Abstract
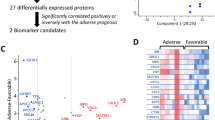
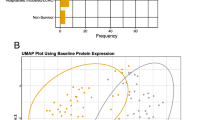
The ongoing COVID-19 pandemic caused by SARS-CoV-2 has prompted global concern due to its profound impact on public health and the economy. Effective treatment of COVID-19 patients in the acute phase or of those with long COVID is a major challenge. Using data-independent acquisition (DIA) technology, we performed proteomic profiling on plasma samples from 22 COVID-19 patients and six healthy controls at Dazhou Central Hospital. Random forest and least absolute shrinkage and selection operator algorithms were used for analysis at various COVID-19 treatment stages. We identified 79 proteins that were differentially expressed between COVID-19 patients and healthy controls, mainly involving pathways associated with cell processes and binding. Across different treatment stages of COVID-19, five proteins—PI16, GPLD1, IGFBP3, KRT19, and VCAM1—were identified as potential molecular markers for dynamic disease monitoring. Furthermore, the proteins BTD, APOM, IGKV2-28, VWF, C4BPA, and C7 were identified as candidate biomarkers for distinguishing between SARS-CoV-2 positivity and negativity. Analysis of protein change profiles between the follow-up and healthy control groups highlighted cardiovascular changes as a concern for patients recovering from COVID-19. Our study revealed the infection profiles of SARS-CoV-2 at the protein expression level comparing different phases of COVID-19. DIA mass spectrometry analysis of plasma samples from COVID-19 patients undergoing treatment identified key proteins involved in signaling pathways that might be used as markers of the recovery phase. These findings provide insight for the development of therapy options and suggest potential blood biomarkers for COVID-19.






Similar content being viewed by others
Data availability
Data from this study are available through the corresponding author.
References
Lu H, Stratton CW, Tang YW (2020) Outbreak of pneumonia of unknown etiology in Wuhan, China: the mystery and the miracle. J Med Virol 92:401–402. https://doi.org/10.1002/jmv.25678
Hui DS, Madani TA, Ntoumi F, Kock R, Dar O, Ippolito G, McHugh TD, Memish ZA, Drosten C, Zumla A, Petersen E (2020) The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health—the latest 2019 novel coronavirus outbreak in Wuhan, China. Int J Infect Dis 91(2020):264–266. https://doi.org/10.1016/j.ijid.2020.01.009
Chipimo PJ, Barradas DT, Kayeyi N, Zulu PM, Muzala K, Mazaba ML, Hamoonga R, Musonda K, Monze M, Kapata N, Sinyange N, Simwaba D, Kapaya F, Mulenga L, Chanda D, Malambo W, Ngosa W, Hines J, Yingst S, Agolory S, Mukonka V (2020) First 100 persons with COVID-19—Zambia, March 18–April 28, 2020. MMWR Morb Mortal Wkly Rep 69:1547–1548. https://doi.org/10.15585/mmwr.mm6942a5
Klauschen F, Muller KR, Binder A, Bockmayr M, Hagele M, Seegerer P, Wienert S, Pruneri G, de Maria S, Badve S, Michiels S, Nielsen TO, Adams S, Savas P, Symmans F, Willis S, Gruosso T, Park M, Haibe-Kains B, Gallas B, Thompson AM, Cree I, Sotiriou C, Solinas C, Preusser M, Hewitt SM, Rimm D, Viale G, Loi S, Loibl S, Salgado R, Denkert C, G. International Immuno-Oncology Biomarker Working (2018) Scoring of tumor-infiltrating lymphocytes: from visual estimation to machine learning. Semin Cancer Biol 52:151–157. https://doi.org/10.1016/j.semcancer.2018.07.001
Yu N, Li W, Kang Q, Xiong Z, Wang S, Lin X, Liu Y, Xiao J, Liu H, Deng D, Chen S, Zeng W, Feng L, Wu J (2020) Clinical features and obstetric and neonatal outcomes of pregnant patients with COVID-19 in Wuhan, China: a retrospective, single-centre, descriptive study. Lancet Infect Dis 20:559–564. https://doi.org/10.1016/S1473-3099(20)30176-6
Vetter P, Vu DL, L’Huillier AG, Schibler M, Kaiser L, Jacquerioz F (2020) Clinical features of covid-19. BMJ 369:m1470. https://doi.org/10.1136/bmj.m1470
Guo CX, He L, Yin JY, Meng XG, Tan W, Yang GP, Bo T, Liu JP, Lin XJ, Chen X (2020) Epidemiological and clinical features of pediatric COVID-19. BMC Med 18:250. https://doi.org/10.1186/s12916-020-01719-2
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395(2020):497–506. https://doi.org/10.1016/S0140-6736(20)30183-5
Yang W, Cao Q, Qin L, Wang X, Cheng Z, Pan A, Dai J, Sun Q, Zhao F, Qu J, Yan F (2020) Clinical characteristics and imaging manifestations of the 2019 novel coronavirus disease (COVID-19): a multi-center study in Wenzhou city, Zhejiang, China. J Infect 80:388–393. https://doi.org/10.1016/j.jinf.2020.02.016
Bhatraju PK, Ghassemieh BJ, Nichols M, Kim R, Jerome KR, Nalla AK, Greninger AL, Pipavath S, Wurfel MM, Evans L, Kritek PA, West TE, Luks A, Gerbino A, Dale CR, Goldman JD, O’Mahony S, Mikacenic C (2020) Covid-19 in critically ill patients in the Seattle Region—case series. N Engl J Med 382:2012–2022. https://doi.org/10.1056/NEJMoa2004500
Guo T, Fan Y, Chen M, Wu X, Zhang L, He T, Wang H, Wan J, Wang X, Lu Z (2020) Cardiovascular implications of fatal outcomes of patients with coronavirus disease 2019 (COVID-19). JAMA Cardiol 5:811–818. https://doi.org/10.1001/jamacardio.2020.1017
Huang L, Zhang X, Zhang X, Wei Z, Zhang L, Xu J, Liang P, Xu Y, Zhang C, Xu A (2020) Rapid asymptomatic transmission of COVID-19 during the incubation period demonstrating strong infectivity in a cluster of youngsters aged 16–23 years outside Wuhan and characteristics of young patients with COVID-19: a prospective contact-tracing study. J Infect 80:e1–e13. https://doi.org/10.1016/j.jinf.2020.03.006
Patel R, Babady E, Theel ES, Storch GA, Pinsky BA, St George K, Smith TC, Bertuzzi S (2020) Report from the American Society for Microbiology COVID-19 international summit, 23 March 2020: value of diagnostic testing for SARS-CoV-2/COVID-19. MBio. https://doi.org/10.1128/mBio.00722-20
Lan J, Ge J, Yu J, Shan S, Zhou H, Fan S, Zhang Q, Shi X, Wang Q, Zhang L, Wang X (2020) Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581:215–220. https://doi.org/10.1038/s41586-020-2180-5
Polack FP, Thomas SJ, Kitchin N, Absalon J, Gurtman A, Lockhart S, Perez JL, Perez Marc G, Moreira ED, Zerbini C, Bailey R, Swanson KA, Roychoudhury S, Koury K, Li P, Kalina WV, Cooper D, Frenck Jr RW, Hammitt LL, Tureci O, Nell H, Schaefer A, Unal S, Tresnan DB, Mather S, Dormitzer PR, Sahin U, Jansen KU, Gruber WC (2020) C.C.T. Group, safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N Engl J Med 383:2603–2615. https://doi.org/10.1056/NEJMoa2034577
Schmith VD, Zhou JJ, Lohmer LRL (2020) The approved dose of Ivermectin alone is not the ideal dose for the treatment of COVID-19. Clin Pharmacol Ther 108:762–765. https://doi.org/10.1002/cpt.1889
Garvin MR, Alvarez C, Miller JI, Prates ET, Walker AM, Amos BK, Mast AE, Justice A, Aronow B, Jacobson D (2020) A mechanistic model and therapeutic interventions for COVID-19 involving a RAS-mediated bradykinin storm. Elife. https://doi.org/10.7554/eLife.59177
Shu T, Ning W, Wu D, Xu J, Han Q, Huang M, Zou X, Yang Q, Yuan Y, Bie Y, Pan S, Mu J, Han Y, Yang X, Zhou H, Li R, Ren Y, Chen X, Yao S, Qiu Y, Zhang DY, Xue Y, Shang Y, Zhou X (2020) Plasma proteomics identify biomarkers and pathogenesis of COVID-19. Immunity 53:1108-1122e1105. https://doi.org/10.1016/j.immuni.2020.10.008
Shen B, Yi X, Sun Y, Bi X, Du J, Zhang C, Quan S, Zhang F, Sun R, Qian L, Ge W, Liu W, Liang S, Chen H, Zhang Y, Li J, Xu J, He Z, Chen B, Wang J, Yan H, Zheng Y, Wang D, Zhu J, Kong Z, Kang Z, Liang X, Ding X, Ruan G, Xiang N, Cai X, Gao H, Li L, Li S, Xiao Q, Lu T, Zhu Y, Liu H, Chen H, Guo T (2020) Proteomic and metabolomic characterization of COVID-19 patient Sera. Cell 182:59-72e15. https://doi.org/10.1016/j.cell.2020.05.032
Chu DKW, Pan Y, Cheng SMS, Hui KPY, Krishnan P, Liu Y, Ng DYM, Wan CKC, Yang P, Wang Q, Peiris M, Poon LLM (2020) Molecular diagnosis of a novel coronavirus (2019-nCoV) causing an outbreak of pneumonia. Clin Chem 66:549–555. https://doi.org/10.1093/clinchem/hvaa029
Echavarria M (2008) Adenoviruses in immunocompromised hosts. Clin Microbiol Rev 21:704–715. https://doi.org/10.1128/CMR.00052-07
Julkunen I (1984) Serological diagnosis of parainfluenza virus infections by enzyme immunoassay with special emphasis on purity of viral antigens. J Med Virol 14:177–187. https://doi.org/10.1002/jmv.1890140212
C. Centers for Disease, Prevention (2006) New laboratory assay for diagnostic testing of avian influenza A/H5 (Asian lineage). MMWR Morb Mortal Wkly Rep 55:127
Paules C, Subbarao K (2017) Influenza. Lancet 390:697–708. https://doi.org/10.1016/S0140-6736(17)30129-0
Griffiths C, Drews SJ, Marchant DJ (2017) Respiratory syncytial virus: infection, detection, and new options for prevention and treatment. Clin Microbiol Rev 30:277–319. https://doi.org/10.1128/CMR.00010-16
Xu X, Chen P, Wang J, Feng J, Zhou H, Li X, Zhong W, Hao P (2020) Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Sci China Life Sci 63:457–460. https://doi.org/10.1007/s11427-020-1637-5
Bernstein LR, Zhang L (2020) Gallium maltolate has in vitro antiviral activity against SARS-CoV-2 and is a potential treatment for COVID-19. Antivir Chem Chemother 28:2040206620983780. https://doi.org/10.1177/2040206620983780
Alhogbani T (2016) Acute myocarditis associated with novel Middle east respiratory syndrome coronavirus. Ann Saudi Med 36:78–80. https://doi.org/10.5144/0256-4947.2016.78
Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z (2020) Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA 323:1061–1069. https://doi.org/10.1001/jama.2020.1585
Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L (2020) Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet 395:507–513. https://doi.org/10.1016/S0140-6736(20)30211-7
Chu Y, Li J, Zeng Z, Huang B, Zhao J, Liu Q, Wu H, Fu J, Zhang Y, Zhang Y, Cai J, Zeng F (2020) A novel model based on CXCL8-derived radiomics for prognosis prediction in colorectal cancer. Front Oncol 10:575422. https://doi.org/10.3389/fonc.2020.575422
Nicora G, Vitali F, Dagliati A, Geifman N, Bellazzi R (2020) Integrated multi-omics analyses in oncology: a review of machine learning methods and tools. Front Oncol 10:1030. https://doi.org/10.3389/fonc.2020.01030
Mao L, Jin H, Wang M, Hu Y, Chen S, He Q, Chang J, Hong C, Zhou Y, Wang D, Miao X, Li Y, Hu B (2020) Neurologic manifestations of hospitalized patients with coronavirus disease 2019 in Wuhan, China. JAMA Neurol 77:683–690. https://doi.org/10.1001/jamaneurol.2020.1127
Liu L, Wang Y, Cao ZY, Wang MM, Liu XM, Gao T, Hu QK, Yuan WJ, Lin L (2015) Up-regulated TLR4 in cardiomyocytes exacerbates heart failure after long-term myocardial infarction. J Cell Mol Med 19:2728–2740. https://doi.org/10.1111/jcmm.12659
Chen L, Li X, Chen M, Feng Y, Xiong C (2020) The ACE2 expression in human heart indicates new potential mechanism of heart injury among patients infected with SARS-CoV-2. Cardiovasc Res 116:1097–1100. https://doi.org/10.1093/cvr/cvaa078
Crackower MA, Sarao R, Oudit GY, Yagil C, Kozieradzki I, Scanga SE, Oliveira-dos-Santos AJ, da Costa J, Zhang L, Pei Y, Scholey J, Ferrario CM, Manoukian AS, Chappell MC, Backx PH, Yagil Y, Penninger JM (2002) Angiotensin-converting enzyme 2 is an essential regulator of heart function. Nature 417:822–828. https://doi.org/10.1038/nature00786
Acknowledgements
We are thankful to all the doctors and nurses of the Infectious Diseases Department at Dazhou Central Hospital for the help they gave us and all the work they accomplished during the epidemic.
Funding
This work was supported by Dazhou Science and Technology Bureau, Dazhou, Sichuan, China (grant numbers ZDYF0001, 22ZDYF0020, 21ZDYF0021, and 21CDDZ-26).
Author information
Authors and Affiliations
Contributions
F.Z.X., F.Z.W., and X.H. conceived the idea and designed the study. X.L., X.H. G.D., and S.L. performed most of the experiments and analyzed and interpreted data. C.L., X.Z., J.L., and S.H. performed part of the experiments and data collection. G.D. and X.L. wrote the original draft and revised the manuscript. F.Z.X., F.Z.W., and X.H. revised the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This study was performed in accordance with the principles of the Declaration of Helsinki. Approval was granted by the Dazhou Central Hospital Ethics Committee (Date: April 15, 2020/no. 000015).
Additional information
Handling Editor: Pablo Pineyro.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Ding, G., Li, S. et al. Proteomic characteristics of the treatment trajectory of patients with COVID-19. Arch Virol 169, 84 (2024). https://doi.org/10.1007/s00705-024-05991-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-024-05991-y