Abstract
Several isotope-labeling strategies have been developed for the study of RNA by nuclear magnetic resonance (NMR) spectroscopy. Here, we report a combined chemical and enzymatic synthesis of [7-15N]-guanosine-5′-triphosphates for incorporation into RNA via T7 RNA polymerase-based in vitro transcription. We showcase the utility of these labels to probe both structure and dynamics in two biologically important RNAs.
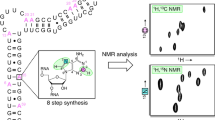
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
RNAs, once thought of as an intermediate in the flow of genetic information from DNA to proteins, are now credited with playing a central role in many cellular functions [1,2,3,4]. As a result, RNAs have increasingly become the target of structural and therapeutic efforts [5,6,7]. Among the different techniques available to study RNA, nuclear magnetic resonance (NMR) spectroscopy is particularly useful [8, 9]. NMR, unlike X-ray crystallography, permits the study of RNA structure and dynamics in solution on a wide-range of timescales spanning picoseconds-to-seconds [10]. Nevertheless, poor chemical shift dispersion and broad linewidths limit the broad application of NMR to understand the structure and dynamics of RNA [11].
Several technological advances have begun to address these problems: use of cryogenic probes, higher magnetic field spectrometers, and the design of multidimensional NMR experiments [12,13,14,15,16,17]. Even with these innovations, spectral crowding and signal overlap still limit the effective analysis of large (> 30 nucleotide, nt) RNAs [18]. To tackle this problem, we and others have developed labeling technologies to incorporate stable isotopes (i.e., 2H, 13C, 19F, and 15N) into RNA to benefit NMR structure and dynamics measurements [7, 19,20,21,22,23,24,25,26,27].
As part of a larger effort to create atom-specifically labeled ribonucleoside-5′-triphosphates (rNTPs) [7, 20,21,22,23,24,25,26,27], we present the chemical synthesis of [7-15N]-guanine from commercially available and inexpensive sodium 15N-nitrite. Then, we used enzymes from the nucleotide salvage biosynthetic pathway to couple the labeled nucleobase to commercially available ribose sources to build the corresponding [7-15N]-guanosine-5′-triphosphates (GTPs) [20, 28, 29]. This labeling scheme leverages the narrow linewidths of the 15N nuclei [30, 31] and will therefore be important when applied to large RNAs. Moreover, the N7 atom in the major groove can serve as a reporter of binding events between nucleic acids and ligands such as metals, drugs, or protein side chains [32, 33].
To showcase the utility of our labels, we incorporated our [7-15N]-GTPs into two RNAs [33, 34] via T7 RNA polymerase (RNAP)-based in vitro transcription and used one (1D)-, two (2D)-, and three (3D)-dimensional NMR experiments to probe RNA structure and dynamics.
Results and discussion
Synthesis
Synthesis of the isotope-labeled nucleobase was carried out in three chemical steps following previously established methods [35] with slight modifications (Scheme 1). The synthesis of [7-15N]-guanine began with the nitrosylation of commercially available 2,6-diaminopyrimidin-4-ol by 15N-labeled sodium nitrite to yield 2,6-diamino-5-[nitroso-15N]pyrimidin-4-ol (1). Reduction by sodium dithionite followed by acidification by sulfuric acid then gave the sulfate salt of 2,6-diamino-5-[amino-15N]pyrimidin-4-ol (2). In the final chemical step, reflux of 2 with formamide followed by formic acid provided the desired [7-15N]-guanine (3). Intermediate compounds 1 and 2 showed the expected 1H NMR spectra (Figs. S1, S2). The formation and purity of 3 was confirmed by 1H, 13C, and 15N NMR (Figs. S3, S4) and high-resolution mass spectroscopy (HR-MS) (Fig. S5).

With the labeled nucleobase 3 in-hand, we utilized enzymes from the nucleotide salvage biosynthetic pathway to couple 3 to either fully protonated [1H6]-d-ribose or fully deuterated [2H6]-d-ribose to form the desired GTPs 4a and 4b, respectively (Scheme 1), as previously described [20]. Briefly, [1H6]- and [2H6]-d-ribose were phosphorylated to ribose-5-phosphate (R5P) by ribokinase (RK, EC 2.7.1.15). Then, R5P was converted to 5-phospho-d-ribosyl-α-1-pyrophosphate (5PRP) by phosphoribosyl pyrophosphate synthetase (PRPPS, EC 2.7.6.1). Coupling of 3 to 5PRP from the previous step was afforded by xanthine-guanine phosphoribosyltransferases (XGPRT, EC 2.4.2.22) to yield [7-15N]-guanosine-5′-monophosphate ([7-15N]-GMP). Guanylate kinase (GK, EC 2.7.4.8) converted GMP to [7-15N]-guanosine-5′-diphosphate ([7-15N]-GDP), which was then phosphorylated by creatine kinase (CK, EC 2.7.3.2) to form the desired GTP products 4a and 4b. Complete conversion of 3 to 4a and 4b was confirmed by 31P NMR (Fig. S6), as previously described [36].
Taken together, our synthetic route provides 4a and 4b in three chemical steps and one enzymatic step with a single chromatographic purification. Furthermore, while the present work used [1H6]- and [2H6]-d-ribose, our method enables the coupling of 3 to any ribose source to create a versatile assortment of atom-specifically labeled rNTPs for use in in vitro transcription and NMR.
NMR characterization of atom-specifically labeled RNA
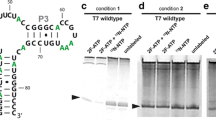
Our reason for synthesizing 4a and 4b was to characterize the structure and dynamics of biologically important RNAs. Thus, we used 4a and 4b along with unlabeled ATP, CTP, and UTP to make two RNAs by in vitro transcription: a 4b-labeled 27 nt fragment from the human cytoplasmic A-site ribosomal RNA (A-site) and a 4a-labeled 35 nt fragment from domain 5 of the group II intron ribozyme from brown algae (domain 5) (Fig. 1).
As a first application, we employed a two-bond (2JH8N7) 2D heteronuclear single quantum coherence (HSQC) experiment on 4a-labeled domain 5 RNA. We obtained a well-resolved 2D spectrum, showing all 13 guanosine H8-N7 resonances (Fig. 2a). A necessary NMR parameter for structure determination is proton–proton distances, which is provided by nuclear Overhauser effect spectroscopy (NOESY) experiments. While these data are informative, crowded proton–proton NOEs can be resolved into a third dimension with 13C- or 15N-editing. As a second application, we employed a 15N-edited 3D NOESY HSQC experiment on 4a-labeled domain 5 to reveal all protons within ~ 5 Å of guanosine H8 [38]. In A-helical RNA, guanosine H8 protons show strong NOE cross-peaks to 5′-neighboring H2′ protons (Fig. 2b), which are traditionally difficult to assign due to severe overlap with other ribose protons (i.e., H3′, H4′, H5′, and H5″) [37,38,39]. We therefore used the chemical shift of guanosine N7 to resolve NOE cross-peaks of H2′ protons to H8. Examples of cross-peaks are shown for helical residues G8 and G22, as well as residues G15 and G26 from the apical loop and internal bulge, respectively (Fig. 2c).
HSQC and 15N-edited NOESY HSQC experiments in 4a-labeled domain 5 RNA. a 2D 1H,15N HSQC spectra showing H8-N7 resonances. b Representation of NOE contacts of H8 protons to H2′ protons for select nucleotides G8, G15, G22, and G26. c 2D 1H,1H slice along a single 15N frequency for the same select nucleotides shown in b. All spectra are annotated with RNA resonance assignments
As a final application, we probed the dynamics of 4b-labeled A-site RNA. Two common relaxation parameters in biomolecules are the longitudinal (R1) and transverse (R2) relaxation rates [40, 41]. While R1 measures the rate of recovery of the z-magnetization to equilibrium, R2 reports on the rate of the decay of x- and y-magnetization [40]. An alternative method to obtain R2 is a transverse rotating-frame (R1ρ) experiment wherein magnetization is aligned along an effective field whose direction is dependent upon the power of the radio frequency (RF) field and its offset [42]. We therefore employed pseudo-2D HSQC-based experiments to determine R1 and R1ρ relaxation rates of guanosine H8 protons (Fig. 3a). We obtained rates for 5 of the 9 guanosines in A-site, with R1 and R1ρ values ranging from 1.75 to 2.00/s and 15.26 to 21.25/s, respectively. Interestingly, G16 showed the highest R1 and lowest R1ρ, indicative of increased flexibility, suggesting the G:C base pair preceding the tetraloop is not stable. These data contribute to our understanding of the dynamic motions within the A-site RNA [43, 44].
Conclusion
We report the synthesis of atom-specifically labeled [7-15N]-GTPs for use in in vitro transcription to make RNA for NMR analysis. Our synthetic routes include a combined chemical and enzymatic approach, using inexpensive commercially available starting materials. To demonstrate the utility of our new labels, we introduced them into two RNAs via in vitro transcription to permit straightforward NMR structure and dynamics measurements. We anticipate these labels will aid efforts to probe in greater detail the structure, interactions, and dynamics of biologically and medically RNAs.
Experimental
Commercially available reagents were used without further purification unless explicitly stated. All reagents used for the synthesis of 3 were purchased from Sigma-Aldrich. All reactions were carried out under nitrogen or argon atmosphere. All non-commercially available enzymes were expressed and purified in-house using established methods [28]. DNA templates for in vitro transcription of RNAs were purchased from Integrated DNA Technologies (IDT, Coralville, IA) and used without further purification. Chromatographic purification was carried out using boronate affinity resin with eluent specified. 1H NMR spectra were recorded on a Bruker Avance I 300 MHz or a Bruker Avance Neo 400 MHz spectrometer, 13C NMR spectra were recorded on a Bruker Avance III 500 MHz spectrometer, 15N NMR spectra were recorded on a Bruker Avance III 700 MHz spectrometer, and 31P NMR spectra were recorded on an Avance III Bruker 800 MHz spectrometer with a triple resonance cryogenic probe. Samples were maintained at a temperature of 25 °C. All NMR experiments for RNA were performed in D2O and all chemical shifts were reported in ppm (parts per million). All RNA spectra were referenced to DSS (4,4-dimethyl-4-silapentane-1-sulfonic acid). Nitrogen-15 and Carbon-13 chemical shifts were indirectly referenced using the ratio of the gyromagnetic ratios of proton to 15N (0.101329118) and 13C (0.251449530), respectively [45, 46]. All NMR experiments for compounds 1, 2, and 3 were performed in DMSO-d6 or D2O. The chemical shifts of compounds 1, 2, and 3 were referenced to the residual protonated solvent signal of DMSO-d6 (2.5 ppm) or D2O (HDO 4.7 ppm) as previously reported [47]. The 15N dimension was referenced using the liquid ammonia referencing implemented in the Topspin software suite.
2,6-Diamino-5-[nitroso-15N]pyrimidin-4-ol (1, C4H5N4 15NO2)
To begin the chemical synthesis of [7-15N]-guanine, 3.65 g 2,6-diaminopyrimidin-4-ol (29.00 mmol) and 2.30 g 15N-labeled sodium nitrite (33.30 mmol) were dissolved in 40 cm3 of 3 M sodium hydroxide. The homogeneous solution obtained from the previous step was then added dropwise to 50 cm3 glacial acetic acid, while stirring and cooling on ice, and gave rise to a pink precipitate. The precipitate was isolated by centrifugation, washed with cold water, ethanol, and diethyl ether, and dried in high vacuum (1 × 10–2 mbar) on a vacuum line for 8 h to give pure compound 1. Yield: 3.55 g (79%); 1H NMR (300 MHz, DMSO-d6): δ = 7.34 (br s, 4H) ppm; HR-MS (ESI–MS): m/z calculated for C4H5N415N1O2 + H+ 157.0486 Da, found 157.0486 Da.
2,6-Diamino-5-[amino-15N]pyrimidin-4-ol (2, C4H7N4 15NO)
After vacuum drying, compound 1 was resuspended in 80 cm3 boiling water and 9.96 g sodium dithionite (57.23 mmol) was added in several portions to give a pink suspension. The reaction was kept at 100 °C and the pink suspension became yellow in color. The mixture was then cooled in an ice bath for 30 min. The yellow solid was collected by filtration and then resuspended in 65 cm3 of 2 M sulfuric acid, heated to 80 °C for 20 min, and cooled on ice to give a sulfate salt. This salt was collected by filtration, washed with ethanol, and dried in high vacuum (1 × 10–2 mbar) on a vacuum line for 12 h to give pure compound 2. Yield: 5.45 g (22.8 mmol); 1H NMR (400 MHz, DMSO-d6): δ = 8.94 (br s, 2H), 6.46 (br s, 2H), 6.34 (br s, 2H) ppm; HR-MS (ESI–MS): m/z calculated for C4H7N415N1O1 + H+ 143.0694 Da, found 143.0693 Da.
[7-15N]-2-Amino-1H-purin-6(9H)-one (3, C5H5N4 15NO)
Compound 2 was dissolved in 30 cm3 formamide and formic acid and then refluxed at 110 °C and 180 °C for 1 and 2 h, respectively, to form a yellow solution. After cooling on ice for 30 min, a milky precipitate formed. The precipitate was filtered, rinsed with water, and then dissolved in 35 cm3 10% aqueous sodium hydroxide. Crude product 3 was precipitated by neutralizing the solution on ice with formic acid. A fine, yellowish powder of crude compound 3 was obtained by filtering the precipitate, which was then washed with water and ethanol to remove any residual formamide contaminant and dried in high vacuum (1 × 10–2 mbar) on a vacuum line for 6 h to yield pure compound 3. Yield: 2.85 g (83%); 1H NMR (400 MHz, DMSO-d6): δ = 8.89 (d, 2JNH = 5.92 Hz, 1H) ppm; 15N NMR (60 MHz, DMSO-d6): δ = 241.39 (15N7) ppm; 1H NMR (700 MHz, 10 mM NaOD in D2O): δ = 7.49 (d, 2JNH = 12 Hz, 1H) ppm; 13C NMR (176 MHz, 10 mM NaOD in D2O): δ = 167.53 (C4), 160.94 (C6), 159.11 (C2), 148.56 (C8), 118.24 (C5) ppm; 15N NMR (70 MHz, 10 mM NaOD in D2O): δ = 221.83 (15N7) ppm; HR-MS (ESI–MS): m/z calculated for C5H5N415N1O1 + H+ 153.0537 Da, found 153.0537 Da.
[7-15N]-[[[[(2R,3S,4R,5R)-5-(2-Amino-6-oxo-1,6-dihydro-9H-purin-9-yl)-3,4-dihydroxytetrahydrofuran-2-yl]methoxy]oxidophosphoryl]oxidophosphoryl]phosphonate (4a, C10H12N4 15NO12P3 4−)
The one-pot reaction was carried out in a 50 cm3 Falcon tube. The reaction mixture consisted of 9.1 mg compound 3 (6.0 mM), fully protonated [1H6]-d-ribose (6 mM), 10 mM MgCl2, deoxyadenosine-5′-triphosphate (0.5 mM), Bovine serum albumin (0.1 mg/cm3), dithiothreitol (10 mM), potassium chloride (100 mM), sodium phosphate monobasic (9.4 mM, pH 6.5), sodium phosphate dibasic (40 mM, pH 6.5), RK (1.0 × 10–5 U/mm3), PRPPS (1.0 × 10–5 U/mm3), XGPRT (0.01 mg/cm3), GK (0.01 mg/cm3), MK (0.01 U/mm3), and CK (0.05 mg/cm3). The reaction was incubated at 37 °C for 12 h. After confirming successful triphosphate formation by 31P NMR, crude compound 4a was purified by boronate affinity chromatography (eluent A: 1 M triethylamine pH 9; eluent B: acidified water pH 4), lyophilized to a powder, and resuspended in Ultrapure water. Yield: 6.8 mg (~ 90%).
[1′,2′,3′,4′,5′,5″-2H6,7-15N]-[[[[(2R,3S,4R,5R)-5-(2-Amino-6-oxo-1,6-dihydro-9H-purin-9-yl)-3,4-dihydroxytetrahydrofuran-2-yl]methoxy]oxidophosphoryl]oxidophosphoryl]phosphonate (4b, C10H6 2H6N4 15NO12P3 4−)
The one-pot reaction was carried out in a 50 cm3 Falcon tube. The reaction mixture consisted of 9.1 mg compound 3 (6.0 mM), fully deuterated [2H6]-d-ribose (6 mM), 10 mM MgCl2, deoxyadenosine-5′-triphosphate (0.5 mM), Bovine serum albumin (0.1 mg/cm3), dithiothreitol (10 mM), potassium chloride (100 mM), sodium phosphate monobasic (9.4 mM, pH 6.5), sodium phosphate dibasic (40 mM, pH 6.5), RK (1.0 × 10–5 U/mm3), PRPPS (1.0 × 10–5 U/mm3), XGPRT (0.01 mg/cm3), GK (0.01 mg/cm3), MK (0.01 U/mm3), and CK (0.05 mg/cm3). The reaction was incubated at 37 °C for 12 h. After confirming successful triphosphate formation by 31P NMR, crude compound 4b was purified by boronate affinity chromatography (eluent A: 1 M triethylamine pH 9; eluent B: acidified water pH 4), lyophilized to a powder, and resuspended in ultrapure water. Yield: 2.5 mg (~ 33%). The low yield of this reaction compared to that of 4a was due to complications with purification, not with conversion of 3 to 4b.
RNA preparation
RNAs were synthesized via in vitro transcription. The reactions were carried out in a 10 cm3 reaction volume at 37 °C. The reactions consisted of transcription buffer (40 mM Tris–HCl pH 8.0, 1 mM spermidine and 0.01% Triton-10), 0.3 µM single strand DNA template, 80 mg/cm3 PEG, 1 mM DTT, 2 U/mm3 thermostable inorganic pyrophosphatase, 10 mg/cm3 T7 RNA polymerase, a total of 5 mM rNTPs (1.25 mM each of unlabeled ATP, CTP, UTP, and either 4a or 4b), and 7.5 mM MgCl2. The concentrations of rNTPs and MgCl2 were chosen following optimization at small (50 mm3) and mid-scale (500 mm3) reactions. All reactions were quenched after 3 h by adding 0.5 mM EDTA. Following transcription, the RNAs were purified via preparative polyacrylamide (12%) gel electrophoresis, electroeluted, and exchanged into deionized water. Then NMR buffer (A-site: 50 mM NaCl, 50 mM Na3PO4, 100 mM EDTA, 0.02% NaN3, 0.1 mM DSS, 10% D2O; domain 5: 100 mM KCl, 10 mM K3PO4, 0.02% NaN3, 0.1 mM DSS, 10% D2O) was added and the samples were lyophilized and resuspended in D2O (99.9%). The concentration of the purified samples ranged between 0.3 and 0.5 mM in 300 mm3.
NMR experiments
All experiments on 4a- and 4b-labeled samples were carried out at 25 °C on a Bruker 600 MHz magnet Avance III spectrometer with a TXI triple resonance probe in 5 mM Bruker optimized Shigemi NMR tubes. 2D-15N Selective Optimized Flip Angle Short Transient (SOFAST) [48] HSQC spectrum of compound 3 was obtained with 16 scans and 1024 × 64 complex points in the 1H and 15N dimensions, respectively. An INEPT delay of 31.25 ms (a coupling constant of 8 Hz) was used for coherence transfer. For the 1H dimension, the spectral width was set to 20 ppm and the carrier to 12 ppm. For the 15N dimension, the spectral width was set to 5 ppm and the carrier to 240 ppm. The two-bond (2JH8N7) 2D HSQC experiment was performed with 128 scans and 100 and 1024 complex points in the 15N and 1H dimensions, respectively. The 15N and 1H carrier frequencies were set to 235.5 ppm and 4.7 ppm, respectively; the spectral width for the 15N and 1H dimensions were set to 9 and 12 ppm, respectively. The 15N-edited 3D NOESY HSQC experiment was carried out on a 600 MHz magnet Avance III spectrometer with a TXI triple resonance probe using previously described pulse sequences [38]. For each 3D data set, 200 × 128 complex points were used for the indirect 1H and 15N dimensions along with 64 transients and 10% non-uniform sine-weighted Poisson-gap sampling [49]. The spectral width of the 15N and 1H dimensions were set to 5.2 and 10.0 ppm, respectively; the carrier was set to 235.1 and 4.7 ppm, for the 15N and 1H dimensions, respectively.
R1 and R1ρ relaxation rates on the guanosine-N7 nitrogen atoms were measured using a pseudo-2D HSQC detected experiments. These experiments were carried out with 16 scans and 1024 time domain points in the 1H dimension, using a spectral width and carrier of 16 and 4.7 ppm, respectively. For the R1 experiments, delay times of 0.02 (X2), 0.04, 0.08, 0.15, 0.30, 0.40, 0.60, 0.80, and 1.00 s were used. For the R1ρ experiments, delay times of 8 (X2), 16, 20, 32, 48, 60, 80, and 100 ms were used. The strength of spin-lock field (ω1) was 1.9 kHz. Calibration of the spin-lock field was carried out as previously described [50]. R1 and R1ρ relaxation rates were determined by fitting intensities to a monoexponential decay and errors were estimated from duplicated delay points. All NMR data were collected at 25 °C with a recycle delay of 1.5 s and analyzed using TopSpin 4.0 and NMRViewJ [51].
References
Abernathy E, Glaunsinger B (2015) Virology 479:600
Serganov A, Patel DJ (2007) Nat Rev Genet 8:776
Lau NC, Lai EC (2005) Genome Biol 6:315
Shang BQ, Li ML, Quan HY, Hou PF, Li ZW, Chu SF, Zheng JN, Bai J (2019) Mol Cancer 18:1
Schoenmaker L, Witzigmann D, Kulkarni JA, Verbeke R, Kersten GF, Jiskoot W, Crommelin DJ (2021) Int J Pharm 601:120586
Abulwerdi FA, Xu W, Ageeli AA, Yonkunas MJ, Arun G, Nam H, Schneekloth JS Jr, Dayie TK, Spector D, Baird N, Le Grice SF (2019) ACS Chem Biol 14:223
Becette OB, Zong G, Chen B, Taiwo KM, Case DA, Dayie TK (2020) Sci Adv 6:eabc6572
Barnwal RP, Yang F, Varani G (2017) Arch Biochem Biophys 628:42
Zhang H, Keane SC (2019) RNA 10:e1541
Liu B, Shi H, Al-Hashimi HM (2021) Curr Opin Struct Biol 70:16
Dayie TK (2008) Int J Mol Sci 9:1214
Alderson TR, Kay LE (2021) Cell 184:577
Luchinat E, Barbieri L, Cremonini M, Banci L (2021) J Biomol NMR 75:97
Banci L, Barbieri L, Calderone V, Cantini F, Cerofolini L, Ciofi-Baffoni S, Felli IC, Fragai M, Lelli M, Luchinat C, Luchinat E (2019) arXiv:1910.07462
Abramov G, Velyvis A, Rennella E, Wong LE, Kay LE (2020) Proc Natl Acad Sci 117:12836
Latham MP, Brown DJ, McCallum SA, Pardi A (2005) ChemBioChem 6:1492
Schnieders R, Keyhani S, Schwalbe H, Fürtig B (2020) Chemistry 26:102
Allain FHT, Varani G (1997) J Mol Biol 267:338
Nußbaumer F, Plangger R, Roeck M, Kreutz C (2020) Angew Chemie Int Ed 59:17062
Longhini AP, LeBlanc RM, Becette O, Salguero C, Wunderlich CH, Johnson BA, D’Souza VM, Kreutz C, Dayie TK (2015) Nucleic Acids Res 44:52
Olenginski LT, Dayie TK (2020) Monatsh Chem 151:1467
Taiwo KM, Becette OB, Zong G, Chen B, Zavalij PY, Dayie TK (2021) Monatsh Chem 152:441
Alvarado LJ, LeBlanc RM, Longhini AP, Keane SC, Jain N, Yildiz ZF, Tolbert BS, D’Souza VM, Summers MF, Kreutz C, Dayie TK (2014) ChemBioChem 15:1573
Alvarado LJ, Longhini AP, LeBlanc RM, Chen B, Kreutz C, Dayie TK (2014) Methods Enzymol 549:133
LeBlanc RM, Longhini AP, Le Grice SF, Johnson BA, Dayie TK (2017) Nucleic Acids Res 45:e146
LeBlanc RM, Longhini AP, Tugarinov V, Dayie TK (2018) J Biomol NMR 71:165
Longhini AP, LeBlanc RM, Dayie TK (2016) Methods 103:11
Arthur PK, Alvarado LJ, Dayie TK (2011) Protein Expr Purif 76:229
Tolbert TJ, Williamson JR (1996) J Am Chem Soc 118:7929
Marchant J, Bax A, Summers MF (2018) J Am Chem Soc 140:6978
Schnieders R, Richter C, Warhaut S, de Jesus V, Keyhani S, Duchardt-Ferner E, Keller H, Wöhnert J, Kuhn L, Breeze AL, Bermerl W, Schalbe H, Fürtig B (2017) J Biomol NMR 69:31
Milecki J (2002) J Label Compd Radiopharm 45:307
Seetharaman M, Eldho NV, Padgett RA, Dayie TK (2006) RNA 12:235
Kondo J, Urzhumtsev A, Westhof E (2006) Nucleic Acids Res 34:676
Lolli M, Medana C, Romagnano S, Castoldi F, Pozzoli S, Vago F, Fanelli R, Airoldi L (1998) J Label Compd Radiopharm 41:243
Spoerner M, Karl M, Lopes P, Hoering M, Loeffel K, Nuehs A, Adelsberger J, Kremer W, Kalbitzer HR (2017) J Biomol NMR 67:1
Wijmenga SS, van Buuren BNM (1998) Prog Nucl Magn Reson Spectrosc 32:287
Hoffman DW (2000) J Biomol NMR 16:165
Dayie KT, Tolbert TJ, Williamson JR (1998) J Magn Reson 130:97
Palmer AG (2004) Chem Rev 104:3623
Dayie KT, Brodsky AS, Williamson JR (2002) J Mol Biol 317:263
Rangadurai A, Szymaski ES, Kimsey IJ, Shi H, Al-Hashimi HM (2019) Prog Nucl Magn Reson Spectrosc 112:55
Lynch SR, Puglisi JD (2001) J Mol Biol 306:1023
Vaiana AC, Westhof E, Auffinger P (2006) Biochimie 88:1061
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995) J Biomol NMR 6:135
Aeschbacher T, Schubert M, Allain FH (2012) J Biomol NMR 52:179
Gottlieb HE, Kotlyar V, Nudelman A (1997) J Org Chem 62:7512
Schanda P, Brutscher B (2005) J Am Chem Soc 127:8014
Hyberts SG, Takeuchi K, Wagner G (2010) J Am Chem Soc 132:2145
Nam H, Becette O, LeBlanc RM, Oh D, Case DA, Dayie TK (2020) J Biomol NMR 74:321
Johnson BA, Blevins RA (1994) J Biomol NMR 4:603
Acknowledgements
This research was financially supported by the National Science Foundation (DBI1040158 to T.K.D.), the National Institutes of Health (U54AI50470 to T.K.D.), the Austrian Science Fund (FWF, project P34770 and P32773 to C.K), and the Austrian Research Promotion Agency FFG (West Austrian BioNMR, 858017 to C.K). We acknowledge Thomas Mueller (University of Innsbruck) for mass spectrometric analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Taiwo, K.M., Olenginski, L.T., Nußbaumer, F. et al. Synthesis of [7-15N]-GTPs for RNA structure and dynamics by NMR spectroscopy. Monatsh Chem 153, 293–299 (2022). https://doi.org/10.1007/s00706-022-02892-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00706-022-02892-1