ20090942
Hoeijmakers JH (2001) Genome maintenance mechanisms for preventing cancer. Nature 411:366–374. doi:10.1038/35077232
Hoek M, Stillman B (2003) Chromatin assembly factor 1 is essential and couples chromatin assembly to DNA replication in vivo. Proc Natl Acad Sci U S A 100:12183–12188. doi:10.1073/pnas.1635158100
Hu CD, Chinenov Y, Kerppola TK (2002) Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol Cell 9(4):789–798. doi:10.1016/S1097-2765(02)00496-3
Hu C, Zhang S, Gao X et al (2012) Roles of Krüppel-associated box (KRAB)-associated co-repressor KAP1 Ser-473 phosphorylation in DNA damage response. J Biol Chem 287:18937–18952. doi:10.1074/jbc.M111.313262
Khurana S, Kruhlak MJ, Kim J et al (2014) A macrohistone variant links dynamic chromatin compaction to BRCA1-dependent genome maintenance. Cell Rep 8:1049–1062. doi:10.1016/j.celrep.2014.07.024
Kruhlak MJ, Celeste A, Dellaire G et al (2006) Changes in chromatin structure and mobility in living cells at sites of DNA double-strand breaks. J Cell Biol 172:823–834. doi:10.1083/jcb.200510015
Kruhlak M, Crouch EE, Orlov M et al (2007) The ATM repair pathway inhibits RNA polymerase I transcription in response to chromosome breaks. Nature 447:730–734. doi:10.1038/nature05842
Lee Y-H, Kuo C-Y, Stark JM et al (2013) HP1 promotes tumor suppressor BRCA1 functions during the DNA damage response. Nucleic Acids Res 41:5784–5798. doi:10.1093/nar/gkt231
LeRoy G, Weston JT, Zee BM et al (2009) Heterochromatin protein 1 is extensively decorated with histone code-like post-translational modifications. Mol Cell Proteomics 8:2432–2442. doi:10.1074/mcp.M900160-MCP200
Lieber MR (2010) The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem 79:181–211. doi:10.1146/annurev.biochem.052308.093131
Liu B, Wang Z, Zhang L et al (2013a) Depleting the methyltransferase Suv39h1 improves DNA repair and extends lifespan in a progeria mouse model. Nat Commun 4:1868. doi:10.1038/ncomms2885
Liu H, Galka M, Mori E et al (2013b) A method for systematic mapping of protein lysine methylation identifies functions for HP1β in DNA damage response. Mol Cell 50:723–735. doi:10.1016/j.molcel.2013.04.025
Lomberk G, Wallrath L, Urrutia R (2006) The heterochromatin protein 1 family. Genome Biol 7:228. doi:10.1186/gb-2006-7-7-228
Luijsterburg MS, Dinant C, Lans H et al (2009) Heterochromatin protein 1 is recruited to various types of DNA damage. J Cell Biol 185:577–586. doi:10.1083/jcb.200810035
Maison C, Almouzni G (2004) HP1 and the dynamics of heterochromatin maintenance. Nat Rev Mol Cell Biol 5:296–304. doi:10.1038/nrm1355
Miller KM, Tjeertes JV, Coates J et al (2010) Human HDAC1 and HDAC2 function in the DNA-damage response to promote DNA nonhomologous end-joining. Nat Struct Mol Biol 17:1144–1151. doi:10.1038/nsmb.1899
Moynahan ME, Jasin M (2010) Mitotic homologous recombination maintains genomic stability and suppresses tumorigenesis. Nat Rev Mol Cell Biol 11:196–207. doi:10.1038/nrm2851
Noon AT, Shibata A, Rief N et al (2010) 53BP1-dependent robust localized KAP-1 phosphorylation is essential for heterochromatic DNA double-strand break repair. Nat Cell Biol 12:177–184. doi:10.1038/ncb2017
Pardo B, Gómez-González B, Aguilera A (2009) DNA repair in mammalian cells: DNA double-strand break repair: how to fix a broken relationship. Cell Mol Life Sci 66:1039–1056. doi:10.1007/s00018-009-8740-3
Rogakou EP, Pilch DR, Orr AH et al (1998) DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J Biol Chem 273:5858–5868
Ryan RF, Schultz DC, Ayyanathan K et al (1999) KAP-1 corepressor protein interacts and colocalizes with heterochromatic and euchromatic HP1 proteins: a potential role for Krüppel-associated box-zinc finger proteins in heterochromatin-mediated gene silencing. Mol Cell Biol 19:4366–4378
San Filippo J, Sung P, Klein H (2008) Mechanism of eukaryotic homologous recombination. Annu Rev Biochem 77:229–257. doi:10.1146/annurev.biochem.77.061306.125255
Shanbhag NM, Rafalska-Metcalf IU, Balane-Bolivar C et al (2010) ATM-dependent chromatin changes silence transcription in cis to DNA double-strand breaks. Cell 141:970–981. doi:10.1016/j.cell.2010.04.038
Sorokin DV, Stixová L, Sehnalová P et al (2015) Localized movement and morphology of UBF1-positive nucleolar regions are changed by γ-irradiation in G 2 phase of the cell cycle. Nucleus 6:301–313. doi:10.1080/19491034.2015.1075111
Stixová L, Sehnalová P, Legartová S et al (2014a) HP1β-dependent recruitment of UBF1 to irradiated chromatin occurs simultaneously with CPDs. Epigenetics Chromatin 7:39. doi:10.1186/1756-8935-7-39
Stixová L, Hrušková T, Sehnalová P, Legartová S, Svidenská S, Kozubek S, Bártová E (2014b) Advanced microscopy techniques used for comparison of UVA- and γ-irradiation-induced DNA damage in the cell nucleus and nucleolus. Folia Biol (Praha) 60(Suppl 1):76–84
Suchánková J, Kozubek S, Legartová S, et al (2015) Distinct kinetics of DNA repair protein accumulation at DNA lesions and cell cycle-dependent formation of γH2AX- and NBS1-positive repair foci. Biol Cell 0:1–15. doi: 10.1111/boc.201500050
Sulli G, Di Micco R, di d’Adda FF (2012) Crosstalk between chromatin state and DNA damage response in cellular senescence and cancer. Nat Rev Cancer 12:709–720. doi:10.1038/nrc3344
Šustáčková G, Kozubek S, Stixová L et al (2012) Acetylation-dependent nuclear arrangement and recruitment of BMI1 protein to UV-damaged chromatin. J Cell Physiol 227:1838–1850. doi:10.1002/jcp.22912
Trembecka-Lucas DO, Dobrucki JW (2012) A heterochromatin protein 1 (HP1) dimer and a proliferating cell nuclear antigen (PCNA) protein interact in vivo and are parts of a multiprotein complex involved in DNA replication and DNA repair. Cell Cycle 11:2170–2175. doi:10.4161/cc.20673
Trembecka-Lucas DO, Szczurek AT, Dobrucki JW (2013) Dynamics of the HP1β-PCNA-containing complexes in DNA replication and repair. Nucleus 4:74–82. doi:10.4161/nucl.23683
Wang D, Zhou J, Liu X, Lu D, Shen C, Du Y, Wei FZ, Song B, Lu X, Yu Y, Wang L, Zhao Y, Wang H, Yang Y, Akiyama Y, Zhang H, Zhu WG (2013) Methylation of SUV39H1 by SET7/9 results in heterochromatin relaxation and genome instability. Proc Natl Acad Sci U S A 110:5516–5521. doi:10.1073/pnas.1216596110
White D, Rafalska-Metcalf IU, Ivanov AV et al (2012) The ATM substrate KAP1 controls DNA repair in heterochromatin: regulation by HP1 proteins and serine 473/824 phosphorylation. Mol Cancer Res 10:401–414. doi:10.1158/1541-7786.MCR-11-0134
Wiernasz E (2004) Recruitment of heterochromatin protein 1 to sites of photooxidative chromatin damage. MSc Thesis, Jagiellonian University, Krakow, Poland
Zarebski M (2008) Method of inflicting local photooxidative damage in live cells and application to studies of repair of plasma memebranes and chromatin. PhD Thesis, Jagiellonian University, Krakow, Poland
Zarebski M, Wiernasz E, Dobrucki JW (2009) Recruitment of heterochromatin protein 1 to DNA repair sites. Cytometry A 75:619–625. doi:10.1002/cyto.a.20734
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was supported by the Czech Science Foundation grant number 302–12-G157. The research leading to these results received funding from the Norwegian Financial Mechanism 2009–2014 under Project Contract No. 7F14369.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Reimer Stick
Rights and permissions
About this article
Cite this article
Bártová, E., Malyšková, B., Komůrková, D. et al. Function of heterochromatin protein 1 during DNA repair. Protoplasma 254, 1233–1240 (2017). https://doi.org/10.1007/s00709-017-1090-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-017-1090-3