Abstract
A new series of xanthone derivatives against the oral human epidermoid carcinoma (KB) cancer cell line is examined to determine the relationship between the structural properties and the biological activity of these compounds—the 3-D quantitative structure–activity relationship (3D-QSAR)—using comparative molecular field analysis (CoMFA) and comparative molecular similarity indices analysis (CoMSIA). The best CoMFA and CoMSIA models were obtained using the atom-based alignment of 33 compounds, 22 training compounds and 11 tested compounds, and these give desirable statistics; those for the CoMFA standard model were: r 2cv = 0.691, r 2 = 0.998, S press = 0.178, s = 0.014 and F = 1080.765, while CoMSIA combined steric, electrostatic, hydrophobic and hydrogen-bond acceptor fields: r 2cv = 0.600, r 2 = 0.988, S press = 0.206, s = 0.034 and F = 284.433. The 3D-QSAR models calculated satisfactory test set activities. The 3D-QSAR contour plots correlated strongly with the experimental data for the binding topology. For this reason, these results would be beneficial for predicting affinities with the compounds of interest, and they are advantageous for guiding the design and synthesis of new and more effective anticancer agents.
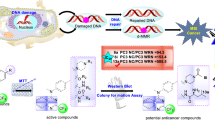
Graphical abstract

A new and more effective anticancer agent of xanthone derivatives against the oral human epidermoid carcinoma (KB) cell line, as investigated by CoMFA and CoMSIA analysis





Similar content being viewed by others
References
Pravda (2006) Soon cancer to be treated with protons. http://english.pravda.ru/news/science/22-08-2006/84027-proton-0. Last accessed 21 June 2008
Bilal A (2005) Treating cancer with stem cells. Medical Engineer website (accessed at http://www.medicalengineer.co.uk/)
Scripture CD, Figg WD (2006) Nat Rev Cancer 6:546
Doddareddy MR, Cho YS, Koh HY, Pae AN (2004) Bioorg Med Chem 12:3977
Bhongade BA, Gadad AK (2004) Bioorg Med Chem 12:2797
Keen N, Taylor S (2004) Nat Rev Cancer 4:927
Faivre S, Kroemer G, Raymond E (2006) Nat Rev Drug Discov 5:671
Suksamrarn S, Komutiban O, Ratananukul P, Chimnoi N, Lartpornmatulee N, Suksamrarn A (2006) Chem Pharm Bull 54:301
Matsumoto K, Akao Y, Kobayashi E, Ohguchi K, Ito T, Tanaka T, Iinuma M, Nozawa Y (2003) J Nat Prod 66:1124
Matsumoto K, Akao Y, Yi H, Ohguchi K, Ito T, Tanaka T, Kobayashi E, Iinuma M, Nozawa Y (2004) Bioorg Med Chem 12:5799
Moongkarndi P, Kosem N, Kaslungka S, Lanratana O, Pongpan N, Neungton N (2004) J Ethnopharmacol 90:161
Ho CK, Huang YL, Chen CC (2002) Planta Med 68:975
Pedro M, Cerqueira F, Sousa ME, Nascimento MS, Pinto M (2002) Bioorg Med Chem 10:3725
Lin CN, Liou SJ, Lee TH, Chang YC, Won SJ (1996) J Pharm Pharmacol 48:539
Liou SS, Shich WL, Cheng TH, Won SJ, Lin CN (1993) J Pharm Pharmacol 45:791
Sultanbawa MUS (1980) Tetrahedron 36:1465
Suksamrarn S, Suwannapoch N, Phakhodee W, Thanuhiranlert J, Ratananukul P, Chimnoi N, Suksamrarn A (2003) Chem Pharm Bull 51:857
Peres V, Nagem TJ (1997) Phytochemistry 44:191
Bennett GJ, Lee HH (1989) Phytochemistry 28:967
Mandal S, Das PC, Joshi PC (1992) J Indian Chem Soc 69:611
Pinto MMM, Sousa ME, Nascimento MSJ (2005) Curr Med Chem 12:2517
Cramer RD, Patterson DEIII, Bunce JD (1988) J Am Chem Soc 110:5959
Klebe G, Abraham U, Mietzner T (1994) J Med Chem 37:4130
Depriest SA, Mayer D, Naylor CB, Marshall GR (1993) J Am Chem Soc 115:5372
Hannongbua S, Lawtrakul L, Sotriffer CA, Rode BM (1996) Quant Struct Act Relat 15:389
Boström J, Böhm M, Gundertofte K, Klebe G (2003) J Chem Inf Comput Sci 43:1020
Kubinyi H (1993) QSAR: Hansch analysis and related approaches. Wiley-VCH, Weinheim
Hansch C, Leo A (1995) Exploring QSAR fundamentals and applications in chemistry and biology. American Chemical Society, Washington, DC
Böhm M, Stürzebecher J, Klebe G (1999) J Med Chem 42:458
Suksamrarn S, Sudta P, Kunchanawatta S, Ratananukul P, Suksamrarn A (2007) Thai patent application, No. 0701003620, 27 June 2007, p 36
Skehan P, Storeng R, Scudiero D, Monks A, McMahon J, Vistica D, Warren JT, Bokesch H, Kenny S, Boyd MR (1990) J Natl Cancer Inst 82:1107
Tripos Associates (2008) SYBYL 7.0 molecular modelling software. Tripos Associates, Inc., St. Louis, MO
Frisch MJ, Trucks GW, Schlegel HB, Scuseria, GE, Robb MA, Cheeseman JR, Montgomery Jr. JA, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M,Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2004) Gaussian 03, revision B.04. Gaussian, Inc., Wallingford, CT
Wold S, Albano C, Dunn J, Edlund U, Esbensen K, Geladi P, Hellberg S, Johansson E, Lindberg W, Sjosstrom M (1987) Multivariate data analysis in chemistry. In: Kowalski B (ed) Chemometrics: mathematics and statistics in chemistry. Reidel, Dordrecht
Acknowledgments
This work is supported by the Thailand Research Fund, the Postgraduate Education and Research Program in Chemistry (PERCH) and in Petroleum and Petrochemical Technology (PPT), and a research team strengthening grant from The National Centre for Genetic Engineering and Biotechnology (BIOTEC). Phornphimon Maitarad is grateful to TGIST for a scholarship and the Graduate school of KU for scholarships and to Ben Parslew for English editing. We would like also to thank the High Performance Computing Center of the National Electronics and Computer Technology Center (NECTEC), the Laboratory for Computational and Applied Chemistry (LCAC), and the computing center of KU for research facilities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Suphavanich, K., Maitarad, P., Hannongbua, S. et al. CoMFA and CoMSIA studies on a new series of xanthone derivatives against the oral human epidermoid carcinoma (KB) cancer cell line. Monatsh Chem 140, 273–280 (2009). https://doi.org/10.1007/s00706-008-0014-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00706-008-0014-5